Abstract
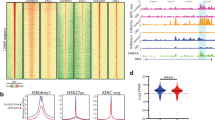
We demonstrate that the binding sites for highly conserved transcription factors vary extensively between human and mouse. We mapped the binding of four tissue-specific transcription factors (FOXA2, HNF1A, HNF4A and HNF6) to 4,000 orthologous gene pairs in hepatocytes purified from human and mouse livers. Despite the conserved function of these factors, from 41% to 89% of their binding events seem to be species specific. When the same protein binds the promoters of orthologous genes, approximately two-thirds of the binding sites do not align.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Bird, C.P., Stranger, B.E. & Dermitzakis, E.T. Curr. Opin. Genet. Dev. 16, 559–564 (2006).
Moses, A.M. et al. PLoS Comput. Biol. 2, e130 (2006).
Prabhakar, S., Noonan, J.P., Paabo, S. & Rubin, E.M. Science 314, 786 (2006).
King, M.C. & Wilson, A.C. Science 188, 107–116 (1975).
Boyer, L.A. et al. Cell 122, 947–956 (2005).
Loh, Y.H. et al. Nat. Genet. 38, 431–440 (2006).
Odom, D.T. et al. Mol. Syst. Biol. 2, 2006.0017 (2006).
Zaret, K.S. Nat. Rev. Genet. 3, 499–512 (2002).
Richert, L. et al. Drug Metab. Dispos. 34, 870–879 (2006).
Qi, Y. et al. Nat. Biotechnol. 24, 963–970 (2006).
Natarajan, A.T. & Darroudi, F. Mutagenesis 6, 399–403 (1991).
Macisaac, K.D. et al. Bioinformatics 22, 423–429 (2006).
Schwartz, S. et al. Genome Res. 13, 103–107 (2003).
Kim, S.H. et al. Diabetes 53, 1375–1384 (2004).
Guccione, E. et al. Nat. Cell Biol. 8, 764–770 (2006).
Acknowledgements
We are grateful to S. Strom and K. Dorko (University of Pittsburgh) for human liver samples (DK92310); T. DiCesare, S. Gupta, R. Kumar, J. Rodriguez and K. Walker for technical assistance and R. Young and G. Gerber for discussions. This work was supported by funding from US National Institutes of Health (grants DK68655 and DK70813 (D.T.O.) and DK076284 (R.D.D.)), Cancer Research UK (D.T.O.) and the Whitaker Foundation (E.F.).
Author information
Authors and Affiliations
Contributions
D.T.O., R.D.D. and E.F. designed experiments; R.D.D. designed the arrays; D.T.O., E.S.J., W.G. and C.M.C. performed experiments; R.D.D., T.W.D, K.D.M., P.A.R. and E.F. analyzed the data and D.T.O, R.D.D., D.K.G. and E.F. created the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Binding to known liver-specific genes. (PDF 198 kb)
Supplementary Fig. 2
Species-specific binding does not arise from technical limitations. (PDF 369 kb)
Supplementary Fig. 3
Distance from binding maxima to motif sequences. (PDF 109 kb)
Supplementary Table 1
Conservation of transcription factors and expression in human and mouse liver. (PDF 88 kb)
Supplementary Table 2
Species-specific binding does not arise from technical limitations. (PDF 266 kb)
Supplementary Table 3
Analysis of motif sequences and alignments. (PDF 583 kb)
Rights and permissions
About this article
Cite this article
Odom, D., Dowell, R., Jacobsen, E. et al. Tissue-specific transcriptional regulation has diverged significantly between human and mouse. Nat Genet 39, 730–732 (2007). https://doi.org/10.1038/ng2047
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng2047