Abstract
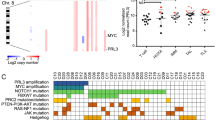
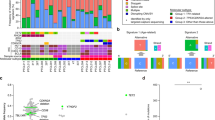
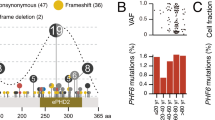
Tumor suppressor genes on the X chromosome may skew the gender distribution of specific types of cancer1,2. T-cell acute lymphoblastic leukemia (T-ALL) is an aggressive hematological malignancy with an increased incidence in males3. In this study, we report the identification of inactivating mutations and deletions in the X-linked plant homeodomain finger 6 (PHF6) gene in 16% of pediatric and 38% of adult primary T-ALL samples. Notably, PHF6 mutations are almost exclusively found in T-ALL samples from male subjects. Mutational loss of PHF6 is importantly associated with leukemias driven by aberrant expression of the homeobox transcription factor oncogenes TLX1 and TLX3. Overall, these results identify PHF6 as a new X-linked tumor suppressor in T-ALL and point to a strong genetic interaction between PHF6 loss and aberrant expression of TLX transcription factors in the pathogenesis of this disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Carrel, L., Cottle, A.A., Goglin, K.C. & Willard, H.F. A first-generation X-inactivation profile of the human X chromosome. Proc. Natl. Acad. Sci. USA 96, 14440–14444 (1999).
Carrel, L. & Willard, H.F. X-inactivation profile reveals extensive variability in X-linked gene expression in females. Nature 434, 400–404 (2005).
Goldberg, J.M. et al. Childhood T-cell acute lymphoblastic leukemia: the Dana-Farber Cancer Institute acute lymphoblastic leukemia consortium experience. J. Clin. Oncol. 21, 3616–3622 (2003).
Aifantis, I., Raetz, E. & Buonamici, S. Molecular pathogenesis of T-cell leukaemia and lymphoma. Nat. Rev. Immunol. 8, 380–390 (2008).
Pui, C.H., Robison, L.L. & Look, A.T. Acute lymphoblastic leukaemia. Lancet 371, 1030–1043 (2008).
Gnirke, A. et al. Solution hybrid selection with ultra-long oligonucleotides for massively parallel targeted sequencing. Nat. Biotechnol. 27, 182–189 (2009).
Lower, K.M. et al. Mutations in PHF6 are associated with Borjeson-Forssman-Lehmann syndrome. Nat. Genet. 32, 661–665 (2002).
Borjeson, M., Forssman, H. & Lehmann, O. An X-linked, recessively inherited syndrome characterized by grave mental deficiency, epilepsy, and endocrine disorder. Acta Med. Scand. 171, 13–21 (1962).
Turner, G. et al. The clinical picture of the Borjeson-Forssman-Lehmann syndrome in males and heterozygous females with PHF6 mutations. Clin. Genet. 65, 226–232 (2004).
Baker, L.A., Allis, C.D. & Wang, G.G. PHD fingers in human diseases: disorders arising from misinterpreting epigenetic marks. Mutat. Res. 647, 3–12 (2008).
Dephoure, N. et al. A quantitative atlas of mitotic phosphorylation. Proc. Natl. Acad. Sci. USA 105, 10762–10767 (2008).
Matsuoka, S. et al. ATM and ATR substrate analysis reveals extensive protein networks responsive to DNA damage. Science 316, 1160–1166 (2007).
Lowndes, N.F. & Toh, G.W. DNA repair: the importance of phosphorylating histone H2AX. Curr. Biol. 15, R99–R102 (2005).
Payer, B. & Lee, J.T. X chromosome dosage compensation: how mammals keep the balance. Annu. Rev. Genet. 42, 733–772 (2008).
Carrel, L. & Willard, H.F. Heterogeneous gene expression from the inactive X chromosome: an X-linked gene that escapes X inactivation in some human cell lines but is inactivated in others. Proc. Natl. Acad. Sci. USA 96, 7364–7369 (1999).
Ferrando, A.A. et al. Gene expression signatures define novel oncogenic pathways in T cell acute lymphoblastic leukemia. Cancer Cell 1, 75–87 (2002).
Soulier, J. et al. HOXA genes are included in genetic and biologic networks defining human acute T-cell leukemia (T-ALL). Blood 106, 274–286 (2005).
Van Vlierberghe, P. et al. The recurrent SET-NUP214 fusion as a new HOXA activation mechanism in pediatric T-cell acute lymphoblastic leukemia. Blood 111, 4668–4680 (2008).
van Grotel, M. et al. The outcome of molecular-cytogenetic subgroups in pediatric T-cell acute lymphoblastic leukemia: a retrospective study of patients treated according to DCOG or COALL protocols. Haematologica 91, 1212–1221 (2006).
Marks, D.I. et al. T-cell acute lymphoblastic leukemia in adults: clinical features, immunophenotype, cytogenetics, and outcome from the large randomized prospective trial (UKALL XII/ECOG 2993). Blood 114, 5136–5145 (2009).
Rumble, S.M. et al. SHRiMP: accurate mapping of short color-space reads. PLOS Comput. Biol. 5, e1000386 (2009).
Khiabanian, H., Van Vlierberghe, P., Palomero, T., Ferrando, A.A. & Rabadan, R. ParMap, an algorithm for the identification of complex genomic variations in nextgen sequencing data. Nature Precedings published online, <http://hdl.handle.net/10101/npre.2010.4145.1> (12 January 2010).
Clappier, E. et al. The C-MYB locus is involved in chromosomal translocation and genomic duplications in human T-cell acute leukemia (T-ALL), the translocation defining a new T-ALL subtype in very young children. Blood 110, 1251–1261 (2007).
Erdogan, F. et al. Impact of low copy repeats on the generation of balanced and unbalanced chromosomal aberrations in mental retardation. Cytogenet. Genome Res. 115, 247–253 (2006).
Lahortiga, I. et al. Duplication of the MYB oncogene in T cell acute lymphoblastic leukemia. Nat. Genet. 39, 593–595 (2007).
Voss, A.K. et al. Protein and gene expression analysis of Phf6, the gene mutated in the Borjeson-Forssman-Lehmann syndrome of intellectual disability and obesity. Gene Expr. Patterns 7, 858–871 (2007).
González-García, S. et al. CSL-MAML-dependent Notch1 signaling controls T lineage-specific IL-7Rα gene expression in early human thymopoiesis and leukemia. J. Exp. Med. 206, 779–791 (2009).
Moffat, J. et al. A lentiviral RNAi library for human and mouse genes applied to an arrayed viral high-content screen. Cell 124, 1283–1298 (2006).
Acknowledgements
This study was supported by the Fund for Scientific Research (FWO) Flanders (postdoctoral grants to P.V.V. and T.T., PhD grant to J.V.d.M., senior clinical investigator award to B.P. and project grants G.0198.08 and G.0869.10N to F.S.); the GOA-UGent (grant no. 12051203); the IWT-Vlaanderen (SBO grant no. 060848); the Children Cancer Fund Ghent (F.S.); Leukemia Research UK (C.J.H.); the Stichting Kinderen Kankervrij (KiKa; grant no. KiKa 2007-012 to L.Z.); the Belgian Program of Interuniversity Poles of Attraction; the Belgian Foundation against Cancer; the Austrian Ministry of Science and Research (GEN-AU Child, GZ 200.136/1-VI/1/2005 to S.S.), the US National Library of Medicine (1R01LM010140-01 to R.R. and H.K.); the ECOG and DCOG tumor banks; grants from Spain's Plan Nacional (BFU 2007-60990 and PlanE2009-0110 to M.L.T.), Comunidad de Madrid (S-SAL0304-2006 to M.L.T.), Fundación MM (M.L.T.), Instituto de Salud Carlos III (RECAVA RD06/0014/1012 to M.L.T.), an Institutional Grant from the Fundación Ramón Areces (M.L.T.), the Alex's Lemonade Stand Foundation Young Investigator Award (T.P.); a US Northeast Biodefense Center ARRA award (U54-AI057158 to R.R.); the US National Institutes of Health (R01CA120196 and R01CA129382 to A.F.); the Rally across America Foundation (A.F); the Swim across America Foundation (A.F.); the Golfers against Cancer Foundation (A.F.); and a Leukemia and Lymphoma Society Scholar Award (A.F.). We thank the Pediatric Cardiosurgery Units from Centro Especial Ramón y Cajal and Ciudad Sanitaria La Paz (Madrid, Spain) for thymus samples.
Author information
Authors and Affiliations
Contributions
P.V.V. performed array-CGH and mutation analysis of PHF6 and wrote the manuscript. T.P. performed exon capture and next-generation sequencing of T-ALL samples and wrote the manuscript. H.K. analyzed next-generation sequencing data. J.V.d.M. performed additional array-CGH analysis and PHF6 mutation screening in T-ALL and BCP-ALL samples. T.T., N.V.R. and A.W.L. performed experiments. M.C. and C.C.-C. performed and analyzed histological and immunohistochemical staining. J.P. collaborated on PHF6 mutation screening in BCP-ALL samples. C.J.H. and C.S. collaborated on additional screening for genomic PHF6 deletions in T-ALL. Y.B., B.D.M. and B.C. collaborated on the PHF6 mutation screening. R.P., M.P., S.S. and J.S. collaborated on the multicenter array-CGH study. S.G.-G. and M.L.T. performed the isolation of T-cell progenitor cells for expression analysis of PHF6. X.Y. performed survival analysis of ECOG T-ALL patients. J.G. provided critical reagents and discussion. E.S. provided samples and correlative clinical data from DCOG. E.P., J.M.R. and P.H.W. provided samples and correlative clinical data from ECOG. J.M. and L.Z. collaborated on the multicenter array-CGH study and PHF6 mutation analysis, provided molecular data on the characterization of T-ALL and performed survival analysis of PHF6 mutations in the DCOG series. R.R. designed and directed the analysis of next-generation sequencing results. F.S. and B.P. designed the studies and directed research. A.F. designed the studies, directed research and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figure 1 and Supplementary Tables 1–7 (PDF 503 kb)
Rights and permissions
About this article
Cite this article
Van Vlierberghe, P., Palomero, T., Khiabanian, H. et al. PHF6 mutations in T-cell acute lymphoblastic leukemia. Nat Genet 42, 338–342 (2010). https://doi.org/10.1038/ng.542
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.542
This article is cited by
-
PHF6 maintains acute myeloid leukemia via regulating NF-κB signaling pathway
Leukemia (2023)
-
Obituary—Pieter Van Vlierberghe (1980–2022)
Leukemia (2023)
-
The Role of PHF6 in Hematopoiesis and Hematologic Malignancies
Stem Cell Reviews and Reports (2023)
-
Börjeson–Forssman–Lehmann syndrome: delineating the clinical and allelic spectrum in 14 new families
European Journal of Human Genetics (2023)
-
Oncogenesis induced by combined Phf6 and Idh2 mutations through increased oncometabolites and impaired DNA repair
Oncogene (2022)