Abstract
Most eukaryotic genomes contain substantial amounts of repetitive DNA organized in the form of constitutive heterochromatin and associated with repressive epigenetic modifications, such as H3K9me3 and C5 cytosine methylation (5mC). In the fungus Neurospora crassa, H3K9me3 and 5mC are catalyzed, respectively, by a conserved SUV39 histone methyltransferase, DIM-5, and a DNMT1-like cytosine methyltransferase, DIM-2. Here we show that DIM-2 can also mediate repeat-induced point mutation (RIP) of repetitive DNA in N. crassa. We further show that DIM-2-dependent RIP requires DIM-5, HP1, and other known heterochromatin factors, implying a role for a repeat-induced heterochromatin-related process. Our previous findings suggest that the mechanism of repeat recognition for RIP involves direct interactions between homologous double-stranded DNA (dsDNA) segments. We thus now propose that, in somatic cells, homologous dsDNA–dsDNA interactions between a small number of repeat copies can nucleate a transient heterochromatic state, which, on longer repeat arrays, may lead to the formation of constitutive heterochromatin.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Almouzni, G. & Probst, A.V. Heterochromatin maintenance and establishment: lessons from the mouse pericentromere. Nucleus 2, 332–338 (2011).
Becker, J.S., Nicetto, D. & Zaret, K.S. H3K9me3-dependent heterochromatin: barrier to cell fate changes. Trends Genet. 32, 29–41 (2016).
Martienssen, R. & Moazed, D. RNAi and heterochromatin assembly. Cold Spring Harb. Perspect. Biol. 7, a019323 (2015).
Padeken, J., Zeller, P. & Gasser, S.M. Repeat DNA in genome organization and stability. Curr. Opin. Genet. Dev. 31, 12–19 (2015).
Plohl, M., Meštrović, N. & Mravinac, B. Centromere identity from the DNA point of view. Chromosoma 123, 313–325 (2014).
Saksouk, N., Simboeck, E. & Déjardin, J. Constitutive heterochromatin formation and transcription in mammals. Epigenetics Chromatin 8, 3 (2015).
Aramayo, R. & Selker, E.U. Neurospora crassa, a model system for epigenetics research. Cold Spring Harb. Perspect. Biol. 5, a017921 (2013).
Bull, J.H. & Wootton, J.C. Heavily methylated amplified DNA in transformants of Neurospora crassa. Nature 310, 701–704 (1984).
Selker, E.U. & Garrett, P.W. DNA sequence duplications trigger gene inactivation in Neurospora crassa. Proc. Natl. Acad. Sci. USA 85, 6870–6874 (1988).
Freitag, M. et al. DNA methylation is independent of RNA interference in Neurospora. Science 304, 1939 (2004).
Chicas, A., Forrest, E.C., Sepich, S., Cogoni, C. & Macino, G. Small interfering RNAs that trigger posttranscriptional gene silencing are not required for the histone H3 Lys9 methylation necessary for transgenic tandem repeat stabilization in Neurospora crassa. Mol. Cell. Biol. 25, 3793–3801 (2005).
Garrick, D., Fiering, S., Martin, D.I. & Whitelaw, E. Repeat-induced gene silencing in mammals. Nat. Genet. 18, 56–59 (1998).
Preis, J.I., Downes, M., Oates, N.A., Rasko, J.E. & Whitelaw, E. Sensitive flow cytometric analysis reveals a novel type of parent-of-origin effect in the mouse genome. Curr. Biol. 13, 955–959 (2003).
van der Maarel, S.M., Tawil, R. & Tapscott, S.J. Facioscapulohumeral muscular dystrophy and DUX4: breaking the silence. Trends Mol. Med. 17, 252–258 (2011).
Galagan, J.E. & Selker, E.U. RIP: the evolutionary cost of genome defense. Trends Genet. 20, 417–423 (2004).
Hane, J. et al. in Genetic Transformation Systems in Fungi (eds. van den Breg, M.A. & Maruthachalam, K.) Vol. 2, 55–68 (Springer International Publishing Switzerland, 2015).
Rossignol, J.L. & Faugeron, G. Gene inactivation triggered by recognition between DNA repeats. Experientia 50, 307–317 (1994).
Gladyshev, E. & Kleckner, N. Direct recognition of homology between double helices of DNA in Neurospora crassa. Nat. Commun. 5, 3509 (2014).
Gladyshev, E. & Kleckner, N. Recombination-independent recognition of DNA homology for repeat-induced point mutation (RIP) is modulated by the underlying nucleotide sequence. PLoS Genet. 12, e1006015 (2016).
Gladyshev, E. & Kleckner, N. Recombination-independent recognition of DNA homology for repeat-induced point mutation. Curr. Genet. http://dx.doi.org/10.1007/s00294-016-0649-4 (2016).
Freitag, M., Williams, R.L., Kothe, G.O. & Selker, E.U. A cytosine methyltransferase homologue is essential for repeat-induced point mutation in Neurospora crassa. Proc. Natl. Acad. Sci. USA 99, 8802–8807 (2002).
Clutterbuck, A.J. Genomic evidence of repeat-induced point mutation (RIP) in filamentous ascomycetes. Fungal Genet. Biol. 48, 306–326 (2011).
Amselem, J., Lebrun, M.H. & Quesneville, H. Whole genome comparative analysis of transposable elements provides new insight into mechanisms of their inactivation in fungal genomes. BMC Genomics 16, 141 (2015).
Kouzminova, E. & Selker, E.U. dim-2 encodes a DNA methyltransferase responsible for all known cytosine methylation in Neurospora. EMBO J. 20, 4309–4323 (2001).
Jamieson, K. et al. Loss of HP1 causes depletion of H3K27me3 from facultative heterochromatin and gain of H3K27me2 at constitutive heterochromatin. Genome Res. 26, 97–107 (2016).
Basenko, E.Y. et al. Genome-wide redistribution of H3K27me3 is linked to genotoxic stress and defective growth. Proc. Natl. Acad. Sci. USA 112, E6339–E6348 (2015).
Zhang, X. et al. Structure of the Neurospora SET domain protein DIM-5, a histone H3 lysine methyltransferase. Cell 111, 117–127 (2002).
Tropschug, M., Barthelmess, I.B. & Neupert, W. Sensitivity to cyclosporin A is mediated by cyclophilin in Neurospora crassa and Saccharomyces cerevisiae. Nature 342, 953–955 (1989).
Roberts, R.J., Vincze, T., Posfai, J. & Macelis, D. REBASE—a database for DNA restriction and modification: enzymes, genes and genomes. Nucleic Acids Res. 43, D298–D299 (2015).
Pontecorvo, G. Structure of heterochromatin. Nature 153, 365–367 (1944).
Dorer, D.R. & Henikoff, S. Expansions of transgene repeats cause heterochromatin formation and gene silencing in Drosophila. Cell 77, 993–1002 (1994).
Malagnac, F. et al. A gene essential for de novo methylation and development in Ascobolus reveals a novel type of eukaryotic DNA methyltransferase structure. Cell 91, 281–290 (1997).
Selker, E.U., Fritz, D.Y. & Singer, M.J. Dense nonsymmetrical DNA methylation resulting from repeat-induced point mutation in Neurospora. Science 262, 1724–1728 (1993).
Belden, W.J., Lewis, Z.A., Selker, E.U., Loros, J.J. & Dunlap, J.C. CHD1 remodels chromatin and influences transient DNA methylation at the clock gene frequency. PLoS Genet. 7, e1002166 (2011).
Ohm, R.A. et al. Diverse lifestyles and strategies of plant pathogenesis encoded in the genomes of eighteen Dothideomycetes fungi. PLoS Pathog. 8, e1003037 (2012).
Bardiya, N. & Shiu, P.K. Cyclosporin A–resistance based gene placement system for Neurospora crassa. Fungal Genet. Biol. 44, 307–314 (2007).
Freitag, M., Hickey, P.C., Raju, N.B., Selker, E.U. & Read, N.D. GFP as a tool to analyze the organization, dynamics and function of nuclei and microtubules in Neurospora crassa. Fungal Genet. Biol. 41, 897–910 (2004).
Freitag, M. & Selker, E.U. Expression and visualization of red fluorescent protein (RFP) in Neurospora crassa. Fungal Genet. Newsl. 52, 14–17 (2005).
McCluskey, K., Wiest, A. & Plamann, M. The Fungal Genetics Stock Center: a repository for 50 years of fungal genetics research. J. Biosci. 35, 119–126 (2010).
Arnaise, S., Zickler, D., Bourdais, A., Dequard-Chablat, M. & Debuchy, R. Mutations in mating-type genes greatly decrease repeat-induced point mutation process in the fungus Podospora anserina. Fungal Genet. Biol. 45, 207–220 (2008).
Acknowledgements
We thank D. Zickler for critically reading the manuscript. This work was supported by grant GM044794 from the US National Institutes of Health to N.K. and research fellowships from the Helen Hay Whitney Foundation, the Howard Hughes Medical Institute and the Charles A. King Trust to E.G.
Author information
Authors and Affiliations
Contributions
E.G. designed, performed and analyzed the data from all experiments. E.G. and N.K. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Analysis of RIP mutation using the 802-bp tester construct.
(a) The 802-bp tester construct is created by replacing the cyclosporin-resistant-1 (csr-1) gene with transgenic DNA containing 802 bp of perfect homology to a neighboring gene (NCU00725). (b) RIP mutations are detected by sequencing the entire tester construct (recovered by PCR) from individual ‘late-arising’ haploid spore clones. (c) When a gene of interest is analyzed for its role in RIP in the heterozygous condition, two configurations are possible: in cis, the wild-type allele of the gene is provided together with the tester construct in the same parental nucleus (maternal or paternal); in trans, the wild-type allele is provided by one parent, whereas the tester construct is provided by the other parent (along with a corresponding gene deletion allele).
Supplementary Figure 2 Numerical analysis of RIP mutation.
(a) RIP is quantified for the crosses in Figure 1b as the mean number of mutations (per spore) in the three regions of interest: the 802-bp endogenous repeat copy (solid blue), the 802-bp ectopic repeat copy (hatched blue) and the 600-bp portion of the linker region (gray). Distributions of per-spore mutation counts are compared for each region of interest by the Kolmogorov–Smirnov test (Online Methods). (b) The mean number of RIP mutations for the crosses in Figure 2d (analyzed as in a). (c) The mean number of RIP mutations for the crosses in Figure 2f (analyzed as in a). (d) RIP mutation of the 450-bp segment of the linker region is quantified for the crosses in Figure 3 as the mean number of mutations (as in a; left) and as the percentage of the mutated DNA strands (as in ref. 18; right). In the latter case, differences in the percentage of mutated strands are evaluated for significance as raw numbers using the χ2 homogeneity test (P value indicated above the line) and Fisher’s exact test (P value below the line). Interspersed homologies are compared to the corresponding instance of random homology in the same genetic background. Error bars, s.e.m. ***P ≤ 0.001, **0.001 < P ≤ 0.01, *0.01 < P ≤ 0.05; NS, P > 0.05.
Supplementary Figure 3 Pairwise correlations of mutation counts observed within the three regions of interest.
(a) All individual C-to-T (C>T; upward tickmarks) and G-to-A (G>A; downward tickmarks) mutations are identified in the sample of 24 progeny spores. Spores are ranked by the total number of mutations. Analyzed crosses X1 (left) and X5 (right). (b) Pearson product–moment correlation coefficients, calculated independently for C-to-T (C>T) and G-to-A (G>A) mutations, for the crosses in a. Only mutations in the three regions of interest (the two 802-bp repeated sequences and the 600-bp segment of the linker region) were analyzed.
Supplementary Figure 4 The H3K27 methyltransferase SET-7 has no detectable role in RIP.
(a) DIM-5-mediated H3K9me3 opposes SET-7-mediated H3K27me3 in N. crassa (refs. 25,26). (b) RIP mutation profile of the 802-bp construct. Cross X32{24}. The number of analyzed spores is provided in brackets. (c) The mean number of RIP mutations for cross X32 (analyzed as in Supplementary Fig. 2a). The wild-type cross X1 is used as the reference.
Supplementary Figure 5 DIM-2-mediated RIP mutations spread into nearby genes.
Eight progeny spores with particularly strong levels of DIM-2-mediated RIP were reanalyzed by sequencing a larger surrounding genomic region (Online Methods). The positions of the neighboring genes (corresponding to the start and stop codons of their ORFs) are indicated. The endogenous copy of the csr-1 gene (1,751 bp) is shown in blue.
Supplementary Figure 6 Detecting low levels of cytosine methylation in vegetative cells of N. crassa.
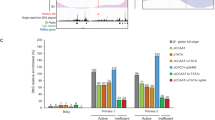
(a) A gel image with 18 individual PCR products corresponding to the three analyzed genomic regions (A, B and C) in Figure 4a. This particular experiment assays BstUI-digested DNA samples from strain T485.4h that carries the 4× array on the dim-2+; dim-5+ background. A representative graphical region of interest (ROI) that was used to quantify PCR yields in ImageJ is shown in yellow. (b) Semiquantitative PCR yields for the three genomic regions were quantified in ImageJ and are reported in raw luminosity units (mean ± s.d.). Top, DNA samples were digested with BstUI; bottom, DNA samples were mock digested. Assayed strains (left to right): FGSC#9720, T485.4h, T486.3h and T402.1h (Supplementary Table 1). The presence of intact transformed DNA was verified by PCR in mock-digested samples using primers NcHis3_F4 and NcHis3_R1 (Supplementary Table 3).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Tables 1–3. (PDF 1336 kb)
Supplementary Data 1
Plasmids used in this study. (ZIP 7 kb)
Supplementary Data 2
Sequence alignments analyzed in this study. (ZIP 265 kb)
Rights and permissions
About this article
Cite this article
Gladyshev, E., Kleckner, N. DNA sequence homology induces cytosine-to-thymine mutation by a heterochromatin-related pathway in Neurospora. Nat Genet 49, 887–894 (2017). https://doi.org/10.1038/ng.3857
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3857
This article is cited by
-
Current genetic strategies to investigate gene functions in Trichoderma reesei
Microbial Cell Factories (2023)
-
Intragenomic rDNA variation - the product of concerted evolution, mutation, or something in between?
Heredity (2023)
-
A thousand-genome panel retraces the global spread and adaptation of a major fungal crop pathogen
Nature Communications (2023)
-
Colletotrichum: species complexes, lifestyle, and peculiarities of some sources of genetic variability
Applied Microbiology and Biotechnology (2020)
-
Diversity of cytosine methylation across the fungal tree of life
Nature Ecology & Evolution (2019)