Abstract
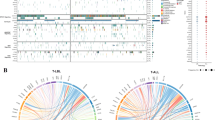
The outcome of treatment-refractory and/or relapsed pediatric T cell acute lymphoblastic leukemia (T-ALL) is extremely poor1, and the genetic basis for this is not well understood. Here we report comprehensive profiling of 121 cases of pediatric T-ALL using transcriptome and/or targeted capture sequencing, through which we identified new recurrent gene fusions involving SPI1 (STMN1-SPI1 and TCF7-SPI1). Cases positive for fusions involving SPI1 (encoding PU.1), accounting for 3.9% (7/181) of the examined pediatric T-ALL cases, showed a double-negative (DN; CD4−CD8−) or CD8+ single-positive (SP) phenotype and had uniformly poor overall survival. These cases represent a subset of pediatric T-ALL distinguishable from the known T-ALL subsets2 in terms of expression of genes involved in T cell precommitment, establishment of T cell identity, and post-β-selection maturation and with respect to mutational profile. PU.1 fusion proteins retained transcriptional activity and, when constitutively expressed in mouse stem/progenitor cells, induced cell proliferation and resulted in a maturation block. Our findings highlight a unique role of SPI1 fusions in high-risk pediatric T-ALL.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Herold, R., von Stackelberg, A., Hartmann, R., Eisenreich, B. & Henze, G. Acute lymphoblastic leukemia-relapse study of the Berlin-Frankfurt-Munster Group (ALL-REZ BFM) experience: early treatment intensity makes the difference. J. Clin. Oncol. 22, 569–570, author reply 570–571 (2004).
Belver, L. & Ferrando, A. The genetics and mechanisms of T cell acute lymphoblastic leukaemia. Nat. Rev. Cancer 16, 494–507 (2016).
Kataoka, K. et al. Integrated molecular analysis of adult T cell leukemia/lymphoma. Nat. Genet. 47, 1304–1315 (2015).
Aplan, P.D. et al. Disruption of the human SCL locus by “illegitimate” V-(D)-J recombinase activity. Science 250, 1426–1429 (1990).
Rubnitz, J.E. et al. Childhood acute lymphoblastic leukemia with the MLL-ENL fusion and t(11;19)(q23;p13.3) translocation. J. Clin. Oncol. 17, 191–196 (1999).
Prasad, R. et al. Leucine-zipper dimerization motif encoded by the AF17 gene fused to ALL-1 (MLL) in acute leukemia. Proc. Natl. Acad. Sci. USA 91, 8107–8111 (1994).
Groupe Français de Cytogénétique Hématologique. t(10;11)(p13-14;q14-21): a new recurrent translocation in T-cell acute lymphoblastic leukemias. Groupe Français de Cytogénétique Hématologique (GFCH). Genes Chromosom. Cancer 3, 411–415 (1991).
Meyer, C. et al. New insights to the MLL recombinome of acute leukemias. Leukemia 23, 1490–1499 (2009).
Babusikova, O., Stevulova, L. & Fajtova, M. Immunophenotyping parameters as prognostic factors in T-acute leukemia patients. Neoplasma 56, 508–513 (2009).
Moreau-Gachelin, F., Tavitian, A. & Tambourin, P. Spi-1 is a putative oncogene in virally induced murine erythroleukaemias. Nature 331, 277–280 (1988).
Moreau-Gachelin, F. et al. Spi-1 oncogene activation in Rauscher and Friend murine virus–induced acute erythroleukemias. Leukemia 4, 20–23 (1990).
Kodandapani, R. et al. A new pattern for helix-turn-helix recognition revealed by the PU.1 ETS-domain-DNA complex. Nature 380, 456–460 (1996).
Carotta, S., Wu, L. & Nutt, S.L. Surprising new roles for PU.1 in the adaptive immune response. Immunol. Rev. 238, 63–75 (2010).
Mak, K.S., Funnell, A.P., Pearson, R.C. & Crossley, M. PU.1 and haematopoietic cell fate: dosage matters. Int. J. Cell Biol. 2011, 808524 (2011).
McKercher, S.R. et al. Targeted disruption of the PU.1 gene results in multiple hematopoietic abnormalities. EMBO J. 15, 5647–5658 (1996).
Scott, E.W., Simon, M.C., Anastasi, J. & Singh, H. Requirement of transcription factor PU.1 in the development of multiple hematopoietic lineages. Science 265, 1573–1577 (1994).
Back, J., Dierich, A., Bronn, C., Kastner, P. & Chan, S. PU.1 determines the self-renewal capacity of erythroid progenitor cells. Blood 103, 3615–3623 (2004).
Nutt, S.L. & Kee, B.L. The transcriptional regulation of B cell lineage commitment. Immunity 26, 715–725 (2007).
Laslo, P., Pongubala, J.M., Lancki, D.W. & Singh, H. Gene regulatory networks directing myeloid and lymphoid cell fates within the immune system. Semin. Immunol. 20, 228–235 (2008).
Champhekar, A. et al. Regulation of early T-lineage gene expression and developmental progression by the progenitor cell transcription factor PU.1. Genes Dev. 29, 832–848 (2015).
Rothenberg, E.V., Moore, J.E. & Yui, M.A. Launching the T-cell-lineage developmental programme. Nat. Rev. Immunol. 8, 9–21 (2008).
Nutt, S.L., Metcalf, D., D'Amico, A., Polli, M. & Wu, L. Dynamic regulation of PU.1 expression in multipotent hematopoietic progenitors. J. Exp. Med. 201, 221–231 (2005).
Del Real, M.M. & Rothenberg, E.V. Architecture of a lymphomyeloid developmental switch controlled by PU.1, Notch and Gata3. Development 140, 1207–1219 (2013).
Dik, W.A. et al. New insights on human T cell development by quantitative T cell receptor gene rearrangement studies and gene expression profiling. J. Exp. Med. 201, 1715–1723 (2005).
Zhang, D.E., Hetherington, C.J., Chen, H.M. & Tenen, D.G. The macrophage transcription factor PU.1 directs tissue-specific expression of the macrophage colony-stimulating factor receptor. Mol. Cell. Biol. 14, 373–381 (1994).
Miyazaki, M. et al. Polycomb group gene mel-18 regulates early T progenitor expansion by maintaining the expression of Hes-1, a target of the Notch pathway. J. Immunol. 174, 2507–2516 (2005).
Masuda, K. et al. Prethymic T-cell development defined by the expression of paired immunoglobulin-like receptors. EMBO J. 24, 4052–4060 (2005).
Ferrando, A.A. et al. Gene expression signatures define novel oncogenic pathways in T cell acute lymphoblastic leukemia. Cancer Cell 1, 75–87 (2002).
Soulier, J. et al. HOXA genes are included in genetic and biologic networks defining human acute T-cell leukemia (T-ALL). Blood 106, 274–286 (2005).
Yui, M.A. & Rothenberg, E.V. Developmental gene networks: a triathlon on the course to T cell identity. Nat. Rev. Immunol. 14, 529–545 (2014).
Zhang, J. et al. The genetic basis of early T-cell precursor acute lymphoblastic leukaemia. Nature 481, 157–163 (2012).
Stehling-Sun, S., Dade, J., Nutt, S.L., DeKoter, R.P. & Camargo, F.D. Regulation of lymphoid versus myeloid fate 'choice' by the transcription factor Mef2c. Nat. Immunol. 10, 289–296 (2009).
Homminga, I. et al. Integrated transcript and genome analyses reveal NKX2-1 and MEF2C as potential oncogenes in T cell acute lymphoblastic leukemia. Cancer Cell 19, 484–497 (2011).
Zhang, J.A., Mortazavi, A., Williams, B.A., Wold, B.J. & Rothenberg, E.V. Dynamic transformations of genome-wide epigenetic marking and transcriptional control establish T cell identity. Cell 149, 467–482 (2012).
Neumann, M. et al. FLT3 mutations in early T-cell precursor ALL characterize a stem cell like leukemia and imply the clinical use of tyrosine kinase inhibitors. PLoS One 8, e53190 (2013).
Zhou, J. et al. PU.1 is essential for MLL leukemia partially via crosstalk with the MEIS/HOX pathway. Leukemia 28, 1436–1448 (2014).
Vlierberghe, P.V. & Ferrando, A. The molecular basis of T cell acute lymphoblastic leukemia. J. Clin. Invest. 122, 3398–3406 (2012).
Weng, A.P. et al. Activating mutations of NOTCH1 in human T cell acute lymphoblastic leukemia. Science 306, 269–271 (2004).
Hebert, J., Cayuela, J.M., Berkeley, J. & Sigaux, F. Candidate tumor-suppressor genes MTS1 (p16INK4A) and MTS2 (p15INK4B) display frequent homozygous deletions in primary cells from T- but not from B-cell lineage acute lymphoblastic leukemias. Blood 84, 4038–4044 (1994).
Van Vlierberghe, P. et al. PHF6 mutations in T-cell acute lymphoblastic leukemia. Nat. Genet. 42, 338–342 (2010).
O'Neil, J. et al. FBW7 mutations in leukemic cells mediate NOTCH pathway activation and resistance to γ-secretase inhibitors. J. Exp. Med. 204, 1813–1824 (2007).
Thompson, B.J. et al. The SCFFBW7 ubiquitin ligase complex as a tumor suppressor in T cell leukemia. J. Exp. Med. 204, 1825–1835 (2007).
Palomero, T. et al. Mutational loss of PTEN induces resistance to NOTCH1 inhibition in T-cell leukemia. Nat. Med. 13, 1203–1210 (2007).
Van Vlierberghe, P. et al. ETV6 mutations in early immature human T cell leukemias. J. Exp. Med. 208, 2571–2579 (2011).
Bar-Eli, M., Ahuja, H., Foti, A. & Cline, M.J. N-RAS mutations in T-cell acute lymphocytic leukaemia: analysis by direct sequencing detects a novel mutation. Br. J. Haematol. 72, 36–39 (1989).
Tosello, V. et al. WT1 mutations in T-ALL. Blood 114, 1038–1045 (2009).
Grossmann, V. et al. Prognostic relevance of RUNX1 mutations in T-cell acute lymphoblastic leukemia. Haematologica 96, 1874–1877 (2011).
Song, M.S. et al. The deubiquitinylation and localization of PTEN are regulated by a HAUSP–PML network. Nature 455, 813–817 (2008).
Aifantis, I., Raetz, E. & Buonamici, S. Molecular pathogenesis of T-cell leukaemia and lymphoma. Nat. Rev. Immunol. 8, 380–390 (2008).
Anderson, M.K., Weiss, A.H., Hernandez-Hoyos, G., Dionne, C.J. & Rothenberg, E.V. Constitutive expression of PU.1 in fetal hematopoietic progenitors blocks T cell development at the pro-T cell stage. Immunity 16, 285–296 (2002).
Laiosa, C.V., Stadtfeld, M., Xie, H., de Andres-Aguayo, L. & Graf, T. Reprogramming of committed T cell progenitors to macrophages and dendritic cells by C/EBPα and PU.1 transcription factors. Immunity 25, 731–744 (2006).
Tydell, C.C. et al. Molecular dissection of prethymic progenitor entry into the T lymphocyte developmental pathway. J. Immunol. 179, 421–438 (2007).
Rosenbauer, F. et al. Lymphoid cell growth and transformation are suppressed by a key regulatory element of the gene encoding PU.1. Nat. Genet. 38, 27–37 (2006).
Sato, Y. et al. Integrated molecular analysis of clear-cell renal cell carcinoma. Nat. Genet. 45, 860–867 (2013).
Suzuki, H. et al. Mutational landscape and clonal architecture in grade II and III gliomas. Nat. Genet. 47, 458–468 (2015).
Haferlach, T. et al. Landscape of genetic lesions in 944 patients with myelodysplastic syndromes. Leukemia 28, 241–247 (2014).
Lawrence, M.S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
Shiraishi, Y. et al. An empirical Bayesian framework for somatic mutation detection from cancer genome sequencing data. Nucleic Acids Res. 41, e89–e89 (2013).
Yoshida, K. et al. Frequent pathway mutations of splicing machinery in myelodysplasia. Nature 478, 64–69 (2011).
Seki, M. et al. Integrated genetic and epigenetic analysis defines novel molecular subgroups in rhabdomyosarcoma. Nat. Commun. 6, 7557 (2015).
Saiki, R. et al. NGS-based copy number analysis in 1,185 patients with myeloid neoplasms. Blood 128, 955 (2016).
Ono, R. et al. Dimerization of MLL fusion proteins and FLT3 activation synergize to induce multiple-lineage leukemogenesis. J. Clin. Invest. 115, 919–929 (2005).
Isoda, T. et al. Process for immune defect and chromosomal translocation during early thymocyte development lacking ATM. Blood 120, 789–799 (2012).
Acknowledgements
We gratefully acknowledge the TCGA Consortium and all its members for making their invaluable data publically available. The results published here are in whole or part based upon data generated by the Therapeutically Applicable Research to Generate Effective Treatment (TARGET) initiative managed by the NCI. Information about TARGET can be found at http://ocg.cancer.gov/programs/target. We are also grateful to M. Matsumura, N. Hoshino, K. Yin, F. Saito, Y. Mori, N. Mizota, and M. Nakamura for their excellent technical assistance. We also wish to express our appreciation to M.-J. Park (Gunma Children's Medical Hospital), K. Nomura (Toyama University), H. Kanegane (Tokyo Medical and Dental University), and K. Kato (Ibaraki Children's Hospital) for collecting samples and T. Yasuda, M. Takeyama, J. Mitsui, and S. Tsuji (University of Tokyo) for next-generation sequencing. This work was supported by KAKENHI (17H04224 (J.T.), 26713037 (M.K.), and 15H05909 (S.O.)) from the Japan Society of Promotion of Science; by Japan Agency for Medical Research and Development (AMED) Practical Research for Innovative Cancer Control (16ck0106066h0003 (M. Sanada)) and Project for Cancer Research and Therapeutic Evolution (P-CREATE) (16cm0106509h001 (J.T.)); and by the Friends of Leukemia Research Fund (147100000868 (M. Seki)). This research also used computational resources of the K computer provided by the RIKEN Advanced Institute for Computational Science through the HPCI System Research project (hp140230 (S.M.), hp160219 (S.M.), and hp150232 (S.M.)).
Author information
Authors and Affiliations
Contributions
Y. Shiraishi, K.C., H.T., T.S., and S.M. developed bioinformatics pipelines. M. Seki, S.K., T. Isobe, K.Y., H.S., Y.F., H.U., K. Kataoka, and Y. Shiozawa performed sequencing data analyses. M. Seki, S.K., T. Isobe, K.Y., M. Sanada, and H. Mano performed sequencing experiments. M. Seki, S.K., L.L., K.M., M.T., C.W., Y.N.-T., A.K., H.K., and A.I. performed functional assays. K.O., T.D., Y. Hashii, and N.K. performed FACS analyses. M. Seki, S.K., T. Isobe, J.T., and S.O. interpreted the results. M.K., Y.A., K. Koh, R.H., M.A., H. Moritake, R.K., T. Imamura, A.S., A.M., K.H., and A. Ohara collected specimens. M. Seki, S.K., J.T., and S.O. generated figures and tables and wrote the manuscript. A. Oka, Y. Hayashi, S.O., and J.T. co-led the entire project. All authors participated in discussions and interpretation of the data and results.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–13. (PDF 12003 kb)
Supplementary Table 1
Patient characteristics of 181 T-ALL cases. (XLSX 30 kb)
Supplementary Table 2
Fusion genes detected by RNA sequencing in 123 pediatric T-ALL cases. (XLSX 18 kb)
Supplementary Table 3
Clinicopathological findings of cases with SPI1 fusion. (XLSX 15 kb)
Supplementary Table 4
The number of SPI1 fusion reads and the number of wild-type SPI1 reads that span the exon–exon junction. (XLSX 14 kb)
Supplementary Table 5
Bait design for targeted capture sequencing. (XLSX 16 kb)
Supplementary Table 6
Mutations detected by targeted capture sequencing. (XLSX 84 kb)
Supplementary Table 7
Structural variants detected by targeted capture sequencing. (XLSX 17 kb)
Rights and permissions
About this article
Cite this article
Seki, M., Kimura, S., Isobe, T. et al. Recurrent SPI1 (PU.1) fusions in high-risk pediatric T cell acute lymphoblastic leukemia. Nat Genet 49, 1274–1281 (2017). https://doi.org/10.1038/ng.3900
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3900
This article is cited by
-
Molecular and clinical analyses of PHF6 mutant myeloid neoplasia provide their pathogenesis and therapeutic targeting
Nature Communications (2024)
-
Pivotal role of BCL11B in the immune, hematopoietic and nervous systems: a review of the BCL11B-associated phenotypes from the genetic perspective
Genes & Immunity (2024)
-
FANCA deficiency promotes leukaemic progression by allowing the emergence of cells carrying oncogenic driver mutations
Oncogene (2023)
-
Transcriptome sequencing identifies novel EVX fusions involved in transcriptional activation of HOX family genes in pediatric immature T-cell acute lymphoblastic leukemia: two cases reports and a literature review
International Journal of Hematology (2023)
-
Novel TENM3–ALK fusion is an alternate mechanism for ALK activation in neuroblastoma
Oncogene (2022)