Abstract
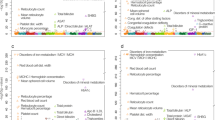
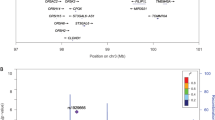
Chronic inflammation has a pathological role in many common diseases and is influenced by both genetic and environmental factors. Here we assess the role of genetic variation in selenoprotein S (SEPS1, also called SELS or SELENOS), a gene involved in stress response in the endoplasmic reticulum and inflammation control. After resequencing SEPS1, we genotyped 13 SNPs in 522 individuals from 92 families. As inflammation biomarkers, we measured plasma levels of IL-6, IL-1β and TNF-α. Bayesian quantitative trait nucleotide analysis identified associations between SEPS1 polymorphisms and all three proinflammatory cytokines. One promoter variant, −105G → A, showed strong evidence for an association with each cytokine (multivariate P = 0.0000002). Functional analysis of this polymorphism showed that the A variant significantly impaired SEPS1 expression after exposure to endoplasmic reticulum stress agents (P = 0.00006). Furthermore, suppression of SEPS1 by short interfering RNA in macrophage cells increased the release of IL-6 and TNF-α. To investigate further the significance of the observed associations, we genotyped −105G → A in 419 Mexican American individuals from 23 families for replication. This analysis confirmed a significant association with both TNF-α (P = 0.0049) and IL-1β (P = 0.0101). These results provide a direct mechanistic link between SEPS1 and the production of inflammatory cytokines and suggest that SEPS1 has a role in mediating inflammation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Walder, K. et al. Tanis: a link between type 2 diabetes and inflammation? Diabetes 51, 1859–1866 (2002).
Gao, Y. et al. Elevation in Tanis expression alters glucose metabolism and insulin sensitivity in H4IIE cells. Diabetes 52, 929–934 (2003).
Kryukov, G.V. et al. Characterization of mammalian selenoproteomes. Science 300, 1439–1443 (2003).
Ye, Y., Shibata, Y., Yun, C., Ron, D. & Rapoport, T.A. A membrane protein complex mediates retro-translocation from the ER lumen into the cytosol. Nature 429, 841–847 (2004).
Karlsson, H.K., Tsuchida, H., Lake, S., Koistinen, H.A. & Krook, A. Relationship between serum amyloid A level and Tanis/SelS mRNA expression in skeletal muscle and adipose tissue from healthy and type 2 diabetic subjects. Diabetes 53, 1424–1428 (2004).
Pahl, H.L. & Baeuerle, P.A. The ER-overload response: activation of NF-kappa B. Trends Biochem. Sci. 22, 63–67 (1997).
Zamani, M., Pociot, F., Raeymaekers, P., Nerup, J. & Cassiman, J.J. Linkage of type I diabetes to 15q26 (IDDM3) in the Danish population. Hum. Genet. 98, 491–496 (1996).
Field, L.L., Tobias, R. & Magnus, T. A locus on chromosome 15q26 (IDDM3) produces susceptibility to insulin-dependent diabetes mellitus. Nat. Genet. 8, 189–194 (1994).
Blacker, D. et al. Results of a high-resolution genome screen of 437 Alzheimer's disease families. Hum. Mol. Genet. 12, 23–32 (2003).
Susi, M., Holopainen, P., Mustalahti, K., Maki, M. & Partanen, J. Candidate gene region 15q26 and genetic susceptibility to coeliac disease in Finnish families. Scand. J. Gastroenterol. 36, 372–374 (2001).
Saadeddin, S.M., Habbab, M.A. & Ferns, G.A. Markers of inflammation and coronary artery disease. Med. Sci. Monit. 8, RA5–12 (2002).
Cheverud, J.M. A simple correction for multiple comparisons in interval mapping genome scans. Heredity 87, 52–58 (2001).
Abecasis, G.R., Cherny, S.S., Cookson, W.O. & Cardon, L.R. Merlin–rapid analysis of dense genetic maps using sparse gene flow trees. Nat. Genet. 30, 97–101 (2002).
Almasy, L. & Blangero, J. Multipoint quantitative-trait linkage analysis in general pedigrees. Am. J. Hum. Genet. 62, 1198–1211 (1998).
Abecasis, G.R., Cookson, W.O. & Cardon, L.R. Pedigree tests of transmission disequilibrium. Eur. J. Hum. Genet. 8, 545–551 (2000).
Blangero, J. et al. Quantitative trait nucleotide analysis using Bayesian model selection. Hum. Biol. (in the press).
Gao, Y. et al. Regulation of the selenoprotein SelS by glucose deprivation and endoplasmic reticulum stress - SelS is a novel glucose-regulated protein. FEBS Lett. 563, 185–190 (2004).
de Maat, M.P. et al. Genetic influence on inflammation variables in the elderly. Arterioscler. Thromb. Vasc. Biol. 24, 2168–2173 (2004).
Pantsulaia, I., Trofimov, S., Kobyliansky, E. & Livshits, G. Genetic and environmental influences on IL-6 and TNF-alpha plasma levels in apparently healthy general population. Cytokine 19, 138–146 (2002).
Williams-Blangero, S. et al. Genetic influences on plasma cytokine variation in a parasitized population. Hum. Biol. 76, 515–525 (2004).
Blangero, J., Williams, J.T. & Almasy, L. Novel family-based approaches to genetic risk in thrombosis. J. Thromb. Haemost. 1, 1391–1397 (2003).
Stengard, J.H. et al. Contributions of 18 additional DNA sequence variations in the gene encoding apolipoprotein E to explaining variation in quantitative measures of lipid metabolism. Am. J. Hum. Genet. 71, 501–517 (2002).
Altshuler, D. et al. The common PPARgamma Pro12Ala polymorphism is associated with decreased risk of type 2 diabetes. Nat. Genet. 26, 76–80 (2000).
Caspersen, C., Pedersen, P.S. & Treiman, M. The sarco/endoplasmic reticulum calcium-ATPase 2b is an endoplasmic reticulum stress-inducible protein. J. Biol. Chem. 275, 22363–22372 (2000).
Kokame, K., Kato, H. & Miyata, T. Identification of ERSE-II, a new cis-acting element responsible for the ATF6-dependent mammalian unfolded protein response. J. Biol. Chem. 276, 9199–9205 (2001).
Roy, B. & Lee, A.S. The mammalian endoplasmic reticulum stress response element consists of an evolutionarily conserved tripartite structure and interacts with a novel stress-inducible complex. Nucleic Acids Res. 27, 1437–1443 (1999).
Arthur, J.R. Selenium supplementation: does soil supplementation help and why? Proc. Nutr. Soc. 62, 393–397 (2003).
Ferencik, M. & Ebringer, L. Modulatory effects of selenium and zinc on the immune system. Folia Microbiol. (Praha) 48, 417–426 (2003).
Kissebah, A.H. et al. Quantitative trait loci on chromosomes 3 and 17 influence phenotypes of the metabolic syndrome. Proc. Natl. Acad. Sci. USA 97, 14478–14483 (2000).
MacCluer, J.W. et al. Genetics of atherosclerosis risk factors in Mexican Americans. Nutr. Rev. 57, S59–S65 (1999).
Buetow, K.H. et al. High-throughput development and characterization of a genomewide collection of gene-based single nucleotide polymorphism markers by chip-based matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Proc. Natl. Acad. Sci. USA 98, 581–584 (2001).
Phillips, P.C. From complex traits to complex alleles. Trends Genet. 15, 6–8 (1999).
Long, A.D., Lyman, R.F., Langley, C.H. & Mackay, T.F. Two sites in the Delta gene region contribute to naturally occurring variation in bristle number in Drosophila melanogaster. Genetics 149, 999–1017 (1998).
Kass, R.E. & Raftery, A.E. Bayes factors. J. Am. Stat. Assoc. 90, 773–795 (1995).
Raftery, A.E. Bayesian model selection in social research. in Sociological Methodology (ed. Marsden, P.V.) 111–195 (Blackwells, Oxford, 1995).
Acknowledgements
This work was supported by grants from the National Institutes of Health. Take Off Pounds Sensibly, Inc. provided funds for establishment of the family database and clinical phenotyping. This work was also supported by grants from the National Center for Research Resources to the General Clinical Research Centers at the Medical College of Wisconsin and the University of Texas Health Science Center San Antonio. The statistical genetics computer package, SOLAR, is supported by a grant from the National Institutes of Mental Health. The supercomputing facilities used for this work at the SBC Genetics Computing Center were supported in part by a gift from the SBC Foundation. Funds for resequencing, genotyping and functional and statistical analyses were provided by ChemGenex Pharmaceuticals Ltd., Australia.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Funds for resequencing, genotyping and functional and statistical analyses were provided by ChemGenex Pharmaceuticals, which has a patent on SELS in relation to its role in multiple diseases. G.R.C. and P.Z. have personal financial interests in ChemGenex. G.R.C. is also the Chief Executive Officer and Managing Director of ChemGenex. P.Z. and J.B. are members of the Scientific Advisory Board of ChemGenex. J.B. also serves as Senior Director of Human Genomics, and K.R.W. serves as Senior Director of Research and Development.
Supplementary information
Supplementary Table 1
Selenoprotein S genetic variation identified in the sample. (PDF 26 kb)
Supplementary Table 2
Distribution of relative pairs in the population sample of 522 individuals from 92 families. (PDF 15 kb)
Supplementary Table 3
siRNA primer sequences. (PDF 13 kb)
Rights and permissions
About this article
Cite this article
Curran, J., Jowett, J., Elliott, K. et al. Genetic variation in selenoprotein S influences inflammatory response. Nat Genet 37, 1234–1241 (2005). https://doi.org/10.1038/ng1655
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1655
This article is cited by
-
Effects of Selenoprotein S Knockdown on Endoplasmic Reticulum Stress in ATDC5 Cells and Gene Expression Profiles in Hypertrophic Chondrocytes
Biological Trace Element Research (2023)
-
Oxidative stress in Hashimoto’s thyroiditis: possible adjuvant therapies to attenuate deleterious effects
Molecular and Cellular Biochemistry (2023)
-
Calcium intake may explain the reduction of colorectal cancer odds by dietary selenium - a case-control study in Poland
BMC Nutrition (2022)
-
Identification of loci associated with susceptibility to bovine paratuberculosis and with the dysregulation of the MECOM, eEF1A2, and U1 spliceosomal RNA expression
Scientific Reports (2021)
-
A Mechanistic Link Between Selenium and Coronavirus Disease 2019 (COVID-19)
Current Nutrition Reports (2021)