Abstract
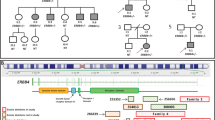
The epilepsies are a common, clinically heterogeneous group of disorders defined by recurrent unprovoked seizures1. Here we describe identification of the causative gene in autosomal-dominant partial epilepsy with auditory features (ADPEAF, MIM 600512), a rare form of idiopathic lateral temporal lobe epilepsy characterized by partial seizures with auditory disturbances2,3. We constructed a complete, 4.2-Mb physical map across the genetically implicated disease-gene region, identified 28 putative genes (Fig. 1) and resequenced all or part of 21 genes before identifying presumptive mutations in one copy of the leucine-rich, glioma-inactivated 1 gene (LGI1) in each of five families with ADPEAF. Previous studies have indicated that loss of both copies of LGI1 promotes glial tumor progression. We show that the expression pattern of mouse Lgi1 is predominantly neuronal and is consistent with the anatomic regions involved in temporal lobe epilepsy. Discovery of LGI1 as a cause of ADPEAF suggests new avenues for research on pathogenic mechanisms of idiopathic epilepsies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Hauser, W.A., Annegers, J.F. & Kurland, L.T. Prevalence of epilepsy in Rochester, Minnesota: 1940–1980. Epilepsia 32, 429–445 (1991).
Ottman, R. et al. Localization of a gene for partial epilepsy to chromosome 10q. Nature Genet. 10, 56–60 (1995).
Winawer, M.R., Ottman, R., Hauser, W.A. & Pedley, T.A. Autosomal dominant partial epilepsy with auditory features: defining the phenotype. Neurology 54, 2173–2176 (2000).
Poza, J.J. et al. Autosomal dominant lateral temporal epilepsy: clinical and genetic study of a large Basque pedigree linked to chromosome 10q. Ann. Neurol. 45, 182–188 (1999).
Michelucci, R. et al. Autosomal dominant partial epilepsy with auditory features: description of a new family. Epilepsia 41, 967–970 (2000).
Mautner, V.F., Lindenau, M., Gottesleben, A., Goetze, G. & Kluwe, L. Supporting evidence of a gene for partial epilepsy on 10q. Neurogenetics 3, 31–34 (2000).
Winawer, M.R. et al. Four new families with autosomal dominant partial epilepsy with auditory features: clinical description and linkage to chromosome 10q24. Epilepsia 43, 55–66 (2002).
Chernova, O.B., Somerville, R.P. & Cowell, J.K. A novel gene, LGI1, from 10q24 is rearranged and downregulated in malignant brain tumors. Oncogene 17, 2873–2881 (1998).
Somerville, R.P., Chernova, O., Liu, S., Shoshan, Y. & Cowell, J.K. Identification of the promoter, genomic structure, and mouse ortholog of LGI1. Mamm. Genome 11, 622–627 (2000).
Kobe, B. & Deisenhofer, J. The leucine-rich repeat: a versatile binding motif. Trends Biochem. Sci. 19, 415–421 (1994).
Kobe, B. & Deisenhofer, J. Proteins with leucine-rich repeats. Curr. Opin. Struct. Biol. 5, 409–416 (1995).
Chang, Z. et al. Molecular and genetic characterization of the Drosophila tartan gene. Dev. Biol. 160, 315–332 (1993).
Battye, R., Stevens, A., Perry, R.L. & Jacobs, J.R. Repellent signaling by Slit requires the leucine-rich repeats. J. Neurosci. 21, 4290–4298 (2001).
Wu, W. et al. Directional guidance of neuronal migration in the olfactory system by the protein Slit. Nature 400, 331–336 (1999).
Kidd, T., Bland, K.S. & Goodman, C.S. Slit is the midline repellent for the robo receptor in Drosophila. Cell 96, 785–794 (1999).
Meisler, M.H., Kearney, J., Ottman, R. & Escayg, A. Identification of epilepsy genes in human and mouse. Annu. Rev. Genet. 35, 567–588 (2001).
Skradski, S.L. et al. A novel gene causing a mendelian audiogenic mouse epilepsy. Neuron 31, 537–544 (2001).
Commission on Classification and Terminology, I.L.A.E. Proposal for revised classification of epilepsies and epileptic syndromes. Epilepsia 30, 389–399 (1989).
Aita, V.M. et al. A comprehensive linkage analysis of chromosome 21q22 supports prior evidence for a putative bipolar affective disorder locus. Am. J. Hum. Genet. 64, 210–217 (1999).
Goring, H.H. & Terwilliger, J.D. Linkage analysis in the presence of errors. III: Marker loci and their map as nuisance parameters. Am. J. Hum. Genet. 66, 1298–1309 (2000).
Kruglyak, L., Daly, M.J., Reeve-Daly, M.P. & Lander, E.S. Parametric and nonparametric linkage analysis: a unified multipoint approach. Am. J. Hum. Genet. 58, 1347–1363 (1996).
Gray, I.C., Nobile, C., Muresu, R., Ford, S. & Spurr, N.K. A 2.4-megabase physical map spanning the CYP2C gene cluster on chromosome 10q24. Genomics 28, 328–332 (1995).
Gray, I.C. et al. An integrated physical and genetic map spanning chromosome band 10q24. Genomics 43, 85–88 (1997).
Nobile, C. et al. A refined physical and EST map spanning 7.4 Mb of 10q24, a region involved in neurological disorders. Mamm. Genome. 9, 835–837 (1998).
Tatusova, T.A. & Madden, T.L. BLAST 2 Sequences, a new tool for comparing protein and nucleotide sequences. FEMS Microbiol. Lett. 174, 247–250 (1999).
Schaeren-Wiemers, N. & Gerfin-Moser, A. A single protocol to detect transcripts of various types and expression levels in neural tissue and cultured cells: in situ hybridization using digoxigenin-labelled cRNA probes. Histochemistry 100, 431–440 (1993).
Engelman, D.M., Steitz, T.A. & Goldman, A. Identifying nonpolar transbilayer helices in amino acid sequences of membrane proteins. Annu. Rev. Biophys. Biophys. Chem. 15, 321–353 (1986).
Juretic, D., Zucic, D., Lucic, B. & Trinajstic, N. Preference functions for prediction of membrane-buried helices in integral membrane proteins. Comput. Chem. 22, 279–294 (1998).
Milpetz, F., Argos, P. & Persson, B. TMAP: a new email and WWW service for membrane-protein structural predictions. Trends Biochem. Sci. 20, 204–205 (1995).
Shah, A.B. et al. Identification and analysis of mutations in the Wilson disease gene (ATP7B): population frequencies, genotype–phenotype correlation, and functional analyses. Am. J. Hum. Genet. 61, 317–328 (1997).
Acknowledgements
This work was supported by grants from the National Institutes of Health, National Institute of Neurological Disorders and Stroke and by funds from the Columbia Genome Center. We thank A. Efstratiadis, I. Dragatsis, I. Lipkin, M. Hornig and H. Scharfman for their critical discussions and helpful suggestions; J. Ju, A.K. Tong and C. Wang for timely technical assistance; P. McCabe, C.D. McNew and S.R. Resor for family referrals and W. Jimenez for assistance with database management. This research would not have been possible without the generous participation of the families described.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kalachikov, S., Evgrafov, O., Ross, B. et al. Mutations in LGI1 cause autosomal-dominant partial epilepsy with auditory features. Nat Genet 30, 335–341 (2002). https://doi.org/10.1038/ng832
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng832
This article is cited by
-
A patient-derived mutation of epilepsy-linked LGI1 increases seizure susceptibility through regulating Kv1.1
Cell & Bioscience (2023)
-
The LGI1 protein: molecular structure, physiological functions and disruption-related seizures
Cellular and Molecular Life Sciences (2022)
-
Long QT syndrome with potassium voltage-gated channel subfamily H member 2 gene mutation mimicking refractory epilepsy: case report
BMC Neurology (2021)
-
Skeletal muscle transcriptome in healthy aging
Nature Communications (2021)
-
Diagnostic Considerations in the Epilepsies—Testing Strategies, Test Type Advantages, and Limitations
Neurotherapeutics (2021)