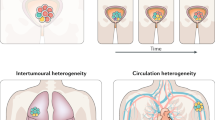
Abstract
Chemotherapy-resistant urothelial carcinoma has no uniformly curative therapy. Understanding how selective pressure from chemotherapy directs the evolution of urothelial carcinoma and shapes its clonal architecture is a central biological question with clinical implications. To address this question, we performed whole-exome sequencing and clonality analysis of 72 urothelial carcinoma samples, including 16 matched sets of primary and advanced tumors prospectively collected before and after chemotherapy. Our analysis provided several insights: (i) chemotherapy-treated urothelial carcinoma is characterized by intra-patient mutational heterogeneity, and the majority of mutations are not shared; (ii) both branching evolution and metastatic spread are very early events in the natural history of urothelial carcinoma; (iii) chemotherapy-treated urothelial carcinoma is enriched with clonal mutations involving L1 cell adhesion molecule (L1CAM) and integrin signaling pathways; and (iv) APOBEC-induced mutagenesis is clonally enriched in chemotherapy-treated urothelial carcinoma and continues to shape the evolution of urothelial carcinoma throughout its lifetime.
Similar content being viewed by others
Main
Urothelial carcinoma results in 15,000 deaths annually in the United States1. Individuals with metastatic urothelial carcinoma are standardly treated with platinum-based chemotherapy2,3,4,5. However, nearly all of these cancers will progress and develop chemotherapy resistance3,6,7. Ultimately, the majority of patients will die of metastatic chemotherapy-resistant urothelial carcinoma2,3,5. Little is known about the clonal architecture of advanced, chemotherapy-treated urothelial carcinoma or the evolutionary dynamics that lead to metastasis and chemotherapy resistance. Large genomic studies like The Cancer Genome Atlas (TCGA) have focused only on untreated, primary tumors8. In particular, the extent to which chemotherapy-treated tumors share the genetic profile of the primary tumor remains unknown.
To understand the relative contributions of different subclones and the effects of chemotherapy as a selective pressure in urothelial carcinoma, we performed whole-exome sequencing and clonality analysis of matched sets of primary, metastatic and germline samples. Because cancers are genetically heterogeneous, we set out to address two fundamental questions: (i) What is the degree of clonal divergence between primary and metastatic urothelial carcinoma? and (ii) How does chemotherapy influence the genomic landscape of tumor cell populations in advanced and metastatic urothelial carcinoma?
We employed the computational framework of CLONET (CLONality Estimate in Tumors) that we developed previously9 (Online Methods) to adjust genomic events for tumor purity and ploidy and then determine the relative abundance of tumor-cell subpopulations through clonality analysis of genomic lesions. By comparing the frequency and patterns of CLONET-adjusted events between primary and metastatic tumors obtained from different anatomical sites and at different time points over each patient's clinical course, we were able to reconstruct phylogenetic trees and compare clonal evolutionary patterns across the study cohort. We then proceeded to examine clonally enriched genomic signatures and trace the evolutionary footprints of mutagenesis mechanisms, including those of the APOBEC3 family of cytidine deaminases, during each cancer's evolution.
Results
Clonal mutational heterogeneity in chemotherapy-treated urothelial carcinoma
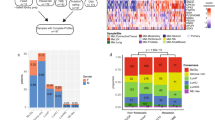
To characterize the clonal architecture of advanced, chemotherapy-treated urothelial carcinoma, we performed whole-exome sequencing of 72 prospectively collected urothelial tumors from 32 patients, including 16 matched sets of primary, metastatic urothelial carcinomas and germline samples and 2 rapid autopsy cases (Fig. 1 and Supplementary Table 1). The study was designed to enrich for patients with advanced disease, as 28 out of 32 (88%) patients either presented with or developed metastatic disease during the study period (Fig. 1 and Supplementary Table 1). Overall, the most frequent mutations and copy number alterations (CNAs) observed in our cohort were consistent with the results of the TCGA data set of untreated urothelial carcinoma8 (Supplementary Fig. 1 and Supplementary Table 2). We observed no statistically significant difference in the number of single-nucleotide variants (SNVs) between pre-chemotherapy and post-chemotherapy tumors (Supplementary Fig. 2).
(a) The bar graph shows the number of tumor samples sequenced from each study subject. The treatment status of each sample is indicated by color within each bar. For each study subject, sex, smoking status, development of metastases and history of intravesical Bacillus Calmette–Guérin (BCG) therapy for pre-existing non-muscle-invasive bladder cancer are represented on the bottom. (b) Schematic illustrating the anatomical sites of primary and metastatic tumor samples. Numbers correspond to the number of tumors at each site, and sites are colored by treatment status of the tumor as in a.
To compare the clonal structure of pre-chemotherapy and post-chemotherapy tumors across the study cohort, we investigated the number of private and shared mutations between the pre-chemotherapy and post-chemotherapy tumors within each patient as a fraction of the total mutational burden (Fig. 2a). On average, only 28.4% (range 0.2–76.4%) of mutations were shared by pre- and post-chemotherapy samples (Fig. 2a). This effect was consistent across primary–primary tumor pairs and primary–metastatic tumor pairs (P = 0.17, Wilcoxon test) (Fig. 2a). Surprisingly, even mutations in previously reported driver genes8, including PIK3CA, KMT2D (MLL2), ATM and TP53, were not consistently shared by matched pre-chemotherapy and post-chemotherapy tumors (Fig. 2b). We confirmed these findings with targeted sequencing of 250 common driver genes, achieving an average coverage of 400× and excellent concordance with variant allele frequencies obtained from whole-exome sequencing (Pearson correlation = 0.93, P < 10−171) (Supplementary Fig. 3). Some post-chemotherapy tumors even evolved to develop different mutations in the same key gene. For example, in patient WCM077, the primary pre-chemotherapy tumor and the pelvic post-chemotherapy lymph node metastasis shared a TP53 p.Tyr234Cys mutation, while the post-chemotherapy lung metastasis had a separate private TP53 p.Gly266Val mutation that was not shared by the primary tumor (Fig. 2b and Supplementary Fig. 4). Taken together, our results demonstrate pronounced mutational heterogeneity in tumor samples from the same patient and suggest that chemotherapy is associated with a significant change in the mutational landscape of advanced urothelial carcinoma.
(a) Percentages of shared and unique SNVs in matched chemotherapy-naive and chemotherapy-treated urothelial carcinoma tumors. (b) Discordance in mutational status for selected driver genes between chemotherapy-naive and chemotherapy-treated urothelial carcinoma tumors. Each column represents a paired set of pre- and post-chemotherapy tumors obtained from an individual subject. Individuals with more than one set of paired tumors are highlighted with matching color designations. Primary–primary tumor pairs are represented on the left, and primary–metastatic tumor pairs are represented on the right. ERCC2 missense mutations were identified in two tumor samples, WCM259_3 (p.Gln755His) and WCM141_1 (p.Thr484Met); matched pre- and post-chemotherapy samples were not available for either patient.
Early branching evolution in chemotherapy-treated urothelial carcinoma
We conducted a phylogenetic analysis of 21 sets of matched tumors from patients for whom at least two tumor samples were available using the parsimony ratchet method10 (Fig. 3 and Supplementary Fig. 5). This analysis identified a pattern of early branching evolution, with several successive waves of clonal expansion occurring early in each patient's urothelial carcinoma. Surprisingly, in every reconstructed evolutionary tree, the resulting position of the primary tumor was as a branch, indicating that the ancestral clone gave rise to multiple cell populations that evolved in parallel during the early stages of tumor evolution.
Urothelial carcinoma phylogenetic trees (top) and the fraction of tumor cells with shared and private clonally adjusted mutations for each tumor sample (bottom) are shown for six patients with three or more tumor samples per patient. All cases showed early metastatic spread, occurring as branching evolution occurred, early in the natural history of the disease. All primary tumors temporally preceded the development of metastasis but were assigned “branch” values on the basis of genomic distance, suggesting early branching evolution and metastatic spread. Different alterations occurring in the same gene are designated with asterisks. LN, lymph node.
In order to better understand the pattern of clonal evolution during the course of chemotherapy, we followed another individual with urothelial carcinoma from the time of diagnosis through death over a period of 16 months. We collected a total of 12 samples from eight anatomical sites and at three different time points: the primary, untreated tumor was obtained by transurethral resection of bladder tumor (TURBT) and was acquired at initial diagnosis; four areas of residual primary bladder tumor and one pelvic lymph node were obtained by radical cystectomy and lymph node dissection following four cycles of gemcitabine–cisplatin chemotherapy; and six metastases were obtained at time of rapid autopsy after docetaxel–ramucirumab therapy (four distant lymph nodes, two liver metastases) (Fig. 4a). We analyzed the genomes of these tumors to characterize the full evolutionary arc of the cancer from the time of diagnosis to death. Metastatic tumors obtained at autopsy harbored high clonal fractions (the percentages of clonal alterations compared to all alterations in a sample) (mean 83%, range 69–91%) (Fig. 4b). Allele-specific copy number analysis identified a subclonal heterozygous deletion of the tumor-suppressor gene CDKN2A in the primary tumor that evolved into a homozygous clonal deletion in the distant metastatic lymph node and liver lesions obtained from the autopsy. This was confirmed by FISH (Fig. 4c), suggesting that CDKN2A loss was selected as the tumor evolved under pressure from chemotherapy. Comparison of clonality-adjusted frequencies of somatic alterations across all tumor samples from this patient showed substantial heterogeneity. This heterogeneity followed several distinct patterns. Certain subclonal mutations (e.g., in RYR2, ANKRD62, NCOA3 and LSS) that were present in the primary tumor persisted and became enriched in the chemotherapy-treated metastatic lesions. Other subclonal mutations (e.g., in POLD2, FOXP1, FGFR4, TRRAP and EGFR) present in the primary tumor were not observed in the metastatic lesions and were considered private to the primary tumor. We also observed other private mutations that were present exclusively in the metastatic lesions (Fig. 4d). In fact, each tumor in this patient harbored a unique set of private mutations (mean 26.3, range 5–138) that was not shared with any of the other tumors (Fig. 4d).
Analysis is shown of 12 tumor samples collected during disease progression and at the time of death of patient WCM117. (a) Top, circles represent the sites of the sequenced tumors. Bottom, timeline and clinical course (vertical lines) in the natural history of the disease. (b) CLONET-adjusted variant allele frequencies of selected mutations. Each dot represents the fraction of cells harboring the corresponding mutation. (c) FISH for CDKN2A (magenta) and reference (green) probes in primary cystectomy tumor 4 (right) and the tumor from the right supraclavicular lymph node collected at the time of autopsy (left). Scale bars, 5 μm. The plot shows allele-specific copy number. Axes correspond to different alleles, and dots represent CDKN2A allele-specific copy number. Arrows indicate the CDKN2A allele-specific copy number in the tumor samples represented in the FISH images. (d) CLONET-adjusted shared and private mutations. The fractions of tumor cells harboring each mutation are represented by shades of green (scale, upper left corner). ND (gray), clonality information was not available. (e) Reconstruction of the cancer evolutionary tree. “WT” denotes the inferred last ancestor with a wild-type genome that acquired a series of mutations during oncogenesis. The length of the branches corresponds to the distance between two tumors based on the number of shared mutations. Samples are color coded consistently across panels: yellow, primary, pre-chemotherapy TURBT sample; orange, four different areas of bladder tumors and a pelvic lymph node metastatic lesion removed through cystectomy and lymph node dissection, respectively, after gemcitabine–cisplatin therapy; green, metastatic lesions after docetaxel–ramucirumab therapy removed during autopsy. LN, lymph node; met, metastasis.
Using clonally adjusted non-silent mutations from each tumor, we reconstructed the evolutionary tree of this patient's urothelial carcinoma (Online Methods) (Fig. 4e). This reconstruction showed a complex branching evolutionary pattern. Early truncal mutations (in RYR2, ANKRD62, NCOA3 and LSS) were present in the initial founder clone and shared by all descendent clones. At each clonal divergence node, additional mutations were acquired, including mutations in driver genes such as TP53 and TSC1. At the time of the patient's initial cancer diagnosis, at least five waves of clonal expansion (each represented by a branching node, numbered 1–5) (Fig. 4e) had already occurred from the lowest common ancestor, as observed in the mutational analysis of the TURBT tumor. This pattern suggests that branching evolution was a very early event in this tumor's development. While the untreated TURBT tumor had a high fraction of tumor cells harboring founder mutations (Fig. 4e), it also had the greatest genetic distance from all other tumors. We attribute this long genetic distance to the high number of private mutations in the TURBT primary tumor sample (138 mutations) (Fig. 4d and Supplementary Table 3), many of which involve genes implicated in cellular responses to cisplatin, including POLD2 and FOXP1 (refs. 11, 12, 13, 14). After neoadjuvant chemotherapy, these mutations disappeared from the evolutionary record, as the cancer cells harboring them were likely eradicated by treatment and thus were not observed subsequently in any of the other tumors (Fig. 4d,e).
One of the earliest cancer cell clades had already separated at the first divergence node, resulting in a population that metastasized to a pelvic lymph node (Fig. 4e). This lymph node was later dissected and removed at the time of radical cystectomy. The surgical removal of this lymph node possibly eliminated mutations from this particular clade, preventing them from entering the genetic pool that later contributed to the development of additional distant metastases. On the other hand, by the time cystectomy took place, distant metastatic spread had already occurred, originating from a different cancer cell clade that branched at divergence node 5 to give rise to both bladder tumor 2 and all of the lymph node and visceral distant metastases that were later collected at the time of the rapid autopsy. This transition from the primary to the metastatic state was marked by the acquisition of a non-silent mutation in TSPAN8 (which encodes tetraspanin 8), a well-recognized pro-metastatic and angiogenesis-promoting gene15,16,17 (Fig. 4d,e). This sequence of events supports a possible role for this mutation as a key driver underlying the metastatic spread of this patient's urothelial carcinoma. Altogether, our data strongly suggest that both branching evolution and metastatic spread are very early events in the natural history of urothelial carcinoma.
Heterogeneity in copy number alterations
To understand how CNAs evolve throughout the lifetime of urothelial carcinoma and during chemotherapy, we conducted a detailed analysis of somatic genomic aberrations. Hierarchical clustering of 44 tumor samples based on allele-specific CNAs (Online Methods) showed two distinct clusters (Fig. 5a). Cluster A was defined by 9p21 (CDKN2A, CDKN2B and MTAP) deletions in the setting of a euploid copy number background. Cluster B was characterized by the enrichment of several amplifications, including those of 1q21.1 (SETDB1 and MLLT11) (P = 0.0002, Fisher's exact test) and 6p22.3 (E2F3) (P = 0.001, Fisher's exact test) (Supplementary Table 4). This cluster was also enriched with TP53 mutations (P = 0.0001, Fisher's exact test). The same CNA clusters were consistently observed when we extended our cohort with TCGA-derived whole-exome sequencing data from untreated urothelial carcinoma samples8 (Supplementary Fig. 6). We also observed an enrichment of tumors belonging to TCGA bladder cancer cluster III (“basal/squamous-like”)8 in our copy number cluster A (P = 0.02, Fisher's exact test) (Supplementary Fig. 7). There was no statistically significant differential enrichment in the number of metastatic samples or chemotherapy-treated samples between the two clusters, suggesting that these clusters reflect a relatively stable feature of urothelial carcinoma biology that is independent of treatment effects or disease stage. Overall, tumor samples from the same patient tended to cluster in the same group despite the presence of some private CNAs. To quantify the degree of within- and between-patient heterogeneity, we interrogated CNAs for a panel of more than 30,000 genes from the Ensembl catalog18. For each pair of tumor samples, we computed the Hamming distance (HD) as the ratio between the number of genes that have different discrete copy numbers and the total number of genes analyzed. We identified a significant difference between intra-patient tumor pairs (median HD = 0.20) and inter-patient tumor pairs (median HD = 0.53) (P = 0.00000003, Wilcoxon test) (Fig. 5b). This limited intra-patient as compared to inter-patient heterogeneity suggests that each patient's cancer is relatively stable during evolution at the copy number level.
(a) Copy number gains are represented in red, and copy number losses are represented in blue. Each column corresponds to one tumor sample. Clinical annotations and TP53 mutational status are given above. Matched samples from the same patient are represented in the same color in the “matched samples” annotation track. Selected genes harboring frequent CNAs are listed on the left. (b) CNA differences (Hamming distance). Dots correspond to sample pairs in a from the same (left) and different (right) patients. Left boxplot: median 0.2, IQR (0.13, 0.28), whiskers (0.05, 0.49). Right boxplot: median 0.53, IQR (0.42, 0.66), whiskers (0.07, 0.93).
We investigated the frequency of combined CNAs and mutations constituting the ATM–RB–FANCC signature that was previously associated with chemotherapy response in the neoadjuvant setting19. We identified this signature in 11 of 15 (73.3%) pre-chemotherapy tumors and in 11 of 29 (37.9%) post-chemotherapy tumors (P = 0.05) in our cohort, supporting the hypothesis that clones harboring these molecular alterations are likely to disappear after treatment, and are superseded by tumor clones with wild-type ATM, RB and FANCC that eventually progress to metastatic, chemotherapy-resistant disease.
Clonal enrichment of mutations in chemotherapy-treated urothelial carcinoma
We hypothesized that the evolution of chemotherapy-treated urothelial carcinoma would proceed in a direction that ultimately leads to the selection of mutations conferring proliferative or chemotherapy-resistance advantages. Using density analysis of CLONET-adjusted variant allele frequencies between pre-chemotherapy and post-chemotherapy tumors, we observed a significant increase in the number of clonal mutations in the post-chemotherapy samples across the study cohort (P = 0.0134, Fisher's exact test) (Fig. 6a and Supplementary Fig. 8), confirming the association between chemotherapy and increased clonality. To dissect the functional impact of these clonally enriched mutations, we conducted gene set enrichment analysis (GSEA) to identify enriched pathways in post-chemotherapy samples (Fig. 6b). This analysis showed a clonal enrichment of mutations in pathways involved in the transmembrane transport of small molecules (odds ratio = 1.9; false discovery rate (FDR) = 0.002), suggesting that mutations in multidrug-resistance genes may have a role in the progression of advanced, chemotherapy-treated urothelial carcinoma. In addition, GSEA demonstrated a significant enrichment for mutations in genes mediating the L1 cell adhesion molecule (L1CAM) (odds ratio = 1.9; FDR = 0.12) and integrin (odds ratio = 2.8; FDR = 0.02) signaling pathways. The majority of mutations identified in the L1CAM and integrin signaling pathways (83% and 90%, respectively) were missense mutations, which can conceivably lead to gain-of-function molecular changes that activate these pathways. These results suggest a role for mutations in the L1CAM and integrin signaling pathways in conferring a selective advantage for resistance to chemotherapy in urothelial carcinoma. Mutations in these pathways may also provide a potential mechanistic link between metastatic spread, the tumor microenvironment and drug resistance, which cooperate to promote tumor survival20,21,22,23,24.
(a) Density plot representing the clonality of non-silent SNVs on the x axis and the density distribution on the y axis. Pre-chemotherapy urothelial carcinoma tumor samples are represented in blue, and post-chemotherapy tumor samples are represented in green. Boxplots (right) represent the percentage of clonal SNVs in pre-chemotherapy and post-chemotherapy tumors, demonstrating a significant increase in clonality in post-chemotherapy tumors. Left boxplot: median 0.5, IQR (0.41, 0.71), whiskers (0.24, 0.88). Right boxplot: median 0.83, IQR (0.61, 0.89), whiskers (0.22, 0.94). (b) Results of GSEA for mutations in chemotherapy-treated urothelial carcinoma. Each node represents one REACTOME pathway. Pathways enriched in post-chemotherapy samples are represented as red nodes. Node size corresponds to the number of genes in each pathway.
Mutagenesis mechanisms driving the evolution of urothelial carcinoma
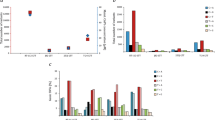
To characterize the evolution of mutational signatures in advanced, chemotherapy-treated urothelial carcinoma, we examined the six possible single-base substitutions (C>A, C>G, C>T, T>A, T >C, and T>G). We identified significant differences in these mutational patterns between chemotherapy-naive and chemotherapy-treated tumors, with a statistically significant enrichment of C>A and C>G changes in the chemotherapy-treated tumors (Fig. 7a).
(a) Composite bar graphs representing the distribution of all possible nucleotide substitutions for non-silent SNVs in sequenced pre-chemotherapy (left) and post-chemotherapy (right) urothelial carcinoma tumors. Asterisks indicate statistically significant changes (significance level = 0.05) in nucleotide substitutions between pre-chemotherapy and post-chemotherapy tumors. (b) Proportion of SNVs occurring in specific nucleotide motif contexts for each category of single-nucleotide substitution. Blue, pre-chemotherapy tumors; green, post-chemotherapy tumors. (c) Four mutational signatures identified in urothelial carcinoma. (d) Significant enrichment of APOBEC signatures in post-chemotherapy tumors. (e) Significant increases in the clonality of APOBEC-induced mutations in post-chemotherapy urothelial carcinoma. Numbers of SNVs, exact P values and total numbers of samples are reported in the Supplementary Note.
To distinguish between potential mutagenic mechanisms responsible for these changes, we matched mutational patterns derived from statistical analysis of nucleotide changes to well-defined signatures of potential mutagens. We observed a significant increase in C>A nucleotide substitutions in tumors treated with cisplatin-based chemotherapy that is consistent with the specific mutagenesis signature induced in Caenorhabditis elegans genome after cisplatin treatment25,26. Further analysis of the context motifs for various base substitutions showed enrichment of C>T or C>G changes at TCW motifs, where W is A or T (Fig. 7b), which is highly suggestive of APOBEC-induced mutagenesis27,28. To confirm this finding, we compared the signatures in our cohort to previously reported Sanger signatures27,28,29 (Online Methods). We observed four distinct signatures in our cohort (Fig. 7c). The first signature was very similar to Sanger signatures 2 and 13, attributed to APOBEC mutagenesis27,28,29,30. We detected three additional signatures corresponding to previously described mutagenic processes associated with age, smoking and ERCC2 mutations27,29. The low frequency of ERCC2 mutations in our cohort of chemotherapy-treated urothelial carcinoma (Supplementary Fig. 1) is consistent with previous reports suggesting that ERCC2 mutations are enriched in responders to cisplatin-based chemotherapy31,32 and are likely to be selected against in tumors that progress through chemotherapy.
Because of the prominence of APOBEC-induced mutagenesis in urothelial carcinoma, we focused on understanding how APOBEC-induced mutations evolve during chemotherapy by comparing the frequency of APOBEC-induced mutations in chemotherapy-naive and chemotherapy-treated tumors. We observed a significant enrichment in APOBEC3-induced mutagenesis (C>T or C>G changes at TCW motifs) in post-chemotherapy tumors (Fig. 7d). To dissect the relative contributions of individual members of the APOBEC3 cytosine deaminase family to this enrichment, we examined the motif contexts favored by individual APOBEC enzymes in mutating respective cytosines33,34. APOBEC3A favors YTCA motifs, while APOBEC3B favors RTCA motifs (wherein Y = pyrimidine and R = purine)33. APOBEC3G induces cytosine substitutions in the single-stranded DNA overhang strand, with a preference for the CCC motif34,35, whereas APOBEC3F preferentially mutates cytosines in the TTC motif (where the underlined C is the mutated nucleotide)36. We detected a significant enrichment in APOBEC3A-induced mutations (P = 0.00001, Fisher's exact test) and a similar enrichment in APOBEC3B mutagenesis (P = 0.0395, Fisher's exact test) in post-chemotherapy tumors. In contrast, APOBEC3G mutagenesis was substantially decreased in post-chemotherapy tumors (Fig. 7d). Furthermore, we observed a corresponding statistically significant increase in the clonality of APOBEC-induced mutations in post-chemotherapy tumors (Fig. 7e). Enrichment analysis of APOBEC-induced mutations highlighted key pathways involved in chemotherapy resistance, including the ABC family of proteins (odds ratio = 2.7; P = 0.038, Fisher's exact test) and homologous recombination DNA-damage repair (odds ratio = 3.8; P = 0.033, Fisher's exact test) (Supplementary Table 5). Our findings suggest that the APOBEC mutational process is not merely a transient event in early urothelial carcinoma oncogenesis but rather continues to shape the evolution of advanced urothelial carcinoma and may promote clonal expansions of chemotherapy-resistant clones.
Discussion
Advanced, chemotherapy-resistant urothelial carcinoma remains a formidable clinical challenge with limited therapeutic options37. Whole-exome analysis of matched samples from the same patient obtained from different anatomical sites and at sequential time points offers a unique opportunity to reconstruct evolutionary dynamics and understand the mutagenic pressures shaping the evolution of primary, untreated urothelial carcinoma to advanced, chemotherapy-treated urothelial carcinoma.
Our analyses identified substantial spatial and temporal heterogeneity between tumors separated in time or by anatomical location within the same patient. The majority of mutations in the post-chemotherapy tumors were not shared with primary, chemotherapy-naive tumors. Branching evolution was the predominant path from primary, chemotherapy-naive to advanced, chemotherapy-treated urothelial carcinoma. Very early in this path, several clonal waves separate from the original founder clone, many of which metastasize early in the tumor's lifetime and continue to evolve in parallel with the primary tumor. Our findings shed light on the importance of addressing gaps in the existing knowledge of the clonality of early events in urothelial carcinoma oncogenesis, including multicentricity and malignant seeding, which could lead to alternative interpretations of the phylogenetic trees. Additionally, our findings suggest that extensive heterogeneity and early branching evolution should be taken into consideration as additional layers of biological complexity that go beyond the traditional two-pathway model of urothelial carcinoma oncogenesis and potentially eclipse grade and stage classifications38.
We demonstrate that chemotherapy-treated urothelial carcinoma is significantly clonally enriched in mutations in L1CAM and integrin signaling pathways. In our study, the majority of these mutations were missense mutations that could potentially lead to activation of these pathways, but the precise functional impact of these mutations warrants future studies. Our results are consistent with data in preclinical models of other tumor types such as cholangiocarcinoma, ovarian carcinoma and pancreatic ductal adenocarcinoma, demonstrating that L1CAM has a key role in cisplatin resistance and in protecting cells from apoptosis39,40,41,42. L1CAM directly binds to integrin receptors via its RGD motif in the sixth immunoglobulin domain43,44, and there is considerable cross-talk between the two pathways45,46. Previous studies demonstrated that integrin signaling has an important role in overriding chemotherapy-induced cell cycle arrest and apoptosis in small-cell lung cancer cells through activation of the phosphatidylinositol-3 kinase (PI3K) pathway47,48. Overexpression of β1-integrin in hepatocellular carcinoma cell lines protected them against apoptosis induced by chemotherapeutic agents by activating MAP kinase signaling49. Stimulating β1-integrin with an antibody ligand in leukemia cells prevented procaspase-8–mediated induction of apoptosis in a PI3K-dependent manner50. Collectively, these observations suggest that alterations in L1CAM and integrin signaling pathways potentially play a key role in chemotherapy resistance in urothelial carcinoma, and provide a mechanistic intersection between the tumor microenvironment and drug resistance through the phenomenon of cell-adhesion-mediated drug resistance (CAM-DR)20,51,52. This phenomenon has been implicated in chemotherapy resistance in several malignancies53,54. L1CAM is potentially targetable with antibodies that have demonstrated efficacy in xenograft animal models of cholangiocarcinomas55,56 and ovarian and pancreatic ductal carcinomas57. Focal adhesion kinase (FAK) inhibitors, which target integrin signaling, have been shown to profoundly sensitize cancer cells to chemotherapy and novel molecular therapeutics, and are currently in early-phase clinical trials58,59. Our results suggest that similar therapeutic approaches merit further study in chemotherapy-resistant urothelial carcinoma.
We demonstrate an increase in APOBEC signatures in chemotherapy-treated tumors. One possible explanation for this interesting finding is that platinum-based chemotherapy increases the formation of APOBEC-mutagenesis-prone single-stranded DNA (ssDNA)60,61. This ssDNA is formed during the 5′→3′ resection that occurs at DNA double-strand breaks during homology-directed repair62. While allowing for error-free repair of these double-strand breaks induced by the excision of platinum–DNA adducts, this process may potentially increase the availability of intermediary ssDNA substrate to APOBEC mutagenesis62. Our results also suggest that APOBEC3A is the main enzyme responsible for mutagenesis in advanced, chemotherapy-treated urothelial carcinoma. This is in accordance with recent data suggesting that APOBEC3A is the key mutagenic cytidine deaminase in most tumor types because of its high proficiency in generating DNA breaks63. These findings suggest a potential mechanism by which chemotherapy acts to increase the genomic diversity of chemotherapy-treated tumors that requires future study. Our data demonstrate that the clonal evolution of chemotherapy-treated urothelial carcinoma is characterized by a dramatic divergence of the mutational landscape in the face of relative stability at the copy number level over the lifetime of each tumor. This finding potentially reflects the dominance of APOBEC-induced mutagenesis in urothelial carcinoma as a mechanism that preferentially induces single-nucleotide changes throughout the tumors' lifetimes and during chemotherapy. An alternative explanation for the marked genomic alterations we observed in chemotherapy-treated samples is that a certain degree of genetic drift occurs over time irrespective of the effect of chemotherapy. However, it is unlikely that genetic drift is the sole mechanism accounting for the genetic heterogeneity we observed, as chemotherapy is a potent selective pressure that is expected to alter evolutionary dynamics, affecting the pace and steering the direction of genetic drift. In fact, our results are consistent with evolutionary models suggesting that cancer's adaptation ensues from the interaction between stochastic processes, such as mutation generation, and clonal selection, which is a deterministic phenomenon64. Our findings support this evolutionary model by demonstrating a complex dynamic interplay between mutagenic mechanisms, such as APOBEC-induced mutagenesis, and extrinsic selective pressures, such as chemotherapy, to constantly shape the clonal evolution of urothelial carcinoma. As genetic information is passed from parent to progeny clones, each process leaves an evolutionary record of molecular alterations in the descendent clones that allows reconstruction of the process. However, it is important to note that this record fully exists only in clones that survive selection. Unfit clones are eliminated from the record and can only be captured by serial sampling of cancer cells throughout the tumor's lifetime, whereas resistant clones are selected to expand and supersede previous clonal waves.
One strength of our study is that it is the most comprehensive investigation (to our knowledge) of the clonal evolution of chemotherapy-resistant urothelial carcinoma. Limitations of our study include a small sample size. Of note, we included muscle-invasive tumor samples from patients who were never treated with chemotherapy as controls.
Our findings have several potential clinical implications. First, genomic divergence between untreated and treated clones suggests that clinically actionable molecular targets in metastatic chemotherapy-treated tumors may be missed when relying only on biopsies of untreated primary tumors at the time of diagnosis. Serial metastatic biopsies during the course of clinical care would be needed to detect the most recent version of the rapidly changing molecular landscape of a given patient's urothelial carcinoma. Second, further study of the functional role of L1CAM and integrin signaling in mediating chemotherapy resistance in urothelial carcinoma could lead to a potential strategy for reversing or preventing chemotherapy resistance by targeting these pathways. Third, despite its initial effectiveness in eliminating cancer cells, platinum-based chemotherapy is associated with unintended significant mutagenic editing of the genomic landscape of post-chemotherapy tumors. Our insight into the nature of these edits is a crucial step toward a complete understanding of the basis of chemotherapy resistance in advanced urothelial carcinoma, which will lay the foundation for the development of rational therapeutic strategies for preventing the emergence of the chemotherapy-resistant state of urothelial carcinoma or for reversing it.
In summary, our results demonstrate that advanced, chemotherapy-treated urothelial carcinoma undergoes extensive and dynamic clonal evolution throughout the lifetime of the tumor, with significant genetic editing that continues during and after chemotherapy. Our findings lay the foundation for an evolutionary understanding of advanced, chemotherapy-treated urothelial carcinoma and present opportunities for advancing cancer precision medicine.
Methods
Patient enrollment and tumor procurement.
All experimental procedures were carried out in accordance with approved guidelines and were approved by the Institutional Review Board (IRB) at Weill Cornell Medicine. Patients in the study signed informed consent under an IRB-approved protocol (IRB #1305013903). Clinical information was collected from the chart. Smoking status was collected from self-administered questionnaires. Tumor samples were obtained from patients through surgical resection or core biopsies.
Rapid autopsy procedures.
The Englander Institute for Precision Medicine at Weill Cornell Medicine, New York Presbyterian has been established to promote personalized medicine focused on molecular diagnostics and therapeutics. Two patients in our series selected the option to be enrolled in the IRB-approved rapid autopsy program. In addition, patients' next-of-kin provided written consent before autopsy. The WCM117 and WCM259 rapid autopsies were conducted within 6 h after death. A systematic autopsy protocol is followed where normal and malignant fresh tissue is collected, allocating samples to be snap frozen or formalin-fixed. The goal is to maximize the amount of tissue collected for research purposes. Once the tissue harvest is complete, the autopsy proceeds in accordance with the protocol established by the WCM Autopsy Service. For our current study, tissue samples from multiple sites were procured from each patient as detailed above. After hematoxylin and eosin (H&E) evaluation and frozen slide annotation, DNA was extracted for whole-exome sequencing (WES).
DNA extraction and next generation sequencing.
In this study, we used a New York State approved whole-exome sequencing assay developed in our Clinical Laboratory Improvement Amendments (CLIA) laboratory called EXaCT-1 (ref. 65). After macrodissection of target lesions, tumor DNA was extracted from formalin-fixed, paraffin-embedded (FFPE) or cored OCT-cryopreserved tumors using the Promega Maxwell 16 MDx (Promega, Madison, WI, USA). Germline DNA was extracted from peripheral blood mononuclear cells using the same method. Pathological review by one of the study pathologists (J.M.M., B.R., M.A.R.) confirmed the diagnosis and determined tumor content. A minimum of 200 ng of DNA was used for whole-exome sequencing. DNA quality was determined by TapeStation Instrument (Agilent Technologies, Santa Clara, CA) and was confirmed by real-time PCR before sequencing. Sequencing was performed using Illumina HiSeq 2500 (2×100 bp). A total of 21,522 genes were analyzed with an average coverage of 85× (range 60–102; Supplementary Table 6) using Agilent HaloPlex Exome (Agilent Technologies, Santa Clara, CA). We developed a targeted sequencing assay of 250 cancer genes (referred to as N250) using hybrid-capture SeqCap EZ Choice Enrichment Kits (Roche Sequencing, Pleasanton, CA) (Supplementary Tables 7 and 8). Sequencing was performed using Illumina HiSeq 2500 (PE 2×75) achieving an average coverage of 400×.
Study sample size definition.
Our study includes 72 samples from 32 patients. However, patient WCM117 comprises 12 samples (17% of the total), and to avoid possible statistical biases in the analysis, we used WCM117 (12 samples) only in Figure 4, as a specific case study to better understand how chemotherapy shapes evolution. Finer analyses such as that of allele specific copy number (Fig. 5a) and SNVs clonality (Fig. 6a) also require an estimate of ploidy and purity. After manual inspection of CLONET outputs, we had 44 samples in 25 patients with reliable ploidy and purity estimates to allow for such analyses. The exact number of samples and patients used in each figure is reported in the Supplementary Note.
Sequence data processing pipeline.
All the study samples data were processed through the computational analysis pipeline of the Institute of Precision Medicine at Weill Cornell, New York Presbyterian Hospital (IPM-Exome-pipeline)65. Raw reads quality was assessed with FASTQC as described previously65. Multi-sample patient data were tested for genotype distance using SPIA66. Pipeline output includes segment DNA copy number data, somatic copy-number aberrations (SCNAs) (Supplementary Table 9), and putative somatic single nucleotide variants (SNVs) (Supplementary Table 10). Finally, to assess tumor ploidy and purity, we applied CLONET9 on segmented data and allelic fraction (AF) of germline heterozygous SNP loci (termed 'informative SNPs'; see “Allele-specific copy number analysis” below). Upon visual inspection of CLONET output, data from 53 of 72 samples were deemed appropriate for downstream copy number analysis; excluded samples were not associated with chemotherapy treatment (P = 0.4384) or biopsy site (P = 1).
We have previously published research on how EXaCT-1 was developed and optimized for use with FFPE and frozen samples, in contrast to many research-grade assays65. To further ensure that our results reflect biological effects rather than technical variability between different sample types, we took special care to account for variations in tumor content for each sample in order to correctly map the clonal evolution of urothelial carcinoma. In particular, we did not observe any significant difference (P = 0.1, Wilcoxon test) when comparing FFPE and fresh sample purity (Supplementary Fig. 9a). Similarly, we did not detect differences (P = 0.137, Wilcoxon test) in the numbers of identified non-silent SNVs between FFPE and fresh samples (Supplementary Fig. 9b).
Allele-specific copy number analysis.
Somatic copy number altered regions are defined by the log2 of the ratio between the tumor and normal local coverage normalized by the global tumor and normal coverage ratio (named log2R). CLONET refines copy number data, adjusting each log2R to account for both aneuploidy and tumor purity. Combining purified log2R values and allele frequency (AF) of informative SNPs, CLONET assigns allele-specific copy number values, represented as a pair (cnA, cnB), to each genomic segment67. Quality filters require at least 10 informative SNPs and a mean coverage of 20 reads to call allele-specific values of a segment. If a segment does not pass filters, adjusted log2R values below −0.4 (above 0.4) were categorized as copy number loss (gain).
Differential copy number analysis between pre- and post-chemotherapy has been performed on a set of 1,160 genes selected among putative cancer genes (COSMIC68 and Intogen69) (CancerGenes) and bladder-specific genes (Supplementary Table 11).
Unsupervised clustering analysis was performed on allele-specific copy number of CancerGenes by means of hierarchical clustering. Briefly, the allele-specific copy number of a gene is represented as a pair of real values. Then, the Euclidean distance on allele-specific copy number calls is used as distance; this approach distinguishes between ambiguous cases such as copy number wild-type status (allele-specific values (1,1)) and copy number neutral loss (2,0), both corresponding to log2R = 0, leading to more informative clusters (Supplementary Table 4). Copy-number-based analysis identified two clusters that we named WCM_A and WCM_B (Fig. 5; Supplementary Fig. 6), which we compared to the original four TCGA clusters (I, II, III, IV) that resulted from an integrated analysis of mRNA, miRNA and protein data8 (Supplementary Fig. 7). Given a TCGA cluster X, we tested the null hypothesis that clusters WCM_A and WCM_B contain the same proportion of samples from X using Fisher's exact test (significance level = 0.05).
Single nucleotide variations analysis.
To improve the quality of SNVs calls in targeted exons, we applied an integrated approach. We first ran both MuTect70 and SNVseeqer71 to nominate putative aberrant genomic positions. Then, we looked closely at the identified positions by means of ASEQ72 in normal and germline samples, executing pileup analysis; for each single nucleotide position identified as putative aberrant, ASEQ returns information about the read count for each of the four bases, A, C, G, and T. To reduce false positives, we required base quality and read quality above 20. Finally, a genomic position is considered aberrant in a tumor sample if the read count of the alternative base is zero in the matched germline and three or more in the tumor. This step allows us to (i) filter out remaining germline SNPs, that is, positions where the alternative base is present in the control sample; and (ii) check for the presence of SNVs with low AFs in patient's multiple samples data. This step is particularly relevant in this study, because SNV identification methods are designed to work on a single normal tumor pair, and they do not consider that samples from the same patient could share the same SNVs, as they are not independent samples. Finally, we annotated genomic position with information relevant for cancer analysis with Oncotator73 (v1.8.0.0). We exploited the last Oncotator datasource corpus including annotations about gene and transcript names, functional consequence (for example, missense or nonsense), the predicted impact on protein function, annotation from cancer-specific resources such as COSMIC or TCGAscape, and possible published results about the specific mutation (Supplementary Table 10). A full description of the resources used by Oncotator is available in the tool help page. We identified 13 possible functional consequences (Supplementary Fig. 10) described in the Sequence Ontology74. To avoid overestimating divergence among samples from the same patient, we were conservative in defining SNVs that more likely produce a change in the protein, as in those that affect the phenotype, and we only considered non-silent mutations as those classified as missense, nonsense, splice-site, nonstop, and start codon. However, for the analysis described in Figure 7, we included all the identified mutations (Supplementary Fig. 10), as mutational mechanisms also affect mutations with neutral functional effects. To further confirm our findings and to check for possible biases introduced by the definition of silent mutations, we also performed the analysis of Figures 2, 3, 4, and 6 using both silent and non-silent SNVs, obtaining comparable results (Supplementary Figs. 11,12,13 and 14).
SNV enrichment analysis.
Gene set enrichment analysis (GSEA) refers to a computational method that identifies sets of genes that are statistically enriched for a given observable variable. In this study, we interrogated the REACTOME pathway database to search for pathways showing a significant increase in the number of SNVs. We applied Fisher's exact test followed by FDR correction by Benjamin-Hochberg (BH) procedure. REACTOME pathways with FDR ≤ 0.2 are reported in Figure 6b. We also checked if, among significant pathways, there is a statistically significant difference between pre- and post- chemotherapy samples. Given a pathway P, Fisher's exact test determines the probability that the number of SNVs in P is different when considering pre- and post- chemo samples. After FDR correction (BH procedure), we highlighted nodes in Figure 6b with FDR ≤ 0.2.
Phylogenetic analysis.
High-quality variants identified in the previous steps were used to reconstruct the phylogenetic tree of each patient using the parsimony ratchet method10 (Figs. 3, 4e and Supplementary Fig. 5). In this representation, each node models a population of tumor cells. Nodes with no children, named leafs, represent cell populations from a tumor sampling, i.e., a tumor biopsy. Internal nodes model inferred tumor cell populations from observed SNVs. The node named WT represents a hypothetical population of wild-type cells (cells with no somatic aberrations). In phylogenetic trees, an edge connects two nodes; the length of an edge is proportional to the number of SNVs. For instance, in Figure 4e, node 1 corresponds to the least common ancestor inferred from all the available biopsies, with the number of mutations proportional to the length of the edge from WT to 1. Node 1 is also connected to the pelvic lymph node metastasis and to the inferred cell population 2. A branch represents a time point in the evolution of the tumor where two distinct cell populations emerge; the length of the branches models the number of SNVs that are private to each population. In Figure 4e, we observed few private mutations in the pelvic lymph node metastasis with respect to the number of SNVs shared by all the other samples, as supported by the heat map in Figure 4d.
Clonality of single nucleotide variants.
Original CLONET implementation allows for the computation of the clonality of an SNV with copy number normal genomic segment, a segment that has an allele-specific copy number (1, 1). Here we extended it to allow for SNV clonality estimation independently from the copy number status of the genomic segments in which the SNV lies. Given the tumor purity P, the allele specific copy number (cnA, cnB), and the number of reads nRef and nAlt supporting the reference and the alternative base, respectively, the clonality of a SNV required to compute expected AF was estimated. We observed that AF could assume only a finite number of values given the DNA copy number state. For instance, let's assume that a locus is aberrant (mutated) in one allele and wild-type (not mutated) in two alleles, such as SNV in a copy number aberrant segment (CN = 3). In this case, with tumor cellularity equal to 100% and clonal SNV, the VAF would be equal to 1/3. Given the number of allele cnSNV harboring a SNV, the expected VAF is defined as:

To estimate cnSNV, we followed parsimony approach assuming cnSNV that best explains VAF adjusted by DNA admixture, computed as previously described9. We computed the clonality of SNVs using the distance between the observed and expected VAF as:

Complete proof is in the Supplementary Note.
Single nucleotide variant signatures.
SNVs are partitioned into six mutation classes (column “Mutation class”, Supplementary Table 10) corresponding to six types of base pair substitution, C>A, C>G, C>T, T >A, T>C, T>G. The null hypothesis that pre- and post-chemotherapy samples are equally likely to harbor SNVs of a specific mutation class is tested with Fisher's exact test (Fig. 7a). The fingerprint of a SNV includes the two bases immediately 5′ and 3′ to each SNV position (column “Genomic context”, Supplementary Table 10) for a total of 96 possible mutation fingerprints27. Fisher's exact test adjusted for multiple hypotheses testing with Benjamin-Hochberg procedure returns the likelihood that a mutation fingerprint is enriched in pre- or post-chemotherapy samples (Fig. 7b). As the set of mutation fingerprints of a tumor sample is a proxy for the mutational processes that shape the cancer genome, we studied the mutational signatures of our study samples and compared them the with the Sanger signatures27, applying the same approach recently proposed29. Briefly, the Sanger signatures were obtained from the identification of 30 mutational processes signatures (named Sanger signatures) upon application of the original tool on more than 10,000 samples from 40 distinct human cancer types27. In our data set, we identify four mutational signatures (Fig. 7c); the Sanger signature analysis shows that APOBEC proteins play a role in the mutational processes shaping urothelial carcinoma genomes. We checked for statistically significant differences between pre- and post-chemotherapy of individual members of the APOBEC family using Fisher's exact test followed by Benjamin-Hochberg FDR correction (Fig. 7d). To test if the clonality of APOBEC-induced SNVs is enriched in post-chemotherapy samples, we dichotomized the SNV clonality levels (threshold 0.6) and then we applied Fisher's exact test (Fig. 7e).
Fluorescence in situ hybridization.
Two 4-μm-thick tissue sections from each block were cut for FISH analysis. CDKN2A deletion was determined using FISH probe (BAC clone RP11-149I2) and a reference probe, located at 9p21. At least 100 nuclei were evaluated per sample using a fluorescence microscope (Olympus BX51; Olympus Optical).
Statistical analysis.
For statistical tests, two-sided Mann–Whitney–Wilcoxon test (referred to as Wilcoxon test in the main text) was used to check for significant differences between two distributions. The two-sided Fisher's exact test was applied to determine whether the deviations between the observed and the expected counts were significant. When appropriate, P values were adjusted for multiple hypotheses testing with Benjamin-Hochberg procedure. Boxplot statistics were computed with the function “boxplot” of R programming language. No statistical methods were used to predetermine study sample size.
URLs.
CLONET, https://bitbucket.org/deid00/clonet/; Oncotator, https://www.broadinstitute.org/oncotator/; Oncotator datasource corpus, http://www.broadinstitute.org/~lichtens/oncobeta/oncotator_v1_ds_Jan262015.tar.gz; REACTOME pathway database, http://www.reactome.org/; Sanger signatures, http://cancer.sanger.ac.uk/cosmic/signatures.
Accession codes.
All BAM files and associated sample information are deposited in dbGaP under accession phs001087.v1.p1.
References
Howlader, N. et al. SEER Cancer Statistics Review, 1975–2010 (National Cancer Institute, Bethesda, Maryland, USA, 2013).
von der Maase, H. et al. Long-term survival results of a randomized trial comparing gemcitabine plus cisplatin, with methotrexate, vinblastine, doxorubicin, plus cisplatin in patients with bladder cancer. J. Clin. Oncol. 23, 4602–4608 (2005).
Clark, P.E. et al. Bladder cancer. J. Natl. Compr. Canc. Netw. 11, 446–475 (2013).
Sternberg, C.N. et al. Randomized phase III trial of high-dose-intensity methotrexate, vinblastine, doxorubicin, and cisplatin (MVAC) chemotherapy and recombinant human granulocyte colony-stimulating factor versus classic MVAC in advanced urothelial tract tumors: European Organization for Research and Treatment of Cancer Protocol no. 30924. J. Clin. Oncol. 19, 2638–2646 (2001).
Sternberg, C.N. et al. Seven year update of an EORTC phase III trial of high-dose intensity M-VAC chemotherapy and G-CSF versus classic M-VAC in advanced urothelial tract tumours. Eur. J. Cancer 42, 50–54 (2006).
Bellmunt, J. et al. Phase III trial of vinflunine plus best supportive care compared with best supportive care alone after a platinum-containing regimen in patients with advanced transitional cell carcinoma of the urothelial tract. J. Clin. Oncol. 27, 4454–4461 (2009).
Bellmunt, J. & Petrylak, D.P. New therapeutic challenges in advanced bladder cancer. Semin. Oncol. 39, 598–607 (2012).
Cancer Genome Atlas Research Network. Comprehensive molecular characterization of urothelial bladder carcinoma. Nature 507, 315–322 (2014).
Prandi, D. et al. Unraveling the clonal hierarchy of somatic genomic aberrations. Genome Biol. 15, 439 (2014).
Nixon, K.C. The parsimony ratchet, a new method for rapid parsimony analysis. Cladistics 15, 407–414 (1999).
Zheng, Y. et al. Identification of Pold2 as a novel interaction partner of protein inhibitor of activated STAT2. Int. J. Mol. Med. 30, 884–888 (2012).
Lee, Y.S., Gregory, M.T. & Yang, W. Human Pol ζ purified with accessory subunits is active in translesion DNA synthesis and complements Pol η in cisplatin bypass. Proc. Natl. Acad. Sci. USA 111, 2954–2959 (2014).
van Boxtel, R. et al. FOXP1 acts through a negative feedback loop to suppress FOXO-induced apoptosis. Cell Death Differ. 20, 1219–1229 (2013).
Choi, E.J. et al. FOXP1 functions as an oncogene in promoting cancer stem cell-like characteristics in ovarian cancer cells. Oncotarget 7, 3506–3519 (2016).
Greco, C. et al. E-cadherin/p120-catenin and tetraspanin Co-029 cooperate for cell motility control in human colon carcinoma. Cancer Res. 70, 7674–7683 (2010).
Wei, L., Li, Y. & Suo, Z. TSPAN8 promotes gastric cancer growth and metastasis via ERK MAPK pathway. Int. J. Clin. Exp. Med. 8, 8599–8607 (2015).
Zöller, M. Tetraspanins: push and pull in suppressing and promoting metastasis. Nat. Rev. Cancer 9, 40–55 (2009).
Kinsella, R.J. et al. Ensembl BioMarts: a hub for data retrieval across taxonomic space. Database (Oxford) 2011, bar030 (2011).
Plimack, E.R. et al. Defects in DNA repair genes predict response to neoadjuvant cisplatin-based chemotherapy in muscle-invasive bladder cancer. Eur. Urol. 68, 959–967 (2015).
Eke, I. & Cordes, N. Focal adhesion signaling and therapy resistance in cancer. Semin. Cancer Biol. 31, 65–75 (2015).
Maiuthed, A. & Chanvorachote, P. Cisplatin at sub-toxic levels mediates integrin switch in lung cancer cells. Anticancer Res. 34, 7111–7117 (2014).
Seguin, L., Desgrosellier, J.S., Weis, S.M. & Cheresh, D.A. Integrins and cancer: regulators of cancer stemness, metastasis, and drug resistance. Trends Cell Biol. 25, 234–240 (2015).
Tsutsumi, S. et al. L1 cell adhesion molecule (L1CAM) expression at the cancer invasive front is a novel prognostic marker of pancreatic ductal adenocarcinoma. J. Surg. Oncol. 103, 669–673 (2011).
Weidle, U.H., Eggle, D. & Klostermann, S. L1-CAM as a target for treatment of cancer with monoclonal antibodies. Anticancer Res. 29, 4919–4931 (2009).
Meier, B. et al. C. elegans whole-genome sequencing reveals mutational signatures related to carcinogens and DNA repair deficiency. Genome Res. 24, 1624–1636 (2014).
Szikriszt, B. et al. A comprehensive survey of the mutagenic impact of common cancer cytotoxics. Genome Biol. 17, 99 (2016).
Alexandrov, L.B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Nik-Zainal, S. et al. Mutational processes molding the genomes of 21 breast cancers. Cell 149, 979–993 (2012).
Kim, J. et al. Somatic ERCC2 mutations are associated with a distinct genomic signature in urothelial tumors. Nat. Genet. 48, 600–606 (2016).
Roberts, S.A. et al. An APOBEC cytidine deaminase mutagenesis pattern is widespread in human cancers. Nat. Genet. 45, 970–976 (2013).
Liu, D. et al. Clinical validation of chemotherapy response biomarker ERCC2 in muscle-invasive urothelial bladder carcinoma. JAMA Oncol. 2, 1094–1096 (2016).
Van Allen, E.M. et al. Somatic ERCC2 mutations correlate with cisplatin sensitivity in muscle-invasive urothelial carcinoma. Cancer Discov. 4, 1140–1153 (2014).
Hoopes, J.I. et al. APOBEC3A and APOBEC3B preferentially deaminate the lagging strand template during DNA replication. Cell Rep. 14, 1273–1282 (2016).
Lackey, L. et al. APOBEC3B and AID have similar nuclear import mechanisms. J. Mol. Biol. 419, 301–314 (2012).
Chelico, L., Pham, P. & Goodman, M.F. Stochastic properties of processive cytidine DNA deaminases AID and APOBEC3G. Phil. Trans. R. Soc. Lond. B 364, 583–593 (2009).
Harris, R.S. & Liddament, M.T. Retroviral restriction by APOBEC proteins. Nat. Rev. Immunol. 4, 868–877 (2004).
Faltas, B.M., Karir, B.S., Tagawa, S.T. & Rosenberg, J.E. Novel molecular targets for urothelial carcinoma. Expert Opin. Ther. Targets 19, 515–525 (2015).
Knowles, M.A. & Hurst, C.D. Molecular biology of bladder cancer: new insights into pathogenesis and clinical diversity. Nat. Rev. Cancer 15, 25–41 (2015).
Yoon, H. et al. L1 cell adhesion molecule and epidermal growth factor receptor activation confer cisplatin resistance in intrahepatic cholangiocarcinoma cells. Cancer Lett. 316, 70–76 (2012).
Stoeck, A. et al. L1-CAM in a membrane-bound or soluble form augments protection from apoptosis in ovarian carcinoma cells. Gynecol. Oncol. 104, 461–469 (2007).
Sebens Müerköster, S. et al. Drug-induced expression of the cellular adhesion molecule L1CAM confers anti-apoptotic protection and chemoresistance in pancreatic ductal adenocarcinoma cells. Oncogene 26, 2759–2768 (2007).
Sebens Müerköster, S. et al. alpha5-integrin is crucial for L1CAM-mediated chemoresistance in pancreatic adenocarcinoma. Int. J. Oncol. 34, 243–253 (2009).
Kiefel, H. et al. L1CAM: a major driver for tumor cell invasion and motility. Cell Adh. Migr. 6, 374–384 (2012).
Gast, D. et al. The RGD integrin binding site in human L1-CAM is important for nuclear signaling. Exp. Cell Res. 314, 2411–2418 (2008).
Voura, E.B., Ramjeesingh, R.A., Montgomery, A.M. & Siu, C.H. Involvement of integrin alpha(v)beta(3) and cell adhesion molecule L1 in transendothelial migration of melanoma cells. Mol. Biol. Cell 12, 2699–2710 (2001).
Burgett, M.E. et al. Direct contact with perivascular tumor cells enhances integrin αvβ3 signaling and migration of endothelial cells. Oncotarget http://dx.doi.org/10.18632/oncotarget.9700 (2016).
Hodkinson, P.S. et al. ECM overrides DNA damage-induced cell cycle arrest and apoptosis in small-cell lung cancer cells through beta1 integrin-dependent activation of PI3-kinase. Cell Death Differ. 13, 1776–1788 (2006).
Sethi, T. et al. Extracellular matrix proteins protect small cell lung cancer cells against apoptosis: a mechanism for small cell lung cancer growth and drug resistance in vivo. Nat. Med. 5, 662–668 (1999).
Zhang, H. et al. Beta 1-integrin protects hepatoma cells from chemotherapy induced apoptosis via a mitogen-activated protein kinase dependent pathway. Cancer 95, 896–906 (2002).
Argyris, E.G. et al. The interferon-induced expression of APOBEC3G in human blood-brain barrier exerts a potent intrinsic immunity to block HIV-1 entry to central nervous system. Virology 367, 440–451 (2007).
Shain, K.H. & Dalton, W.S. Cell adhesion is a key determinant in de novo multidrug resistance (MDR): new targets for the prevention of acquired MDR. Mol. Cancer Ther. 1, 69–78 (2001).
Damiano, J.S. Integrins as novel drug targets for overcoming innate drug resistance. Curr. Cancer Drug Targets 2, 37–43 (2002).
Hazlehurst, L.A. & Dalton, W.S. Mechanisms associated with cell adhesion mediated drug resistance (CAM-DR) in hematopoietic malignancies. Cancer Metastasis Rev. 20, 43–50 (2001).
Hehlgans, S., Haase, M. & Cordes, N. Signalling via integrins: implications for cell survival and anticancer strategies. Biochim. Biophys. Acta 1775, 163–180 (2007).
Cho, S. et al. Generation, characterization and preclinical studies of a human anti-L1CAM monoclonal antibody that cross-reacts with rodent L1CAM. MAbs 8, 414–425 (2016).
Min, J.K. et al. L1 cell adhesion molecule is a novel therapeutic target in intrahepatic cholangiocarcinoma. Clin. Cancer Res. 16, 3571–3580 (2010).
Schäfer, H. et al. Combined treatment of L1CAM antibodies and cytostatic drugs improve the therapeutic response of pancreatic and ovarian carcinoma. Cancer Lett. 319, 66–82 (2012).
Golubovskaya, V.M. Targeting FAK in human cancer: from finding to first clinical trials. Front. Biosci. (Landmark Ed.) 19, 687–706 (2014).
Schultze, A. & Fiedler, W. Therapeutic potential and limitations of new FAK inhibitors in the treatment of cancer. Expert Opin. Investig. Drugs 19, 777–788 (2010).
Rezaee, M., Sanche, L. & Hunting, D.J. Cisplatin enhances the formation of DNA single- and double-strand breaks by hydrated electrons and hydroxyl radicals. Radiat. Res. 179, 323–331 (2013).
Zellweger, R. et al. Rad51-mediated replication fork reversal is a global response to genotoxic treatments in human cells. J. Cell Biol. 208, 563–579 (2015).
Roberts, S.A. & Gordenin, D.A. Hypermutation in human cancer genomes: footprints and mechanisms. Nat. Rev. Cancer 14, 786–800 (2014).
Chan, K. et al. An APOBEC3A hypermutation signature is distinguishable from the signature of background mutagenesis by APOBEC3B in human cancers. Nat. Genet. 47, 1067–1072 (2015).
Lipinski, K.A. et al. Cancer evolution and the limits of predictability in precision cancer medicine. Trends Cancer 2, 49–63 (2016).
Beltran, H. et al. Whole-exome sequencing of metastatic cancer and iomarkers of treatment response. JAMA Oncol. 1, 466–474 (2015).
Demichelis, F. et al. SNP panel identification assay (SPIA): a genetic-based assay for the identification of cell lines. Nucleic Acids Res. 36, 2446–2456 (2008).
Beltran, H. et al. Divergent clonal evolution of castration-resistant neuroendocrine prostate cancer. Nat. Med. 22, 298–305 (2016).
Futreal, P.A. et al. A census of human cancer genes. Nat. Rev. Cancer 4, 177–183 (2004).
Rubio-Perez, C. et al. In silico prescription of anticancer drugs to cohorts of 28 tumor types reveals targeting opportunities. Cancer Cell 27, 382–396 (2015).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Jiang, Y., Soong, T.D., Wang, L., Melnick, A.M. & Elemento, O. Genome-wide detection of genes targeted by non-Ig somatic hypermutation in lymphoma. PLoS One 7, e40332 (2012).
Romanel, A., Lago, S., Prandi, D., Sboner, A. & Demichelis, F. ASEQ: fast allele-specific studies from next-generation sequencing data. BMC Med. Genomics 8, 9 (2015).
Ramos, A.H. et al. Oncotator: cancer variant annotation tool. Hum. Mutat. 36, E2423–E2429 (2015).
Eilbeck, K. et al. The sequence ontology: a tool for the unification of genome annotations. Genome Biol. 6, R44 (2005).
Acknowledgements
We would like to thank our patients and their families for participation in this study. We would like to thank B. Sleckman for constructive review of the manuscript. We would also like to acknowledge D. Scherr and C. Barbieri for contributing samples, our research and clinical pathology fellows J. Fontugne, M. Kossai, C. Pauli, K. Hennrick and K. Park for their assistance during rapid autopsies, and S.S. Chae and D. Wilkes for technical assistance and constructive comments. We would also like to thank T.Y. MacDonald, J. Padilla and T. Fedrizzi for technical assistance. Work was partially supported by the Translational Research Program at WCMC Pathology and Laboratory Medicine. This work was supported by a Conquer Cancer Foundation and the John and Elizabeth Leonard Family Foundation Young Investigator Award (B.M.F.), NIH/NCATS KL2TR000458 (B.M.F.), Early Detection Research Network US NCI 5U01 CA111275-09 (J.M.M., M.A.R. and F.D.), Damon Runyon Cancer Research Foundation Clinical Investigator Award CI-67-13 (H.B.), and H2020 European Research Council ERC-CoG 648670 (F.D.).
Author information
Authors and Affiliations
Contributions
Initiation and design of study: B.M.F., H.B., M.A.R. and F.D. Subject enrollment and sample and clinical data collection: B.M.F., H.B., D.M.N., S.T.T., A.M.M., C.S., J.M.M. and B.R. Statistical and bioinformatics analyses: D.P., O.E., A.S. and F.D. Supervision of research: B.M.F., M.A.R., J.R. and F.D. Writing of the first draft of the manuscript: B.M.F., D.P., H.B., M.A.R. and F.D. All authors contributed to the writing and editing of the revised manuscript and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Analysis of mutated genes as reported in TCGA bladder study.
Each row represents a gene harboring frequent SNVs (middle panel) or somatic copy number (bottom panel). Annotation rows include, from top to bottom, tumor ploidy, treatment information, biopsy site, patient’s gender, matched samples. Top bars report the otal number of non-silent SNVs in each sample. Bars on the left indicate the per-sample frequency of aberration in the cohort. NA, not available.
Supplementary Figure 2 SNVs burden comparison.
Per sample number of all SNVs (left boxplot) and only non-silent SNVs (right boxplot) in pre-chemotherapy and post-chemotherapy tumors. No statistical difference.
Supplementary Figure 3 Single nucleotide variants (SNV) validation by targeted sequencing (N250 panel).
Each dot represents a genomic position corresponding to an SNV sequenced with WES and with N250 approaches. Reported values are the ratios between the number of reads supporting the alternative base and the coverage. SNVs from the same patient are represented with the same color.
Supplementary Figure 4 Discordance in the SNVs between chemotherapy-naive and chemotherapy-treated UC tumors.
(a) Percentage of shared and unique SNVs in matched chemotherapy-naive and chemotherapy-treated UC tumors (same of Fig. 2a). (b) Detail of the specific amino acid change in matched chemotherapy-naive and chemotherapy-treated UC tumors. Each column represents a matched pair of pre- and post- chemotherapy tumors. Row reports the specific amino acid change. The figure highlights the wide divergence in the mutational landscape of pre- and post-chemotherapy samples.
Supplementary Figure 5 Phylogenetic trees of patients with two sequenced tumor samples.
Phylogenetic trees from 15 patients with two samples per patient.
Supplementary Figure 6 Hierarchical clusters of 131 UC tumor samples by copy-number alterations.
The plot extends Fig. 5a considering samples from both WCM and TCGA cohorts. Copy number gains are represented in red and copy number losses are represented in blue. Each column represents one tumor sample. Clinical annotations include treatment, biopsy site, cohort, and the presence of TP53 SNV.
Supplementary Figure 7 Hierarchical clusters of 86 UC tumor samples by copy-number alterations.
The plot shows allele specific copy number clusters as in Fig. 5a while considering samples from the TCGA cohort. Copy number gains are represented in red and copy number losses are represented in blue. Each column represents one tumor sample. Each row represents one of the 2503 genes used to compute TCGA clusters (Fig. S4.1 from Nature 507, 315–322, 2014)). Annotations include presence of TP53 SNV and TCGA clusters.
Supplementary Figure 8 Pairwise comparison of SNVs clonality.
Top right scatter plot reports the clonality of SNVs in matched pre-chemotherapy and post-chemotherapy samples. Bottom and left boxplots show clonality of SNVs that are private to pre-chemotherapy and post-chemotherapy samples, respectively.
Supplementary Figure 9 Comparison of FFPE and Fresh samples.
(left) Boxplot of the purity in FFPE and Fresh samples (p>0.05, Wilcoxon test). (right) Boxplot of the number of non-silent SNVs in FFPE and Fresh samples (p>0.05, Wilcoxon test).
Supplementary Figure 10 Landscape of somatic SNVs identified.
Landscape of somatic SNVs identified.
Supplementary Figure 11 Comparison of shared and local SNVs considering silent and non-silent SNVs.
Comparison of the percentage of shared and local SNVs in matched chemotherapy-naive and chemotherapy-treated UC tumors when considering only non-silent SNVs (top) or all SNVs (bottom).
Supplementary Figure 12 Early branching evolution in UC using silent and non-silent SNVs.
Phylogenetic trees (top), shared and private clonally-adjusted mutations (bottom) from 6 patients with three or more tumor samples per patient.
Supplementary Figure 13 Phylogenetic tree of case WCM117 using silent and non-silent SNVs.
Silent and non-silent SNVs across 12 tumor samples (top) collected at various time points and from various anatomical locations. Fractions of tumor cells harboring each mutation represented by shades of green. Reconstruction of evolutionary tree (bottom).
Supplementary Figure 14 Clonal enrichment of mutations in chemotherapy-treated UC considering silent and non-silent SNVs.
Clonal enrichment of mutations in chemotherapy-treated UC considering silent and non-silent SNVs.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–14 and Supplementary Note (PDF 3323 kb)
Supplementary Tables 1–11
Supplementary Tables 1–11 (XLSX 5139 kb)
Rights and permissions
About this article
Cite this article
Faltas, B., Prandi, D., Tagawa, S. et al. Clonal evolution of chemotherapy-resistant urothelial carcinoma. Nat Genet 48, 1490–1499 (2016). https://doi.org/10.1038/ng.3692
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3692
This article is cited by
-
Establishment and characterization of multiple patient-derived organoids from a case of advanced endometrial cancer
Human Cell (2024)
-
ARHGAP6 inhibits bladder cancer cell viability, migration, and invasion via β-catenin signaling and enhances mitomycin C sensitivity
Human Cell (2023)
-
The evolution of non-small cell lung cancer metastases in TRACERx
Nature (2023)
-
Stepwise molecular mechanisms responsible for chemoresistance in bladder cancer cells
Cell Death Discovery (2022)
-
Serial ctDNA analysis predicts clinical progression in patients with advanced urothelial carcinoma
British Journal of Cancer (2022)