Abstract
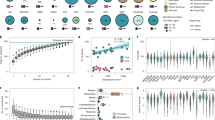
Ulcerative colitis is a chronic, relapsing inflammatory condition of the gastrointestinal tract with a complex genetic and environmental etiology. In an effort to identify genetic variation underlying ulcerative colitis risk, we present two distinct genome-wide association studies of ulcerative colitis and their joint analysis with a previously published scan1, comprising, in aggregate, 2,693 individuals with ulcerative colitis and 6,791 control subjects. Fifty-nine SNPs from 14 independent loci attained an association significance of P < 10−5. Seven of these loci exceeded genome-wide significance (P < 5 × 10−8). After testing an independent cohort of 2,009 cases of ulcerative colitis and 1,580 controls, we identified 13 loci that were significantly associated with ulcerative colitis (P < 5 × 10−8), including the immunoglobulin receptor gene FCGR2A, 5p15, 2p16 and ORMDL3 (orosomucoid1-like 3). We confirmed association with 14 previously identified ulcerative colitis susceptibility loci, and an analysis of acknowledged Crohn's disease loci showed that roughly half of the known Crohn's disease associations are shared with ulcerative colitis. These data implicate approximately 30 loci in ulcerative colitis, thereby providing insight into disease pathogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
Change history
29 March 2011
In the version of this article initially published, Kathryn Roeder's affiliation was incorrect. Her correct affiliation is the Department of Statistics, Carnegie Mellon University, Pittsburgh, Pennsylvania, USA. The error has been corrected in the HTML and PDF versions of the article.
References
Silverberg, M.S. et al. Ulcerative colitis-risk loci on chromosomes 1p36 and 12q15 found by genome-wide association study. Nat. Genet. 41, 216–220 (2009).
Barrett, J.C. et al. Genome-wide association defines more than 30 distinct susceptibility loci for Crohn's disease. Nat. Genet. 40, 955–962 (2008).
Fisher, S.A. et al. Genetic determinants of ulcerative colitis include the ECM1 locus and five loci implicated in Crohn's disease. Nat. Genet. 40, 710–712 (2008).
Franke, A. et al. Replication of signals from recent studies of Crohn's disease identifies previously unknown disease loci for ulcerative colitis. Nat. Genet. 40, 713–715 (2008).
Franke, A. et al. Sequence variants in IL10, ARPC2 and multiple other loci contribute to ulcerative colitis susceptibility. Nat. Genet. 40, 1319–1323 (2008).
Zhernakova, A. et al. Genetic analysis of innate immunity in Crohn's disease and ulcerative colitis identifies two susceptibility loci harboring CARD9 and IL18RAP. Am. J. Hum. Genet. 82, 1202–1210 (2008).
de Bakker, P.I. et al. Practical aspects of imputation-driven meta-analysis of genome-wide association studies. Hum. Mol. Genet. 17, R122–R128 (2008).
Franke, A. et al. Systematic association mapping identifies NELL1 as a novel IBD disease gene. PLoS One 2, e691 (2007).
Goyette, P. et al. Gene-centric association mapping of chromosome 3p implicates MST1 in IBD pathogenesis. Mucosal Immunol. 1, 131–138 (2008).
Barrett, J.C. et al. Genome-wide association study of ulcerative colitis identifies three new susceptibility loci, including the HNF4A region. Nat. Genet. 41, 1330–1334 (2009).
Kugathasan, S. et al. Loci on 20q13 and 21q22 are associated with pediatric-onset inflammatory bowel disease. Nat. Genet. 40, 1211–1215 (2008).
Asano, K. et al. A genome-wide association study identifies three new susceptibility loci for ulcerative colitis in the Japanese population. Nat. Genet. 41, 1325–1329 (2009).
Imielinski, M. et al. Common variants at five new loci associated with early-onset inflammatory bowel disease. Nat. Genet. 41, 1335–1340 (2009).
McGovern, D.P. et al. Genetic epistasis of IL23/IL17 pathway genes in Crohn's disease. Inflamm. Bowel Dis. 15, 883–889 (2009).
Sun, S.C. Deubiquitylation and regulation of the immune response. Nat. Rev. Immunol. 8, 501–511 (2008).
Graham, R.R. et al. Genetic variants near TNFAIP3 on 6q23 are associated with systemic lupus erythematosus. Nat. Genet. 40, 1059–1061 (2008).
Trynka, G. et al. Coeliac disease-associated risk variants in TNFAIP3 and REL implicate altered NF-κB signalling. Gut 58, 1078–1083 (2009).
Burke, J.E. & Dennis, E.A. Phospholipase A2 structure/function, mechanism, and signaling. J. Lipid Res. 50 Suppl, S237–S242 (2009).
Hirschfield, G.M. et al. Primary biliary cirrhosis associated with HLA, IL12A, and IL12RB2 variants. N. Engl. J. Med. 360, 2544–2555 (2009).
Verlaan, D.J. et al. Allele-specific chromatin remodeling in the ZPBP2/GSDMB/ORMDL3 locus associated with the risk of asthma and autoimmune disease. Am. J. Hum. Genet. 85, 377–393 (2009).
Moffatt, M.F. et al. Genetic variants regulating ORMDL3 expression contribute to the risk of childhood asthma. Nature 448, 470–473 (2007).
Kitamura, M. Biphasic, bidirectional regulation of NF-κB by endoplasmic reticulum stress. Antioxid. Redox Signal. 11, 2353–2364 (2009).
Todd, D.J., Lee, A.H. & Glimcher, L.H. The endoplasmic reticulum stress response in immunity and autoimmunity. Nat. Rev. Immunol. 8, 663–674 (2008).
Hjelmqvist, L. et al. ORMDL proteins are a conserved new family of endoplasmic reticulum membrane proteins. Genome Biol. 3, RESEARCH0027 (2002).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Plenge, R.M. et al. TRAF1-C5 is a risk locus for rheumatiod arthritis: a genomewide study. N. Engl. J. Med. 357, 1199–1209 (2007).
Wang, Y. et al. Activation of ATF6 and an ATF6 DNA binding site by the endoplasmic reticulum stress response. J. Biol. Chem. 275, 27013–27020 (2000).
Acknowledgements
This study was supported in part by US National Center for Research Resources (NCRR) grant M01-RR00425 to the Cedars-Sinai General Research Center Genotyping core; US National Institutes of Health/NIDDK grant P01-DK046763; Diabetes Endocrinology Research Center grant DK063491; Cedars-Sinai Medical Center Inflammatory Bowel Disease Research Funds. Additional funding was provided by grants DK76984 (M.C.D.) and DK084554 (M.C.D. and D.P.B.M.). Cardiovascular Health Study research reported in this article was supported by contract numbers N01-HC-85079 through N01-HC-85086, N01-HC-35129, N01 HC-15103, N01 HC-55222, N01-HC-75150 and N01-HC-45133; grant numbers U01 HL080295 and R01 HL087652 from the US National Heart, Lung, and Blood Institute; and additional contribution from the US National Institute of Neurological Disorders and Stroke. A full list of principal Cardiovascular Health Study investigators and institutions can be found at http://www.chs-nhlbi.org/pi.htm. A.G. is supported by the Crohn's and Colitis Foundation of America. R.J.X. and M.J.D. are supported by grants DK83756, DK086502 and DK043351 (NIDDK).
The NIDDK IBD Genetics Consortium is funded by the following grants: DK062431 (S.R.B.), DK062422 (J.H.C.), DK062420 (R.H.D.), DK062432 (J.D.R.), DK062423 (M.S.S.), DK062413 (D.P.B.M.) and DK062429 (J.H.C.). J.H.C. is also funded by Bohmfalk Funds for Medical Research, Burroughs Wellcome Medical Foundation and the Crohn's and Colitis Foundation of America. J.D.R. is also funded by grants from the US National Institute of Allergy and Infectious Diseases (AI065687; AI067152) and from the US National Institute of Diabetes and Digestive and Kidney Diseases (DK064869).
Activities in Sweden were supported by the Swedish Society of Medicine, the Bengt Ihre Foundation, the Karolinska Institutet, the Swedish National Program for IBD Genetics, the Swedish Organization for IBD, the Swedish Medical Research Council, the Soderbergh Foundation and the Swedish Cancer Foundation. Support for genotyping and genetic data analysis was provided by the Singapore National Cancer Centre, Singapore General Hospital and the Singapore Millennium Foundation (to S.P.) and the Agency for Science Technology and Research (A*STAR), Singapore (to M.L.H. and M.S.). Genotyping and DNA handling at the Genome Institute of Singapore were performed by W.Y. Meah, K.K. Heng, H.B. Toh, X. Lin, S. Rajaram, D. Tan and C.H. Wong. We are grateful to the funders and investigators of the Epidemiological Investigation of Rheumatoid Arthritis for providing genotype data from healthy Swedish individuals.
Author information
Authors and Affiliations
Consortia
Contributions
D.P.B.M., M.J.D., R.J.X., J.D.R., J.H.C., P.G., R.H.D. and M.S. participated in the study design and conception. D.P.B.M., A.G., M.J.D., R.J.X., J.D.R. and M.S. wrote the manuscript with contributions from R.H.D. and J.I.R. D.P.B.M., L.T., K.D.T., C.Li, C.B., P.R.F., M.C., M.D., J.H., M.L.H., M.L., L.P., A.A., E.C., A.L., O.P., E.-J.B., C.D., D.W.H., D.J.d.J., P.C.S., R.K.W., Y.S., M.S.S., J.H.C., S.R.B., L.P.S., R.H.D., M.C.D., N.L.G., T.H., A.I., G.Y.M., D.S.S., E.A.V., S.R.T., V.A., C.W. and S.P. performed patient diagnosis, patient enrollment and collection of clinical data. Replication genotyping was performed by C. Lagacé, C.R.S. and C.B. in the laboratory of J.D.R. Expression analysis, immunohistochemistry and shRNA studies were designed by A.G. and R.J.X. and performed by C. Li and A.G. M.J.D., J.E., B.M.N., K.R., J.W., J.D.R., P.G., T.G. and R.T.H.O. provided statistical analyses. All authors contributed to the final paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK).
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–5 and Supplementary Figures 1 and 2 (PDF 2209 kb)
Rights and permissions
About this article
Cite this article
McGovern, D., Gardet, A., Törkvist, L. et al. Genome-wide association identifies multiple ulcerative colitis susceptibility loci. Nat Genet 42, 332–337 (2010). https://doi.org/10.1038/ng.549
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.549