Abstract
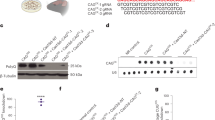
Inactivation of the mouse homologue of the Huntington disease gene (Hdh) results in early embryonic lethality. To investigate the normal function of Hdh in the adult and to evaluate current models for Huntington disease (HD), we have used the Cre/loxP site-specific recombination strategy to inactivate Hdh expression in the forebrain and testis, resulting in a progressive degenerative neuronal phenotype and sterility. On the basis of these results, we propose that huntingtin is required for neuronal function and survival in the brain and that a loss-of-function mechanism may contribute to HD pathogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Reddy, P.H., Williams, M. & Tagle, D.A. Recent advances in understanding the pathogenesis of Huntington's disease. Trends Neurosci. 22, 248–255 (1999).
DiFiglia, M. et al. Aggregation of huntingtin in neuronal intranuclear inclusions and dystrophic neurites in brain. Science 277, 1990–1993 (1997).
Li, S.H. et al. Huntington's disease gene (IT15) is widely expressed in human and rat tissues. Neuron 11, 985–993 (1993).
Strong, T.V. et al. Widespread expression of the human and rat Huntington's disease gene in brain and nonneural tissues. Nature Genet. 5, 259–265 (1993).
Sharp, A.H. et al. Widespread expression of Huntington's disease gene (IT15) protein product. Neuron 14, 1065–1074 (1995).
DiFiglia, M. et al. Huntingtin is a cytoplasmic protein associated with vesicles in human and rat brain neurons. Neuron 14, 1075–1081 (1995).
Gutekunst, C.A. et al. Identification and localization of huntingtin in brain and human lymphoblastoid cell lines with anti-fusion protein antibodies. Proc. Natl Acad. Sci. USA 92, 8710–8714 (1995).
Ferrante, R.J. et al. Heterogeneous topographic and cellular distribution of huntingtin expression in the normal human neostriatum. J. Neurosci. 17, 3052–3063 (1997).
Velier, J. et al. Wild-type and mutant huntingtins function in vesicle trafficking in the secretory and endocytic pathways. Exp. Neurol. 152, 34–40 (1998).
MacDonald, M.E. & Gusella, J.F. Huntington's disease: translating a CAG repeat into a pathogenic mechanism. Curr. Opin. Neurobiol. 6, 638–643 (1996).
Duyao, M.P. et al. Inactivation of the mouse Huntington's disease gene homolog Hdh. Science 269, 407–410 (1995).
Zeitlin, S., Liu, J.P., Chapman, D.L., Papaioannou, V.E. & Efstratiadis, A. Increased apoptosis and early embryonic lethality in mice nullizygous for the Huntington's disease gene homologue. Nature Genet. 11, 155–163 (1995).
Nasir, J. et al. Targeted disruption of the Huntington's disease gene results in embryonic lethality and behavioral and morphological changes in heterozygotes. Cell 81, 811–823 (1995).
O'Kusky, J.R., Nasir, J., Cicchetti, F., Parent, A. & Hayden, M.R. Neuronal degeneration in the basal ganglia and loss of pallido-subthalamic synapses in mice with targeted disruption of the Huntington's disease gene. Brain Res. 818, 468–479 (1999).
Sauer, B. & Henderson, N. Site-specific DNA recombination in mammalian cells by the Cre recombinase of bacteriophage P1. Proc. Natl Acad. Sci. USA 85, 5166–5170 (1988).
Mayford, M., Wang, J., Kandel, E.R. & O'Dell, T.J. CaMKII regulates the frequency-response function of hippocampal synapses for the production of both LTD and LTP. Cell 81, 891–904 (1995).
Tsien, J.Z. et al. Subregion- and cell type-restricted gene knockout in mouse brain. Cell 87, 1317–1326 (1996).
van den Akker, E. et al. Targeted inactivation of Hoxb8 affects survival of a spinal ganglion and causes aberrant limb reflexes. Mech. Dev. 89, 103–114 (1999).
Lalonde, R. Motor abnormalities in weaver mutant mice. Exp. Brain Res. 65, 479–481 (1987).
Mangiarini, L. et al. Exon 1 of the HD gene with an expanded CAG repeat is sufficient to cause a progressive neurological phenotype in transgenic mice. Cell 87, 493–506 (1996).
Matsuoka, Y., Kitamura, Y., Tooyama, I., Kimura, H. & Taniguchi, T. In vivo hypoxia-induced neuronal damage with an enhancement of neuronal nitric oxide synthase immunoreactivity in hippocampus. Exp. Neurol. 146, 57–66 (1997).
Santarosa, R. et al. Hyperplasia and apoptosis. Opposing cellular processes that regulate the response of the rabbit bladder to transient outlet obstruction. Lab. Invest. 70, 503–510 (1994).
Schmued, L.C., Albertson, C. & Slikker, W. Jr Fluoro-Jade: a novel fluorochrome for the sensitive and reliable histochemical localization of neuronal degeneration. Brain Res. 751, 37–46 (1997).
Bordelon, Y.M. & Chesselet, M.-F. Early effects of intrastriatal injections of quinolinic acid on microtubule-associated protein-2 and neuropeptides in rat basal ganglia. Neuroscience 93, 843–853 (1999).
Sibilia, M., Steinbach, J.P., Stingl, L., Aguzzi, A. & Wagner, E.F. A strain-independent postnatal neurodegeneration in mice lacking the EGF receptor. EMBO J. 17, 719–731 (1998).
Doherty, P., Cohen, J. & Walsh, F.S. Neurite outgrowth in response to transfected NCAM changes during development and is modulated by polysialic acid. Neuron 5, 209–219 (1990).
Hynes, R.O. & Lander, A.D. Contact and adhesive specificities in the associations, migrations and targeting of cells and axons. Cell 68, 303–322 (1992).
Haney, C.A. et al. Heterophilic binding of L1 on unmyelinated sensory axons mediates schwann cell adhesion and is required for axonal survival. J. Cell Biol. 146, 1173–1183 (1999).
Lagenaur, C. & Lemmon, V. An L1-like molecule, the 8D9 antigen, is a potent substrate for neurite extension. Proc. Natl Acad. Sci. USA 84, 7753–7747 (1987).
Landmesser, L., Schultz, K. & Rutishauser, U. Distinct roles for adhesion molecules during innervation of embryonic chick muscle. Dev. Biol. 130, 645–670 (1988).
Dragatsis, I., Efstratiadis, A. & Zeitlin, S. Mouse mutant embryos lacking huntingtin are rescued from lethality by wild-type extraembryonic tissues. Development 125, 1529–1539 (1998).
White, J.K. et al. Huntingtin is required for neurogenesis and is not impaired by the Huntington's disease CAG expansion. Nature Genet. 17, 404–410 (1997).
Metzler, M. et al. Life without huntingtin: normal differentiation into functional neurons. J. Neurochem. 72, 1009–1018 (1999).
Block-Galarza, J. et al. Fast transport and retrograde movement of huntingtin and HAP 1 in axons. Neuroreport 8, 2247–2251 (1997).
Sapp, E. et al. Axonal transport of N-terminal huntingtin suggests early pathology of corticostriatal projections in Huntington disease. J. Neuropathol. Exp. Neurol. 58, 165–173 (1999).
Narain, Y., Wyttenbach, A., Rankin, J., Furlong, R.A. & Rubinsztein, D.C. A molecular investigation of true dominance in Huntington's disease. J. Med. Genet. 36, 739–746 (1999).
Wheeler, V.C. et al. Long glutamine tracts cause nuclear localization of a novel form of huntingtin in medium spiny striatal neurons in HdhQ92 and HdhQ111 knock-in mice. Hum. Mol. Genet. 9, 503–513 (2000).
Rich, T. et al. Disassembly of nuclear inclusions in the dividing cell–a novel insight into neurodegeneration. Hum. Mol. Genet. 8, 2451–2459 (1999).
Dragatsis, I., Dietrich, P. & Zeitlin, S. Expression of the Huntingtin-associated protein 1 gene in the developing and adult mouse. Neurosci. Lett. 282, 37–40 (2000).
Sittler, A. et al. SH3GL3 associates with the Huntingtin exon 1 protein and promotes the formation of polygln-containing protein aggregates. Mol. Cell 2, 427–436 (1998).
Sathasivam, K. et al. Formation of polyglutamine inclusions in non-CNS tissue. Hum. Mol. Genet. 8, 813–822 (1999).
Levine, M.S. et al. Enhanced sensitivity to N-methyl-D-aspartate receptor activation in transgenic and knockin mouse models of Huntington's disease. J. Neurosci. Res. 58, 515–532 (1999).
Dragatsis, I. & Zeitlin, S. CaMKIIσ-cre transgene expression and recombination patterns in the mouse brain. Genesis 26, 133–135 (2000).
Persichetti, F. et al. Normal and expanded Huntington's disease gene alleles produce distinguishable proteins due to translation across the CAG repeat. Mol. Med. 1, 374–383 (1995).
Crawley, J.N. Behavioral phenotyping of transgenic and knockout mice: experimental design and evaluation of general health, sensory functions, motor abilities, and specific behavioral tests. Brain Res. 835, 18–26 (1999).
Rigamonti, D. et al. Wild-type huntingtin protects from apoptosis upstream of caspase-3. J. Neurosci. 20, 3705–3713 (2000).
Acknowledgements
We thank R.S. Fisher and P. Dietrich for histological examinations; M. Mendelsohn for blastocyst injections; K. Ratnam and T. Kolar for technical assistance; M. MacDonald, J. Goldman and T. Jessell for antibodies; A. Efstratiadis for helpful discussions and support; D. Wolgemuth and E.X. Wu for suggestions; and P. Dietrich, J.-P. Liu, A. Yamamoto and R. Hen for critical reading of the manuscript. We are grateful for the generous support provided to S.Z. and M.S.L. by the Hereditary Disease Foundation and the Hereditary Disease Foundation's Cure Huntington's Disease Initiative.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Dragatsis, I., Levine, M. & Zeitlin, S. Inactivation of Hdh in the brain and testis results in progressive neurodegeneration and sterility in mice. Nat Genet 26, 300–306 (2000). https://doi.org/10.1038/81593
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/81593
This article is cited by
-
Chemical engineering of therapeutic siRNAs for allele-specific gene silencing in Huntington’s disease models
Nature Communications (2022)
-
Antisense Oligonucleotide Therapy: From Design to the Huntington Disease Clinic
BioDrugs (2022)
-
RNA-seq analysis reveals significant transcriptome changes in huntingtin-null human neuroblastoma cells
BMC Medical Genomics (2021)
-
Huntingtin structure is orchestrated by HAP40 and shows a polyglutamine expansion-specific interaction with exon 1
Communications Biology (2021)
-
A high-throughput screening to identify small molecules that suppress huntingtin promoter activity or activate huntingtin-antisense promoter activity
Scientific Reports (2021)