Abstract
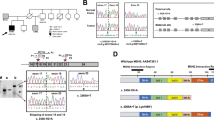
The mechanism by which germline mutations of DNA mismatch repair genes cause susceptibility to tumour formation is not yet understood. Studies in vitro indicate that heterozygosity for these mutations, unlike homozygosity, does not affect mismatch repair. Surprisingly, no loss of heterozygosity at the predisposing loci has so far been described in hereditary nonpolyposis colorectal cancers. Here, we show that loss of heterozygosity (LOH) of markers within or adjacent to the MLH1 gene on chromosome 3p occurs nonrandomly in tumours from members of families in which the disease phenotype cosegregates with MLH1. In every informative case, the loss affects the wild type allele. These results suggest that DNA mismatch repair genes resemble tumour suppressor genes in that two hits are required to cause a phenotypic effect.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Fishel, R. et al. The human mutator gene homolog MSH2 and its association with hereditary nonpolyposis colon cancer. Cell 75, 1027–1038 (1993).
Leach, F.S. et al. Mutations of a mutS homolog in hereditary nonpolyposis colorectal cancer. Cell 75, 1215–1225 (1993).
Papadopoulos, N. et al. Mutation of a mutL homolog in hereditary colon cancer. Science 263, 1625–1629 (1994).
Bronner, C.A. et al. Mutation in the DNA mismatch repair gene homologue hMLH1 is associated with hereditary non-polyposis colon cancer. Nature 368, 258–261 (1994).
Nicolaides, N et al. Mutations of two PMS homologues in hereditary nonpolyposis colon cancer. Nature 371, 75–80 (1994).
Lynch, H.T. et al. Genetics, natural history, tumour spectrum, and pathology of hereditary nonpolyposis colorectal cancer: an updated review. Gastroenterology 104, 1535–1549 (1993).
Peltomäki, P. et al. Genetic mapping of a locus predisposing to human colorectal cancer. Science 260, 810–812 (1993).
Aaltonen, L.A. et al. Clues to the pathogenesis of familial colorectal cancer. Science 260, 812–816 (1993).
Lindblom, A., Tannergård, P., Werelius, B. & Nordenskjöld, M. Genetic mapping of a second locus predisposing to hereditary non-polyposis colorectal cancer. Nature Genet. 5, 279–282 (1993).
Parsons, R. et al. Hypemnutability and mismatch repair deficiency in RER+ tumour cells. Cell 75, 1227–1236 (1993).
Ionov, Y., Peinado, M.A., Malkhosyan, S., Shibata, D. & Perucho, M. Ubiquitous somatic mutations in simple repeated sequences reveal a new mechanism for colonic carcinogenesis. Nature 363, 558–561 (1993).
Thibodeau, S.N., Bren, G. & Schaid, D. Microsatellite instability in cancer of the proximal colon. Science 260, 816–819 (1993).
Risinger, J.I. et al. Genetic instability of microsatellites in endometrial carcinoma. Cancer Res. 53, 5100–5103 (1993).
Aaltonen, L.A. et al. Replication errors in benign and malignant tumours from hereditary nonpolyposis colorectal cancer patients. Cancer Res. 54, 1645–1648 (1994).
Liu, B. et al. hMSH2 mutations in hereditary nonpolyposis colorectal cancer. Cancer Res. 54, 4590–4594 (1994).
Nyström-Lahti, M. et al. Mismatch repair genes on chromosomes 2p and 3p account for a major share of hereditary nonpolyposis colorectal cancer. Am. J. hum. Genet. 55, 659–665 (1994).
Nyström-Lahti, M. et al. Close linkage to chromosome 3p and conservation of ancestral founding haplotype in hereditary nonpolyposis colorectal cancer families. Proc. natn. Acad. Sci. U.S.A. 91, 6054–6058 (1994).
Mecklin, J-P, Järvinen, H.J. & Peltokallio, P. Cancer family syndrome. Genetic analysis of 22 Finnish kindreds. Gastroenterology 90, 328–333 (1986).
Mecklin, J-P. Frequency of hereditary colorectal carcinoma. Gastroenterology 93, 1021–1025 (1987).
Peltomäki, P. et al. Evidence supporting exclusion of the DCC gene and a portion of chromosome 18q as the locus for susceptibility to hereditary nonpolyposis colorectal carcinoma in five kindreds. Cancer Res. 51, 4135–4140 (1991).
Peltomäki, P. et al. Evidence that the MCC-APC gene region in 5q21 is not the site for susceptibility to hereditary nonpolyposis colorectal carcinoma. Cancer Res. 52, 4530–4533 (1992).
Ganly, P.S., Jarad, N., Rudd, R.M. & Rabbits, P.H. PCR-based RFLP analysis allows genotyping of the short arm of chromosome 3 in small biopsies from patients with lung cancer. Genomics 12, 221–228 (1992).
Knudson, A.G. All in the (cancer) family. Nature Genet. 5, 103–104 (1993).
Bodmer, W., Bishop, T. & Karran, P. Genetic steps in colorectal cancer. Nature Genet. 6, 217–219 (1994).
Jiricny, J. Colon cancer and DNA repair: have mismatches met their match?Trends Genet. 10, 164–168 (1994).
Kouri, M. Diploid predominance in hereditary nonpolyposis colorectal carcinoma evaluated by flow cytometry. Cancer 65, 1825–1829 (1990).
Lothe, R.A. et al. Genomic instability in colorectal cancer; relationship to clinicopathological variables and family history. Cancer Res. 53, 5849–5852 (1993).
Wu, C. et al. DNA alterations in cells from hereditary non-polyposis colorectal cancer patients. Oncogene 9, 991–994 (1994).
Knudson, A.G. Mutation and cancer: statistical study on retinoblastoma. Proc. natn. Acad. Sci. U.S.A. 68, 820–823 (1971).
Gyapay, G. et al. The 1993-1994 Généthon human genetic linkage map. Nature Genet. 7, 246–339 (1994).
Vogelstein, B. et al. Genetic alterations during colorectal tumour development. New Engl. J. Med. 319, 525–532 1988).
Sambrook, J., Fritsch, E.F. & Maniatis, T. Molecular cloning, a laboratory manual. (Cold Spring Harbor Laboratory Press/New York, 1989).
Isola, J., DeVries, S., Chu, L, Ghazvini, S. & Waldman, F. Analysis of changes in DNA sequence copy number by comparative genomic hybridization in archival paraffin-embedded tumour samples. Am. J. Pathol. (in the press).
Weissenbach, J. et al. Second generation linkage map of the human genome. Nature 359, 794–801 (1992).
Jones, M.H., Yamakawa, K. & Nakamura, Y. Isolation and characterization of 19 dinucleotide repeat polymorphisms on chromosome 3p. Hum. molec. Genet. 1, 131–133 (1992).
Spirio, L., Joslyn, G., Nelson, L., Leppert, M. & White, R. A CA repeat 30-70 kb downstream from the adenomatous polyposis gene. Nucl. Acids Res. 19, 6348 (1991).
Jones, M.H. & Nakamura, Y. Detection of loss of heterozygosity at the human TP53 locus using a dinucleotide repeat polymorphism. Genes Chrom.Cancer 5, 89–90 (1992).
Risinger, J.I. & Boyd, J. Dinucleotide repeat polymorphism in the human DCC gene at chromosome 18q21. Hum. molec. genet. 1, 657 (1992).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Hemminki, A., Peltomäki, P., Mecklin, JP. et al. Loss of the wild type MLH1 gene is a feature of hereditary nonpolyposis colorectal cancer. Nat Genet 8, 405–410 (1994). https://doi.org/10.1038/ng1294-405
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ng1294-405
This article is cited by
-
Lynch syndrome, molecular mechanisms and variant classification
British Journal of Cancer (2023)
-
Histomorphological and molecular genetic characterization of different intratumoral regions and matched metastatic lymph nodes of colorectal cancer with heterogenous mismatch repair protein expression
Journal of Cancer Research and Clinical Oncology (2023)
-
Association of a novel point mutation in MSH2 gene with familial multiple primary cancers
Journal of Hematology & Oncology (2017)
-
Opportunities for immunotherapy in microsatellite instable colorectal cancer
Cancer Immunology, Immunotherapy (2016)
-
Milestones of Lynch syndrome: 1895–2015
Nature Reviews Cancer (2015)