Abstract
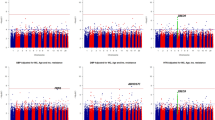
Numerous genetic loci have been associated with systolic blood pressure (SBP) and diastolic blood pressure (DBP) in Europeans1,2,3. We now report genome-wide association studies of pulse pressure (PP) and mean arterial pressure (MAP). In discovery (N = 74,064) and follow-up studies (N = 48,607), we identified at genome-wide significance (P = 2.7 × 10−8 to P = 2.3 × 10−13) four new PP loci (at 4q12 near CHIC2, 7q22.3 near PIK3CG, 8q24.12 in NOV and 11q24.3 near ADAMTS8), two new MAP loci (3p21.31 in MAP4 and 10q25.3 near ADRB1) and one locus associated with both of these traits (2q24.3 near FIGN) that has also recently been associated with SBP in east Asians. For three of the new PP loci, the estimated effect for SBP was opposite of that for DBP, in contrast to the majority of common SBP- and DBP-associated variants, which show concordant effects on both traits. These findings suggest new genetic pathways underlying blood pressure variation, some of which may differentially influence SBP and DBP.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Ehret, G. et al. Genetic variants in novel pathways influence blood pressure and cardiovascular disease risk. Nature advance online publication, doi: 10.1038/nature10405 (11 September 2011).
Levy, D. et al. Genome-wide association study of blood pressure and hypertension. Nat. Genet. 41, 677–687 (2009).
Newton-Cheh, C. et al. Genome-wide association study identifies eight loci associated with blood pressure. Nat. Genet. 41, 666–676 (2009).
Lawes, C.M. et al. Blood pressure and the global burden of disease 2000. Part II: estimates of attributable burden. J. Hypertens. 24, 423–430 (2006).
Rose, G. Strategies of prevention: the individual and the population. in Coronary Heart Disease Epidemiology: from Aetiology to Public Health (ed. Marmot M, E.P.) 631–41 (Oxford University Press, Oxford, UK, 2005).
Domanski, M.J. et al. Independent prognostic information provided by sphygmomanometrically determined pulse pressure and mean arterial pressure in patients with left ventricular dysfunction. J. Am. Coll. Cardiol. 33, 951–958 (1999).
Domanski, M. et al. Pulse pressure and cardiovascular disease-related mortality: follow-up study of the Multiple Risk Factor Intervention Trial (MRFIT). J. Am. Med. Assoc. 287, 2677–2683 (2002).
Franklin, S.S. et al. Single versus combined blood pressure components and risk for cardiovascular disease: the Framingham Heart Study. Circulation 119, 243–250 (2009).
Lewington, S., Clarke, R., Qizilbash, N., Peto, R. & Collins, R. Age-specific relevance of usual blood pressure to vascular mortality: a meta-analysis of individual data for one million adults in 61 prospective studies. Lancet 360, 1903–1913 (2002).
Devlin, B. & Roeder, K. Genomic control for association studies. Biometrics 55, 997–1004 (1999).
Kato, N. et al. Meta-analysis of genome-wide association studies identifies common variants associated with blood pressure variation in east Asians. Nat. Genet. 43, 531–538 (2011).
Sesso, H.D. et al. Systolic and diastolic blood pressure, pulse pressure, and mean arterial pressure as predictors of cardiovascular disease risk in men. Hypertension 36, 801–807 (2000).
Darne, B., Girerd, X., Safar, M., Cambien, F. & Guize, L. Pulsatile versus steady component of blood pressure: a cross-sectional analysis and a prospective analysis on cardiovascular mortality. Hypertension 13, 392–400 (1989).
Blacher, J. & Safar, M.E. Large-artery stiffness, hypertension and cardiovascular risk in older patients. Nat. Clin. Pract. Cardiovasc. Med. 2, 450–455 (2005).
Dart, A.M. & Kingwell, B.A. Pulse pressure–a review of mechanisms and clinical relevance. J. Am. Coll. Cardiol. 37, 975–984 (2001).
Johnson, A.D. et al. Genome-wide meta-analyses identifies seven loci associated with platelet aggregation in response to agonists. Nat. Genet. 42, 608–613 (2010).
Soranzo, N. et al. A novel variant on chromosome 7q22.3 associated with mean platelet volume, counts, and function. Blood 113, 3831–3837 (2009).
Soranzo, N. et al. A genome-wide meta-analysis identifies 22 loci associated with eight hematological parameters in the HaemGen consortium. Nat. Genet. 41, 1182–1190 (2009).
Oudit, G.Y. et al. Phosphoinositide 3-kinase γ-deficient mice are protected from isoproterenol-induced heart failure. Circulation 108, 2147–2152 (2003).
Perrino, C. et al. Dynamic regulation of phosphoinositide 3-kinase-γ activity and β-adrenergic receptor trafficking in end-stage human heart failure. Circulation 116, 2571–2579 (2007).
Wågsäter, D. et al. ADAMTS-4 and -8 are inflammatory regulated enzymes expressed in macrophage-rich areas of human atherosclerotic plaques. Atherosclerosis 196, 514–522 (2008).
Ellis, P.D., Chen, Q., Barker, P.J., Metcalfe, J.C. & Kemp, P.R. Nov gene encodes adhesion factor for vascular smooth muscle cells and is dynamically regulated in response to vascular injury. Arterioscler. Thromb. Vasc. Biol. 20, 1912–1919 (2000).
Shimoyama, T. et al. CCN3 inhibits neointimal hyperplasia through modulation of smooth muscle cell growth and migration. Arterioscler. Thromb. Vasc. Biol. 30, 675–682 (2010).
Heath, E. et al. Abnormal skeletal and cardiac development, cardiomyopathy, muscle atrophy and cataracts in mice with a targeted disruption of the Nov (Ccn3) gene. BMC Dev. Biol. 8, 18 (2008).
Cools, J. et al. Fusion of a novel gene, BTL, to ETV6 in acute myeloid leukemias with a t(4;12)(q11-q12;p13). Blood 94, 1820–1824 (1999).
Kamatani, Y. et al. Genome-wide association study of hematological and biochemical traits in a Japanese population. Nat. Genet. 42, 210–215 (2010).
Dorn, G.W. II. Adrenergic signaling polymorphisms and their impact on cardiovascular disease. Physiol. Rev. 90, 1013–1062 (2010).
Kitsios, G.D. & Zintzaras, E. Synopsis and data synthesis of genetic association studies in hypertension for the adrenergic receptor family genes: the CUMAGAS-HYPERT database. Am. J. Hypertens. 23, 305–313 (2010).
Rohrer, D.K., Chruscinski, A., Schauble, E.H., Bernstein, D. & Kobilka, B.K. Cardiovascular and metabolic alterations in mice lacking both β1- and β2-adrenergic receptors. J. Biol. Chem. 274, 16701–16708 (1999).
Cheng, G., Qiao, F., Gallien, T.N., Kuppuswamy, D. & Cooper, G. IV. Inhibition of β-adrenergic receptor trafficking in adult cardiocytes by MAP4 decoration of microtubules. Am. J. Physiol. Heart Circ. Physiol. 288, H1193–H1202 (2005).
Li, Y. & Abecasis, G.R. Mach 1.0: Rapid haplotype reconstruction and missing genotype inference. Am. J. Hum. Genet. S79, 2290 (2006).
Marchini, J., Howie, B., Myers, S., McVean, G. & Donnelly, P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nat. Genet. 39, 906–913 (2007).
Servin, B. & Stephens, M. Imputation-based analysis of association studies: candidate regions and quantitative traits. PLoS Genet. 3, e114 (2007).
Acknowledgements
A number of the participating studies and authors are members of the CHARGE and Global BPgen consortia. Many funding mechanisms by the US National Institutes of Health and National Heart, Lung, and Blood Institute, European and private funding agencies contributed to this work, and a full list of acknowledgements is provided in the Supplementary Note.
Author information
Authors and Affiliations
Consortia
Contributions
ICBP-GWAS PP and MAP Working and Writing Sub-Group (alphabetical order): M.J.C., P.E. (co-chair), T.J., P.B.M., P.F.O., M.D.T. (co-chair), C.M.v.D. (co-chair), G.C.V., L.V.W. ICBP-GWAS Steering Committee (alphabetical order): G.R.A., M. Bochud, M. Boehnke, M.J.C. (co-chair), A.C., G.B.E., P.E., T.B.H., M.-R.J., A.D.J., T.J., M.G.L., L.L., D.L. (co-chair), P.B.M.(co-chair), C.N.-C. (co-chair), B.M.P., K.M.R., A.V.S., M.D.T., C.M.v.D., G.C.V. Analysis: L.V.W., G.C.V., P.F.O., T.J. Expression analyses: V.E., P.H., A.D.J., D.L., J.H.L., C.P.N., A. Plump, P.A.C.'t H., K.W.v.D. Cohort contributions (alphabetical order): Study concept/design: AGES: T.A., V.G., T.B.H., L.L., A.V.S. AortaGen Consortium: G.F.M. ARIC: E.B., A.C., S.K.G. ASPS: H. Schmidt, R.S. BLSA: L.F. B58C-T1DGC: D.P.S. B58C-WTCCC: D.P.S. BHS: L.J.P. CARDIoGRAM Consortium: N.J.S. C4D Consortium: R. Clarke, CHS: J.C.B., N.L.G., B.M.P., K.M.R., K.D.T. CHARGE Consortium Heart Failure Working Group: N.L.S. CoLaus: V.M., P. Vollenweider, G. Waeber CROATIA-Korcula: C.H. CROATIA-Split: M. Boban, I.R. CROATIA-Vis: A.F.W. DeCode Genetics: H.H., K.S., G.T., U.T. DGI controls: D.A., L.G., C.N.-C. ENGAGE: J.E., I.R.K. EGCUT: H.A., A.M. EPIC: K.-T.K. ERF: B.A.O. Fenland: N.J.W. FUSION: M. Boehnke, F.S.C., R.N.B., J.T. INGI CARL: A.P.d'A., P. Gasparini INGI-FVG: A.P.d'A., P. Gasparini INCHIANTI: S. Bandinelli., Y.M. KORA S3: C.G., M. Laan, E.O. KORA F4: T.M., H.-E.W. LifeLines: R.P.S., M.M.v.d.K. LOLIPOP: J.C.C., P.E., J.S.K. LBC1921/LBC1936: I.J.D., J.M.S. MICROS: A. Pfeufer MESA: G.L.B., X.G., W.P. MIGen controls: O.M., C.J.O., V.S., D. Siscovick NESDA: B.W.P., H. Snieder NEURO-CHARGE Consortium: M. Breteler, M. Fornage NFBC1966: M.-R.J., P.Z. NSPHS: U.B.G. NTR: D.I.B., E.J.C.d.G. ORCADES: H.C., J.F.W. PROCARDIS controls: M. Farrall, A. Hamsten, J.F.P., H.W. PROSPER/PHASE: B.B., J.W.J., D. Stott RSI/RSII/RSIII: A. Hofman, C.M.v.D., J.C.M.W. SardiNIA: G.R.A., M.U. SHIP: M.D., H.K.K., R.R., U.V., H.V. SUVIMAX: P. Galan, S. Hercberg, P.M. TwinsUK: T.D.S. WGHS: P.M.R. YFS: M.K., T.L., O.T.R., J.V. Phenotype data acquisition and quality control: AGES: T.A., V.G., T.B.H., L.L. ARIC: A.C., S.K.G., A.C.M., D.C.R. ASPS: M. Loitfelder, R.S. BLSA: S.S.N. B58C-T1DGC: D.P.S. B58C-WTCCC: D.P.S. BHS: J.P.B., J.H. C4D Consortium: R. Clarke, J.C.H. CHS: B.M.P. CoLaus: M. Bochud, V.M., P. Vollenweider CROATIA-Korcula: C.H., O.P. CROATIA-Split: M. Boban, I.R. DGI controls: L.G., C.N.-C. EGCUT: H.A., A.K., A.M., M.-L.T. EPIC: N.J.W. Fenland: N.J.W. FHS: S.-J.H., M.G.L., D.L., R.S.V., T.J.W. FUSION: J.T. INGI CARL: A.F., F.F., P. Gasparini, S.U. INGI FVG: A.F., F.F., P. Gasparini, S.U. INGI-Val Borbera: C. Masciullo, C.S., D.T. INCHIANTI: A.M.C. KORA S3: C.G. KORA F4: A.D. LifeLines: M.M.v.d.K. LOLIPOP: J.C.C., J.S.K., J.S. LBC1921/LBC1936: I.J.D., L.M.L., J.M.S. MICROS: M. Facheris, A. Pfeufer MESA: G.L.B., X.G., W.P. MIGen controls: G.L., O.M., C.J.O., V.S., D. Siscovick NESDA: X. Lu, I.M.N., B.W.P., H. Snieder NEURO-CHARGE Consortium: M. Breteler, S.D., A.L.D., M. Fornage NFBC1966: P.E., M.-R.J., J. Laitinen, A. Pouta, P.Z. NSPHS: U.B.G. NTR: D.I.B., E.J.C.d.G., G. Willemsen ORCADES: S.H.W., J.F.W. PROCARDIS controls: J.F.P. PROSPER/PHASE: D. Stott, S.T. RSI/RSII/RSIII: F.U.S.M.-R., E.J.G.S., C.M.v.D., G.C.V., J.C.M.W. SardiNIA: M.O., M.U. SHIP: M.D., R.R., H.V. SUVIMAX: P. Galan, M. Lathrop TwinsUK: T.D.S. WGHS: P.M.R. YFS: M.K., T.L., O.T.R., J.V. Genotype data acquisition and quality control: AGES: A.V.S. ARIC: A.C., G.B.E., S.K.G., A.C.M., D.C.R., G.S. ASPS: P. Gider, H. Schmidt, M.Z. BLSA: D. Hernandez B58C-T1DGC: S. Heath, W.L.M. B58C-WTCCC: W.L.M. BHS: J.P.B., R.J.W. C4D Consortium: J.C.H., H.O. CHS: J.C.B., N.L.G., K.D.T. CoLaus: V.M., P. Vollenweider CROATIA-Korcula: C.H., O.P. CROATIA-Split: I.R. CROATIA-Vis: V.V. DGI controls: D.A., B.F.V. EGCUT: T.E., T.H. EPIC: N.J.W. Fenland: R.J.F.L., J. Luan, N.J.W. FHS: S.-J.H., M.G.L. FUSION: F.S.C. INGI CARL: A.P.d'A. INGI FVG: A.P.d'A. INGI Val Borbera: C. Masciullo, C.S., D.T. INCHIANTI: A.S. KORA S3: C.G., M. Laan, E.O. KORA F4: T.M., H.-E.W. LifeLines: B.Z.A. LOLIPOP: J.C.C., J.S.K., J.S., W.Z. LBC1921/LBC1936: G.D., I.J.D. MICROS: I.P. MESA: G.L.B., Y.-D.I.C., X.G. MIGen controls: G.L., O.M., C.J.O., V.S., D.S. NESDA: J.F., X. Lu, I.M.N., B.W.P., H. Snieder NFBC1966: P.E., M.-R.J., J. Laitinen, P.Z. NTR: D.I.B., E.J.C.d.G., J.-J.H., G. Willemsen ORCADES: H.C., J.F.W. PROCARDIS controls: A.G., J.F.P. PROSPER/PHASE: S.T. RSI/RSII/RSIII: F.R., A.G.U. SardiNIA: G.R.A. SHIP: H.K.K., U.V., H.V. SUVIMAX: S. Heath, M. Lathrop TwinsUK: M.M., S.-Y.S., N.S., F.Z. WGHS: D.I.C., A.N.P. YFS: T.L., O.T.R. Data analysis: AGES: T.A., A.V.S. ARIC: A.C., G.B.E., A.C.M., V.P., D.C.R., G.S. ASPS: P. Gider, H. Schmidt, M.Z. BLSA: T.T. B58C-T1DGC: D.P.S. B58C-WTCCC: D. Hadley, D.P.S. BHS: A.W.M., L.J.P., R.J.W. C4D Consortium: J.C.H., H.O., CHS: J.C.B., N.L.G., K.M.R. CoLaus: J.S.B., S. Bergmann, M. Bochud, T.J. CROATIA-Korcula: C.H., O.P. CROATIA-Split: C.H. CROATIA-Vis: V.V. DGI controls: P.A., C.N.-C., B.F.V. EchoGen Consortium: J.F.F. EGCUT: T.E., T.H. ENGAGE: M.P. EPIC: I.B., R.J.F.L., N.J.W., J.H.Z. ERF: A.C.J.W.J., Y.A. Fenland: R.J.F.L., J. Luan FHS: S.-J.H., M.G.L. FUSION: A.U.J. INGI CARL: N.P. INGI FVG: N.P. INGI Val Borbera: T.C., G.P., C.S., D.T. KORA S3: S.E., S.S. KORA F4: B.K. LifeLines: B.Z.A. LOLIPOP: J.C.C., J.S.K., X. Li, J.S., W.Z. LBC1921/LBC1936: L.M.L. MICROS: F.D.G.M. MESA: Y-D.I.C., X.G., W.P. MIGen controls: G.L. NESDA: J.F., X. Lu NEURO-CHARGE Consortium: S.D., A.L.D., M. Fornage NFBC1966: P.F.O. NSPHS:W.I. NTR: J.-J.H. ORCADES: P.N., S.H.W., J.F.W. PROCARDIS controls: M. Farrall, A.G., J.F.P. PROSPER/PHASE: J.W.J., S.T. RSI/RSII/RSIII: N.A., S.K., C.M.v.D., G.C.V. SardiNIA: J.L.B.-G. SHIP: U.V. SUVIMAX: T.J., P.M. TwinsUK: N.S., F.Z. WGHS: D.I.C., L.M.R., YFS: T.L., O.T.R.
Corresponding authors
Ethics declarations
Competing interests
A.C. is managed by Johns Hopkins Medicine. I.B. and spouse own stock in Incyte Ltd and GlaxoSmithKline. A.N.P. is an employee of Amgen. G.F.M. is owner of Cardiovascular Engineering, Inc, a company that designs and manufactures devices that measure vascular stiffness. The company uses these devices in clinical studies that evaluate the effects of diseases and interventions on vascular stiffness. V.M. is an employee of GlaxoSmithKline plc. A. Plump is an employee of Merck and Co, Inc.
Additional information
A full list of members is provided in the Supplementary Note.
A full list of members is provided in the Supplementary Note.
A full list of members is provided in the Supplementary Note.
A full list of members is provided in the Supplementary Note.
A full list of members is provided in the Supplementary Note.
A full list of members is provided in the Supplementary Note.
A full list of members is provided in the Supplementary Note.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4, Supplementary Tables 1–3 and Supplementary Note. (PDF 3579 kb)
Rights and permissions
About this article
Cite this article
Wain, L., Verwoert, G., O'Reilly, P. et al. Genome-wide association study identifies six new loci influencing pulse pressure and mean arterial pressure. Nat Genet 43, 1005–1011 (2011). https://doi.org/10.1038/ng.922
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.922
This article is cited by
-
Epigenome-wide association study in Chinese monozygotic twins identifies DNA methylation loci associated with blood pressure
Clinical Epigenetics (2023)
-
Polymorphisms of microRNAs are associated with salt sensitivity in a Han Chinese population: the EpiSS study
Journal of Human Hypertension (2022)
-
Expression of cardiovascular-related microRNAs is altered in L-arginine:glycine amidinotransferase deficient mice
Scientific Reports (2022)
-
Association of Circulating Biomarkers of lnc-IGSF3-1:1, SCOC-AS1, and SLC8A1-AS1 with Salt Sensitivity of Blood Pressure in Chinese Population
Journal of Cardiovascular Translational Research (2022)
-
Arterial Stiffness and the Canonical WNT/β-catenin Pathway
Current Hypertension Reports (2022)