Abstract
Generalist and specialist species differ in the breadth of their ecological niches. Little is known about the niche width of obligate human pathogens. Here we analyzed a global collection of Mycobacterium tuberculosis lineage 4 clinical isolates, the most geographically widespread cause of human tuberculosis. We show that lineage 4 comprises globally distributed and geographically restricted sublineages, suggesting a distinction between generalists and specialists. Population genomic analyses showed that, whereas the majority of human T cell epitopes were conserved in all sublineages, the proportion of variable epitopes was higher in generalists. Our data further support a European origin for the most common generalist sublineage. Hence, the global success of lineage 4 reflects distinct strategies adopted by different sublineages and the influence of human migration.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Futuyma, D.J. & Moreno, G. The evolution of ecological specialization. Annu. Rev. Ecol. Syst. 19, 207–233 (1988).
Woolhouse, M.E., Webster, J.P., Domingo, E., Charlesworth, B. & Levin, B.R. Biological and biomedical implications of the co-evolution of pathogens and their hosts. Nat. Genet. 32, 569–577 (2002).
Woolhouse, M.E., Taylor, L.H. & Haydon, D.T. Population biology of multihost pathogens. Science 292, 1109–1112 (2001).
Kirzinger, M.W. & Stavrinides, J. Host specificity determinants as a genetic continuum. Trends Microbiol. 20, 88–93 (2012).
Vouga, M. & Greub, G. Emerging bacterial pathogens: the past and beyond. Clin. Microbiol. Infect. 22, 12–21 (2016).
Brites, D. & Gagneux, S. Co-evolution of Mycobacterium tuberculosis and Homo sapiens. Immunol. Rev. 264, 6–24 (2015).
Borrell, S. & Gagneux, S. Infectiousness, reproductive fitness and evolution of drug-resistant Mycobacterium tuberculosis. Int. J. Tuberc. Lung Dis. 13, 1456–1466 (2009).
Merker, M. et al. Evolutionary history and global spread of the Mycobacterium tuberculosis Beijing lineage. Nat. Genet. 47, 242–249 (2015).
Luo, T. et al. Southern East Asian origin and coexpansion of Mycobacterium tuberculosis Beijing family with Han Chinese. Proc. Natl. Acad. Sci. USA 112, 8136–8141 (2015).
Cowley, D. et al. Recent and rapid emergence of W-Beijing strains of Mycobacterium tuberculosis in Cape Town, South Africa. Clin. Infect. Dis. 47, 1252–1259 (2008).
de Jong, B.C., Antonio, M. & Gagneux, S. Mycobacterium africanum—review of an important cause of human tuberculosis in West Africa. PLoS Negl. Trop. Dis. 4, e744 (2010).
Firdessa, R. et al. Mycobacterial lineages causing pulmonary and extrapulmonary tuberculosis, Ethiopia. Emerg. Infect. Dis. 19, 460–463 (2013).
Gagneux, S. Host–pathogen coevolution in human tuberculosis. Phil. Trans. R. Soc. Lond. B 367, 850–859 (2012).
Fenner, L. et al. HIV infection disrupts the sympatric host–pathogen relationship in human tuberculosis. PLoS Genet. 9, e1003318 (2013).
Gagneux, S. et al. Variable host–pathogen compatibility in Mycobacterium tuberculosis. Proc. Natl. Acad. Sci. USA 103, 2869–2873 (2006).
Baker, L., Brown, T., Maiden, M.C. & Drobniewski, F. Silent nucleotide polymorphisms and a phylogeny for Mycobacterium tuberculosis. Emerg. Infect. Dis. 10, 1568–1577 (2004).
Reed, M.B. et al. Major Mycobacterium tuberculosis lineages associate with patient country of origin. J. Clin. Microbiol. 47, 1119–1128 (2009).
Demay, C. et al. SITVITWEB—a publicly available international multimarker database for studying Mycobacterium tuberculosis genetic diversity and molecular epidemiology. Infect. Genet. Evol. 12, 755–766 (2012).
Coscolla, M. & Gagneux, S. Consequences of genomic diversity in Mycobacterium tuberculosis. Semin. Immunol. 26, 431–444 (2014).
Coscolla, M. & Gagneux, S. Does M. tuberculosis genomic diversity explain disease diversity? Drug Discov. Today Dis. Mech. 7, e43–e59 (2010).
Comas, I. et al. Out-of-Africa migration and Neolithic coexpansion of Mycobacterium tuberculosis with modern humans. Nat. Genet. 45, 1176–1182 (2013).
Coscolla, M. et al. Novel Mycobacterium tuberculosis complex isolate from a wild chimpanzee. Emerg. Infect. Dis. 19, 969–976 (2013).
Abadia, E. et al. Resolving lineage assignation on Mycobacterium tuberculosis clinical isolates classified by spoligotyping with a new high-throughput 3R SNPs based method. Infect. Genet. Evol. 10, 1066–1074 (2010).
Filliol, I. et al. Global phylogeny of Mycobacterium tuberculosis based on single nucleotide polymorphism (SNP) analysis: insights into tuberculosis evolution, phylogenetic accuracy of other DNA fingerprinting systems, and recommendations for a minimal standard SNP set. J. Bacteriol. 188, 759–772 (2006).
Sreevatsan, S. et al. Restricted structural gene polymorphism in the Mycobacterium tuberculosis complex indicates evolutionarily recent global dissemination. Proc. Natl. Acad. Sci. USA 94, 9869–9874 (1997).
Homolka, S. et al. High resolution discrimination of clinical Mycobacterium tuberculosis complex strains based on single nucleotide polymorphisms. PLoS One 7, e39855 (2012).
Coll, F. et al. A robust SNP barcode for typing Mycobacterium tuberculosis complex strains. Nat. Commun. 5, 4812 (2014).
Tsolaki, A.G. et al. Functional and evolutionary genomics of Mycobacterium tuberculosis: insights from genomic deletions in 100 strains. Proc. Natl. Acad. Sci. USA 101, 4865–4870 (2004).
Comas, I., Homolka, S., Niemann, S. & Gagneux, S. Genotyping of genetically monomorphic bacteria: DNA sequencing in Mycobacterium tuberculosis highlights the limitations of current methodologies. PLoS One 4, e7815 (2009).
Yeboah-Manu, D. et al. Genotypic diversity and drug susceptibility patterns among M. tuberculosis complex isolates from South-Western Ghana. PLoS One 6, e21906 (2011).
Malla, B. et al. First insights into the phylogenetic diversity of Mycobacterium tuberculosis in Nepal. PLoS One 7, e52297 (2012).
Fenner, L. et al. Mycobacterium tuberculosis transmission in a country with low tuberculosis incidence: role of immigration and HIV infection. J. Clin. Microbiol. 50, 388–395 (2012).
Ballif, M. et al. Genetic diversity of Mycobacterium tuberculosis in Madang, Papua New Guinea. Int. J. Tuberc. Lung Dis. 16, 1100–1107 (2012).
Wampande, E.M. et al. Long-term dominance of Mycobacterium tuberculosis Uganda family in peri-urban Kampala-Uganda is not associated with cavitary disease. BMC Infect. Dis. 13, 484 (2013).
Ley, S.D. et al. Diversity of Mycobacterium tuberculosis and drug resistance in different provinces of Papua New Guinea. BMC Microbiol. 14, 307 (2014).
Stucki, D. et al. Two new rapid SNP-typing methods for classifying Mycobacterium tuberculosis complex into the main phylogenetic lineages. PLoS One 7, e41253 (2012).
Bouakaze, C. et al. Matrix-assisted laser desorption ionization–time of flight mass spectrometry–based single nucleotide polymorphism genotyping assay using iPLEX gold technology for identification of Mycobacterium tuberculosis complex species and lineages. J. Clin. Microbiol. 49, 3292–3299 (2011).
World Health Organization. Global Tuberculosis Control—Surveillance, Planning, Financing (WHO, 2015).
Möller, M., de Wit, E. & Hoal, E.G. Past, present and future directions in human genetic susceptibility to tuberculosis. FEMS Immunol. Med. Microbiol. 58, 3–26 (2010).
Caws, M. et al. The influence of host and bacterial genotype on the development of disseminated disease with Mycobacterium tuberculosis. PLoS Pathog. 4, e1000034 (2008).
Asante-Poku, A. et al. Mycobacterium africanum is associated with patient ethnicity in Ghana. PLoS Negl. Trop. Dis. 9, e3370 (2015).
Ng, P.C. & Henikoff, S. SIFT: predicting amino acid changes that affect protein function. Nucleic Acids Res. 31, 3812–3814 (2003).
Ellstrand, N.C. & Elam, D.R. Population genetic consequences of small population size: implications for plant conservation. Annu. Rev. Ecol. Syst. 24, 217–242 (1993).
Lanzas, F., Karakousis, P.C., Sacchettini, J.C. & Ioerger, T.R. Multidrug-resistant tuberculosis in Panama is driven by clonal expansion of a multidrug-resistant Mycobacterium tuberculosis strain related to the KZN extensively drug-resistant M. tuberculosis strain from South Africa. J. Clin. Microbiol. 51, 3277–3285 (2013).
Bryant, J.M. et al. Inferring patient to patient transmission of Mycobacterium tuberculosis from whole genome sequencing data. BMC Infect. Dis. 13, 110 (2013).
Bryant, J.M. et al. Whole-genome sequencing to establish relapse or re-infection with Mycobacterium tuberculosis: a retrospective observational study. Lancet Respir. Med. 1, 786–792 (2013).
Gardy, J.L. et al. Whole-genome sequencing and social-network analysis of a tuberculosis outbreak. N. Engl. J. Med. 364, 730–739 (2011).
Casali, N. et al. Evolution and transmission of drug-resistant tuberculosis in a Russian population. Nat. Genet. 46, 279–286 (2014).
Walker, T.M. et al. Whole-genome sequencing to delineate Mycobacterium tuberculosis outbreaks: a retrospective observational study. Lancet Infect. Dis. 13, 137–146 (2013).
Roetzer, A. et al. Whole genome sequencing versus traditional genotyping for investigation of a Mycobacterium tuberculosis outbreak: a longitudinal molecular epidemiological study. PLoS Med. 10, e1001387 (2013).
Farhat, M.R. et al. Genomic analysis identifies targets of convergent positive selection in drug-resistant Mycobacterium tuberculosis. Nat. Genet. 45, 1183–1189 (2013).
Clark, T.G. et al. Elucidating emergence and transmission of multidrug-resistant tuberculosis in treatment experienced patients by whole genome sequencing. PLoS One 8, e83012 (2013).
Jamieson, F.B. et al. Whole-genome sequencing of the Mycobacterium tuberculosis Manila sublineage results in less clustering and better resolution than mycobacterial interspersed repetitive-unit-variable-number tandem-repeat (MIRU-VNTR) typing and spoligotyping. J. Clin. Microbiol. 52, 3795–3798 (2014).
Pérez-Lago, L. et al. Whole genome sequencing analysis of intrapatient microevolution in Mycobacterium tuberculosis: potential impact on the inference of tuberculosis transmission. J. Infect. Dis. 209, 98–108 (2014).
Guerra-Assunção, J.A. et al. Recurrence due to relapse or reinfection with Mycobacterium tuberculosis: a whole-genome sequencing approach in a large, population-based cohort with a high HIV infection prevalence and active follow-up. J. Infect. Dis. 211, 1154–1163 (2015).
Guerra-Assunção, J.A. et al. Large-scale whole genome sequencing of M. tuberculosis provides insights into transmission in a high prevalence area. eLife 4, e05166 (2015).
Comas, I. et al. Human T cell epitopes of Mycobacterium tuberculosis are evolutionarily hyperconserved. Nat. Genet. 42, 498–503 (2010).
Coscolla, M. et al. M. tuberculosis T cell epitope analysis reveals paucity of antigenic variation and identifies rare variable TB antigens. Cell Host Microbe 18, 538–548 (2015).
Deitsch, K.W., Lukehart, S.A. & Stringer, J.R. Common strategies for antigenic variation by bacterial, fungal and protozoan pathogens. Nat. Rev. Microbiol. 7, 493–503 (2009).
Ernst, J.D. et al. Meeting Report: NIH Workshop on the Tuberculosis Immune Epitope Database. Tuberculosis (Edinb.) 88, 366–370 (2008).
Pepperell, C.S. et al. The role of selection in shaping diversity of natural M. tuberculosis populations. PLoS Pathog. 9, e1003543 (2013).
Zhang, Y.J. et al. Global assessment of genomic regions required for growth in Mycobacterium tuberculosis. PLoS Pathog. 8, e1002946 (2012).
Esparza, M. et al. PstS-1, the 38-kDa Mycobacterium tuberculosis glycoprotein, is an adhesin, which binds the macrophage mannose receptor and promotes phagocytosis. Scand. J. Immunol. 81, 46–55 (2015).
Bekmurzayeva, A., Sypabekova, M. & Kanayeva, D. Tuberculosis diagnosis using immunodominant, secreted antigens of Mycobacterium tuberculosis. Tuberculosis (Edinb.) 93, 381–388 (2013).
Nagai, S., Wiker, H.G., Harboe, M. & Kinomoto, M. Isolation and partial characterization of major protein antigens in the culture fluid of Mycobacterium tuberculosis. Infect. Immun. 59, 372–382 (1991).
Juárez, M.D., Torres, A. & Espitia, C. Characterization of the Mycobacterium tuberculosis region containing the mpt83 and mpt70 genes. FEMS Microbiol. Lett. 203, 95–102 (2001).
Coppola, M. et al. Synthetic long peptide derived from Mycobacterium tuberculosis latency antigen Rv1733c protects against tuberculosis. Clin. Vaccine Immunol. 22, 1060–1069 (2015).
Araujo, L.S. et al. Profile of interferon-γ response to latency-associated and novel in vivo expressed antigens in a cohort of subjects recently exposed to Mycobacterium tuberculosis. Tuberculosis (Edinb.) 95, 751–757 (2015).
Yu, Y., Harris, A.J., Blair, C. & He, X. RASP (Reconstruct Ancestral State in Phylogenies): a tool for historical biogeography. Mol. Phylogenet. Evol. 87, 46–49 (2015).
Brudey, K. et al. Mycobacterium tuberculosis complex genetic diversity: mining the fourth international spoligotyping database (SpolDB4) for classification, population genetics and epidemiology. BMC Microbiol. 6, 23 (2006).
Comas, I. et al. Population genomics of Mycobacterium tuberculosis in Ethiopia contradicts the virgin soil hypothesis for human tuberculosis in sub-Saharan Africa. Curr. Biol. 25, 3260–3266 (2015).
Bos, K.I. et al. Pre-Columbian mycobacterial genomes reveal seals as a source of New World human tuberculosis. Nature 514, 494–497 (2014).
Kay, G.L. et al. Eighteenth-century genomes show that mixed infections were common at time of peak tuberculosis in Europe. Nat. Commun. 6, 6717 (2015).
Lazzarini, L.C. et al. Discovery of a novel Mycobacterium tuberculosis lineage that is a major cause of tuberculosis in Rio de Janeiro, Brazil. J. Clin. Microbiol. 45, 3891–3902 (2007).
Perdigão, J. et al. Unraveling Mycobacterium tuberculosis genomic diversity and evolution in Lisbon, Portugal, a highly drug resistant setting. BMC Genomics 15, 991 (2014).
Mokrousov, I. et al. Latin-American–Mediterranean lineage of Mycobacterium tuberculosis: human traces across pathogen's phylogeography. Mol. Phylogenet. Evol. 99, 133–143 (2016).
Baily, S.L. & Miguez, E.J. Mass Migration to Modern Latin America. (Scholarly Resources,1993).
Bates, J.H. & Stead, W.W. The history of tuberculosis as a global epidemic. Med. Clin. North Am. 77, 1205–1217 (1993).
Gagneux, S. & Small, P.M. Global phylogeography of Mycobacterium tuberculosis and implications for tuberculosis product development. Lancet Infect. Dis. 7, 328–337 (2007).
Steiner, A., Stucki, D., Coscolla, M., Borrell, S. & Gagneux, S. KvarQ: targeted and direct variant calling from fastq reads of bacterial genomes. BMC Genomics 15, 881 (2014).
Bolger, A.M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Tamura, K. et al. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011).
Paradis, E., Claude, J. & Strimmer, K. APE: Analyses of Phylogenetics and Evolution in R language. Bioinformatics 20, 289–290 (2004).
Excoffier, L., Laval, G. & Schneider, S. Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evol. Bioinform. Online 1, 47–50 (2007).
Waterhouse, A.M., Procter, J.B., Martin, D.M., Clamp, M. & Barton, G.J. Jalview Version 2—a multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191 (2009).
Hartl, D. & Clarck, A.G. Principles of Population Genetics (Sinauer, 2007).
Lew, J.M., Kapopoulou, A., Jones, L.M. & Cole, S.T. TubercuList—10 years after. Tuberculosis (Edinb.) 91, 1–7 (2011).
Ronquist, F. & Huelsenbeck, J.P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572–1574 (2003).
Altschul, S.F., Gish, W., Miller, W., Myers, E.W. & Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Acknowledgements
We thank S. Lecher, S. Li and J. Zallet for technical support. Calculations were performed at the sciCORE scientific computing core facility at the University of Basel. This work was supported by the Swiss National Science Foundation (grants 310030_166687 (S.G.) and 320030_153442 (M.E.) and Swiss HIV Cohort Study grant 740 to L.F.), the European Research Council (309540-EVODRTB to S.G.), TB-PAN-NET (FP7-223681 to S.N.), PathoNgenTrace projects (FP7-278864-2 to S.N.), SystemsX.ch (S.G.), the German Center for Infection Research (DZIF; S.N.), the Novartis Foundation (S.G.), the Natural Science Foundation of China (91631301 to Q.G.), and the National Institute of Allergy and Infectious Diseases (5U01-AI069924-05) of the US National Institutes of Health (M.E.).
Author information
Authors and Affiliations
Contributions
D.S., D. Brites, L.J., S.N. and S.G. planned the experiments. D.S., L.J., D. Brites, A.T., L.F., L.R., S.B., M. Ballif, Q.L., T.L., Q.G., M.K.-M., M. Bonnet, M.E., R.M., H.M., M.M., G.T.V., J.F., M.G., J.T., F.J., J.L.G., A.A.-P., D.Y.-M., E.W., W.S., M.J., W.H.B., I.B., J.B., M.S., S.E.G.V., P. Suffys, A.K., R.W., L.G.-B., B.M., S.D.L., H.-P.B., B.C.d.J., K.T., E.S.-P., M. Bonnet, A.G.-B., M.F., V.N.P.B., K.E., I.A., P.W.N., G.R., F.G., S. Akter, F.N., L.S.-I., N.E.N., A.R., M.H., D.M.C., G.S., S.H., D. Bakonyte, P. Stakenas, R.D., V.C., O.M., S. Al-Hajoj, L.O., F.B., E.J.C., L.D., P. Supply and I.C. contributed reagents and performed the experiments. D.S., L.J., D. Brites, M.C., S.N. and S.G. analyzed the data. D.S., L.J., D. Brites, S.N. and S.G. wrote the manuscript. All authors critically reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
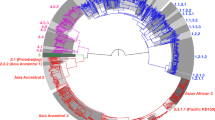
Supplementary Figure 1 Maximum-likelihood phylogeny of 72 MTBC lineage 4 isolates and 9,455 variable single-nucleotide positions.
Sublineage names were adapted from Coll et al. Newly identified or newly named sublineages are labeled in red. Sublineage-specific markers are indicated in black boxes on the branches. Gray boxes on branches indicate large sequence polymorphisms (LSP/RD) described previously. Black circles indicate clade-specific SNPs published previously.
Supplementary Figure 2 Principal component analysis of 9,455 variable single-nucleotide positions in 72 lineage 4 strains.
Each dot represents one of 72 MTBC lineage 4 strains. Colors correspond to those in Figure 1.
Supplementary Figure 3 Mean pairwise SNP distances within and between isolates of the ten sublineages of lineage 4.
Purple circles represent mean pairwise numbers of SNPs for isolates within sublineages, and blue squares represent mean pairwise numbers of SNPs between isolates of the corresponding sublineage and all other lineage 4 isolates. Error bars are single standard deviation. Total mean pairwise distances between sublineages were significantly higher than total pairwise distances within sublineages (Wilcoxon rank-sum test, P < 0.0001).
Supplementary Figure 4 Isolates of three sublineages of MTBC lineage 4 (L4.1.2/Haarlem, L4.3/LAM and L4.10/PGG3) were observed in more than 40 countries each.
L4.3/LAM was found at a proportion of 30–100% in more than 50% of the countries of its occurrence. The differences in proportions among sublineages were statistically significant (χ2 test, P = 0.001).
Supplementary Figure 5 Additional heat maps of the proportions of three sublineages that were intermediate in distribution and numbers of countries of occurrence.
Intensity of red corresponds to the proportion of each sublineage among all lineage 4 isolates. Scale is identical to that in Figure 3. Countries with fewer than three isolates in total are filled white. Gray fill means the sublineage is not found in the country. A total of 3,366 isolates were used.
Supplementary Figure 6 World maps showing the proportions of all sublineages, normalized by TB prevalence (WHO, 2013) and area of the country.
Intensity of red corresponds to the proportion of each sublineage among all lineage 4 isolates. Countries with fewer than three isolates in total are filled white. Gray fill means that the sublineage is not found in the country. A total of 3,366 isolates were used. (A) Specialist sublineages. (B) Generalist sublineages. (C) Intermediate sublineages.
Supplementary Figure 7 Minimum spanning tree (MST) based on 24 MIRU-VNTR typing data for a worldwide collection of 2,132 L4.3/LAM strains.
The length of the branches (continuous, dashed and dotted lines) denotes the number of allele changes between two patterns: solid lines represent 1, 2 or 3 changes; gray dashed lines represent 4 changes; and gray dotted lines represent 5 or more changes. Strains selected for genome sequencing are colored in red.
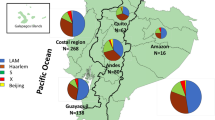
Supplementary Figure 8 Geographical distribution of the 293 L4.3/LAM isolates from 57 countries.
The 293 L4.3/LAM strains selected for genome sequencing covered all continents and included 129 strains from Africa (22 countries), 42 strains from the Americas (9 countries), 36 strains from Asia (12 countries), 83 strains from Europe (12 countries) and 3 strains from Oceania (2 countries).
Supplementary Figure 9 Maximum-likelihood phylogeny of 293 L4.3/LAM strains.
Label colors indicate continent of strain origin (blue, Europe/Mediterranean; red, sub-Saharan Africa; yellow, Americas; pink, Asia). Numbers on nodes show branch support from bootstrapping (500 pseudoreplicates). Circles indicate clades with deleted regions of difference (RD). H37Rv was used as the outgroup.
Supplementary Figure 10 Genome-based Bayesian phylogeny of 203 strains of the L4.6.1/Uganda sublineage.
H37Rv was used as the outgroup. Numbers on the nodes indicate posterior probabilities. Continent colors correspond to those in Supplementary Figure 9.
Supplementary Figure 11 Genome-based maximum-likelihood phylogeny of 203 strains of the L4.6.1/Uganda sublineage.
H37Rv was used as the outgroup. Number on the nodes indicate bootstrap support. Continent colors correspond to those in Supplementary Figure 9.
Supplementary Figure 12 Maximum-likelihood phylogeny of 228 strains of the L4.2/Haarlem sublineage based on an alignment of 15,567 variable positions.
H37Rv was used as the outgroup. Numbers on the nodes indicate posterior probabilities. Continent colors correspond to those in Supplementary Figure 9.
Supplementary Figure 13 Maximum-likelihood phylogeny of 301 strains of the L4.10/PGG3 sublineage based on an alignment of 25,678 variable positions.
train of L4.2/Haarlem sublineage was used as the outgroup. Number on the node indicate bootstrap support.
Supplementary Figure 14 Genome-based phylogeny of 293 strains of the L4.3/LAM sublineage.
As in Figure 6 but with reconstructed ancestral geographical regions using a maximum-parsimony method (S-DIVA). Bayesian phylogeny with label colors indicating continent of strain origin (blue, Europe/Mediterranean; red, sub-Saharan Africa; yellow, Americas; pink, Asia). Numbers on nodes indicate posterior probabilities from MrBayes. Pie charts show the reconstructed ancestral geographical regions of the internal nodes using a maximum-parsimony method (S-DIVA in RASP software). The hypothetical L4.3/LAM ancestor is indicated, and a European origin for this ancestor was supported. The colors correspond to continents.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–14 and Supplementary Tables 1–4, 6 and 8. (PDF 2896 kb)
Supplementary Table 5
Distribution of MTBC lineage 4 clinical isolates per country and per sublineage. (XLSX 24 kb)
Supplementary Table 7
Whole-genome sequence data accession codes. (XLSX 72 kb)
Supplementary Table 9
Description of epitopes containing non-singleton nonsynonymous mutations in the four sublineages analyzed. (XLSX 16 kb)
Rights and permissions
About this article
Cite this article
Stucki, D., Brites, D., Jeljeli, L. et al. Mycobacterium tuberculosis lineage 4 comprises globally distributed and geographically restricted sublineages. Nat Genet 48, 1535–1543 (2016). https://doi.org/10.1038/ng.3704
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3704
This article is cited by
-
Characterization of pig tonsils as niches for the generation of Streptococcus suis diversity
Veterinary Research (2024)
-
Whole Genome Sequence Dataset of Mycobacterium tuberculosis Strains from Patients of Campania Region
Scientific Data (2024)
-
Molecular investigations of Mycobacterium tuberculosis genotypes among baseline and follow-up strains circulating in four regions of Eswatini
BMC Infectious Diseases (2023)
-
Molecular epidemiology and transmission dynamics of multi-drug resistant tuberculosis strains using whole genome sequencing in the Amhara region, Ethiopia
BMC Genomics (2023)
-
An overview of tuberculosis outbreaks reported in the years 2011–2020
BMC Infectious Diseases (2023)