Abstract
The microRNA miR-210 is a signature of hypoxia. We found robust increase in the abundance of miR-210 (>100-fold) in activated T cells, especially in the TH17 lineage of helper T cells. Hypoxia acted in synergy with stimulation via the T cell antigen receptor (TCR) and coreceptor CD28 to accelerate and increase Mir210 expression. Mir210 was directly regulated by HIF-1α, a key transcriptional regulator of TH17 polarization. Unexpectedly, we identified Hif1a as a target of miR-210, which suggested negative feedback by miR-210 in inhibiting HIF-1α expression. Deletion of Mir210 promoted TH17 differentiation under conditions of limited oxygen. In experimental colitis, miR-210 reduced the abundance of Hif1a transcripts and the proportion of cells that produced inflammatory cytokines and controlled disease severity. Our study identifies miR-210 as an important regulator of T cell differentiation in hypoxia, which can limit immunopathology.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Caldwell, C.C. et al. Differential effects of physiologically relevant hypoxic conditions on T lymphocyte development and effector functions. J. Immunol. 167, 6140–6149 (2001).
McNamee, E.N., Korns Johnson, D., Homann, D. & Clambey, E.T. Hypoxia and hypoxia-inducible factors as regulators of T cell development, differentiation, and function. Immunol. Res. 55, 58–70 (2013).
Ye, J. Emerging role of adipose tissue hypoxia in obesity and insulin resistance. Int. J. Obes. 33, 54–66 (2009).
Bedogni, B. et al. The hypoxic microenvironment of the skin contributes to Akt-mediated melanocyte transformation. Cancer Cell 8, 443–454 (2005).
Taylor, C.T. & Colgan, S.P. Hypoxia and gastrointestinal disease. J. Mol. Med. 85, 1295–1300 (2007).
Semenza, G.L. Hypoxia-inducible factor 1 (HIF-1) pathway. Sci. STKE 2007, cm8 (2007).
Wang, G.L., Jiang, B.H., Rue, E.A. & Semenza, G.L. Hypoxia-inducible factor 1 is a basic-helix-loop-helix-PAS heterodimer regulated by cellular O2 tension. Proc. Natl. Acad. Sci. USA 92, 5510–5514 (1995).
Maxwell, P.H. et al. The tumour suppressor protein VHL targets hypoxia-inducible factors for oxygen-dependent proteolysis. Nature 399, 271–275 (1999).
Jaakkola, P. et al. Targeting of HIF-α to the von Hippel-Lindau ubiquitylation complex by O2-regulated prolyl hydroxylation. Science 292, 468–472 (2001).
Bruning, U. et al. MicroRNA-155 promotes resolution of hypoxia-inducible factor 1α activity during prolonged hypoxia. Mol. Cell. Biol. 31, 4087–4096 (2011).
Korn, T., Bettelli, E., Oukka, M. & Kuchroo, V.K. IL-17 and Th17 cells. Annu. Rev. Immunol. 27, 485–517 (2009).
Weaver, C.T., Hatton, R.D., Mangan, P.R. & Harrington, L.E. IL-17 family cytokines and the expanding diversity of effector T cell lineages. Annu. Rev. Immunol. 25, 821–852 (2007).
Fife, B.T. et al. Interactions between PD-1 and PD-L1 promote tolerance by blocking the TCR-induced stop signal. Nat. Immunol. 10, 1185–1192 (2009).
Wu, S. et al. A human colonic commensal promotes colon tumorigenesis via activation of T helper type 17 T cell responses. Nat. Med. 15, 1016–1022 (2009).
Dang, E.V. et al. Control of TH17/Treg balance by hypoxia-inducible factor 1. Cell 146, 772–784 (2011).
Shi, L.Z. et al. HIF1alpha-dependent glycolytic pathway orchestrates a metabolic checkpoint for the differentiation of TH17 and Treg cells. J. Exp. Med. 208, 1367–1376 (2011).
Bartel, D.P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
Leung, A.K. & Sharp, P.A. MicroRNA functions in stress responses. Mol. Cell 40, 205–215 (2010).
Chan, Y.C., Banerjee, J., Choi, S.Y. & Sen, C.K. miR-210: the master hypoxamir. Microcirculation 19, 215–223 (2012).
Camps, C. et al. hsa-miR-210 Is induced by hypoxia and is an independent prognostic factor in breast cancer. Clin. Cancer Res. 14, 1340–1348 (2008).
Giannakakis, A. et al. miR-210 links hypoxia with cell cycle regulation and is deleted in human epithelial ovarian cancer. Cancer Biol. Ther. 7, 255–264 (2008).
Kulshreshtha, R. et al. A microRNA signature of hypoxia. Mol. Cell. Biol. 27, 1859–1867 (2007).
Mok, Y. et al. MiR-210 is induced by Oct-2, regulates B cells, and inhibits autoantibody production. J. Immunol. 191, 3037–3048 (2013).
O'Connell, R.M., Rao, D.S., Chaudhuri, A.A. & Baltimore, D. Physiological and pathological roles for microRNAs in the immune system. Nat. Rev. Immunol. 10, 111–122 (2010).
Baumjohann, D. & Ansel, K.M. MicroRNA-mediated regulation of T helper cell differentiation and plasticity. Nat. Rev. Immunol. 13, 666–678 (2013).
Kuchen, S. et al. Regulation of microRNA expression and abundance during lymphopoiesis. Immunity 32, 828–839 (2010).
Chong, M.M. et al. Canonical and alternate functions of the microRNA biogenesis machinery. Genes Dev. 24, 1951–1960 (2010).
Thai, T.H. et al. Regulation of the germinal center response by microRNA-155. Science 316, 604–608 (2007).
Fraser, J.D., Irving, B.A., Crabtree, G.R. & Weiss, A. Regulation of interleukin-2 gene enhancer activity by the T cell accessory molecule CD28. Science 251, 313–316 (1991).
Stittrich, A.B. et al. The microRNA miR-182 is induced by IL-2 and promotes clonal expansion of activated helper T lymphocytes. Nat. Immunol. 11, 1057–1062 (2010).
Kane, L.P., Andres, P.G., Howland, K.C., Abbas, A.K. & Weiss, A. Akt provides the CD28 costimulatory signal for up-regulation of IL-2 and IFN-γ but not TH2 cytokines. Nat. Immunol. 2, 37–44 (2001).
Huang, X. et al. Hypoxia-inducible mir-210 regulates normoxic gene expression involved in tumor initiation. Mol. Cell 35, 856–867 (2009).
Ciofani, M. et al. A validated regulatory network for Th17 cell specification. Cell 151, 289–303 (2012).
Park, C.Y. et al. A resource for the conditional ablation of microRNAs in the mouse. Cell Rep 1, 385–391 (2012).
Wang, X. & El Naqa, I.M. Prediction of both conserved and nonconserved microRNA targets in animals. Bioinformatics 24, 325–332 (2008).
Krek, A. et al. Combinatorial microRNA target predictions. Nat. Genet. 37, 495–500 (2005).
John, B. et al. Human microRNA targets. PLoS Biol. 2, e363 (2004).
Lewis, B.P., Shih, I.H., Jones-Rhoades, M.W., Bartel, D.P. & Burge, C.B. Prediction of mammalian microRNA targets. Cell 115, 787–798 (2003).
Betel, D., Koppal, A., Agius, P., Sander, C. & Leslie, C. Comprehensive modeling of microRNA targets predicts functional non-conserved and non-canonical sites. Genome Biol. 11, R90 (2010).
Huang, X. & Zuo, J. Emerging roles of miR-210 and other non-coding RNAs in the hypoxic response. Acta Biochim. Biophys. Sin. 10.1093/abbs/gmt141 (6 January 2014).
Tsuchiya, S. et al. MicroRNA-210 regulates cancer cell proliferation through targeting fibroblast growth factor receptor-like 1 (FGFRL1). J. Biol. Chem. 286, 420–428 (2011).
Favaro, E. et al. MicroRNA-210 regulates mitochondrial free radical response to hypoxia and krebs cycle in cancer cells by targeting iron sulfur cluster protein ISCU. PLoS ONE 5, e10345 (2010).
Chi, S.W., Zang, J.B., Mele, A. & Darnell, R.B. Argonaute HITS-CLIP decodes microRNA-mRNA interaction maps. Nature 460, 479–486 (2009).
Loeb, G.B. et al. Transcriptome-wide miR-155 binding map reveals widespread noncanonical microRNA targeting. Mol. Cell 48, 760–770 (2012).
Ikejiri, A. et al. Dynamic regulation of Th17 differentiation by oxygen concentrations. Int. Immunol. 24, 137–146 (2012).
Westermann, J. & Bode, U. Distribution of activated T cells migrating through the body: a matter of life and death. Immunol. Today 20, 302–306 (1999).
Vaupel, P., Thews, O., Kelleher, D.K. & Hoeckel, M. Current status of knowledge and critical issues in tumor oxygenation. Results from 25 years research in tumor pathophysiology. Adv. Exp. Med. Biol. 454, 591–602 (1998).
Henze, A.T. et al. Prolyl hydroxylases 2 and 3 act in gliomas as protective negative feedback regulators of hypoxia-inducible factors. Cancer Res. 70, 357–366 (2010).
Minamishima, Y.A. et al. A feedback loop involving the Phd3 prolyl hydroxylase tunes the mammalian hypoxic response in vivo. Mol. Cell. Biol. 29, 5729–5741 (2009).
Stiehl, D.P. et al. Increased prolyl 4-hydroxylase domain proteins compensate for decreased oxygen levels. Evidence for an autoregulatory oxygen-sensing system. J. Biol. Chem. 281, 23482–23491 (2006).
Park, C.Y., Choi, Y.S. & McManus, M.T. Analysis of microRNA knockouts in mice. Hum. Mol. Genet. 19, R169–R175 (2010).
Wang, R. et al. The transcription factor Myc controls metabolic reprogramming upon T lymphocyte activation. Immunity 35, 871–882 (2011).
Hoyer, K.K., Kuswanto, W.F., Gallo, E. & Abbas, A.K. Distinct roles of helper T-cell subsets in a systemic autoimmune disease. Blood 113, 389–395 (2009).
Steiner, D.F. et al. MicroRNA-29 regulates T-box transcription factors and interferon-gamma production in helper T cells. Immunity 35, 169–181 (2011).
Wang, H. et al. Tonic ubiquitylation controls T-cell receptor: CD3 complex expression during T-cell development. EMBO J. 29, 1285–1298 (2010).
Yamaji, O. et al. The development of colitogenic CD4+ T cells is regulated by IL-7 in collaboration with NK cell function in a murine model of colitis. J. Immunol. 188, 2524–2536 (2012).
Acknowledgements
We thank A. Roque for animal husbandry; R. Locksley and Z. Wang for access to the cell-sorting facility and assistance with cell sorting; J.J. O'Shea (US National Institutes of Health) for CP550690; K. Shokat (University of California, San Francisco) for GDC-0941 and MLN0128; R. Wang (St. Jude Children's Research Hospital) for organs from Hif1af/fCD4-Cre mice; M. Matloubian (University of California, San Francisco) for RNA from mice infected with lymphocytic choriomeningitis virus; A. Abbas (University of California, San Francisco) for Il2−/− DO11.10 mice; K. Shokat (University of California, San Francisco); J.J. O'Shea (US National Institutes of Health) for kinase inhibitors; and K. Mark Ansel and L. Jeker for critical reading of the manuscript. Supported by the Arthritis Foundation (H.W.), Deutsche Forschungsgemeinschaft (H.F.), the National Basic Research Program of China (2013CB967002 to L.W.) and the Keck Foundation.
Author information
Authors and Affiliations
Contributions
H.W. and H.F. planned and did experiments, analyzed and interpreted data and wrote the manuscript; M.O. did histology analysis; L.W. contributed critical prepublication data; M.T.M. provided guidance and direction and the Mir210-targeted mice; and A.W. supervised the work, helped conceive of the experiments and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Global miRNA expression–profiling studies reveal robust miR-210 induction after T cell activation.
(a) The expression of miR-210 in resting T cells, activated T cells and different helper T cell subsets. The primary data was from ref.57. (b) The expression of miR-210 in CD4+ and CD8+ thymocytes and activated T cells. The small RNA deep sequencing data was from ref.58,59.
Supplementary Figure 2 miR-210 induction in primary T cells in vitro and in vivo.
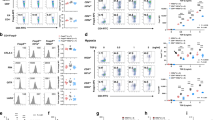
(a) Adoptively transferred T cells were harvested from various tissues 3 weeks after adoptive transfer. The surface-expression of CD44 and CD62L on adoptively transferred T cells were measured by flow cytometry. (b) Quantitative RT-PCR analysis of miR-210 abundance in resting or anti-CD3–anti-CD28 stimulated mouse CD8+ T cells. Cells were stimulated for a period of 4 d under normoxic (21% O2) or hypoxic (1% O2) conditions and RNA was harvested at the indicated time points. Relative expression was normalized to sno202. (c) Quantitative RTPCR analysis of miR-210 transcripts in P14 transgenic CD8+ T cells from mice infected with LCMV. P14 transgenic CD8+ T cells were injected into BoyJ congenic host mice. The chimeric mice were LCMV-infected 24 hrs after the adoptive transfer and P14 transgenic CD8+ T cells were harvested at the indicated time points, followed by RNA isolation and RT-PCR analysis of miR-210 abundance. Relative expression was normalized to sno202. (d) Quantitative RT-PCR analysis of miR-210 or miR155 abundance in resting or anti-CD3–anti-CD28 stimulated mouse or human primary T cells. Mouse CD4+ (left) and CD8+ (middle) T cells as well as human CD4+ (right) T cells were stimulated for a period of 4 d under normoxic (21% O2) or hypoxic (1% O2) conditions and RNA was harvested at the indicated time points. Relative expression is normalized to sno202 and the ratio of hypoxic vs. normoxic expression of miR-210 or miR-155 at the indicated timepoints is depicted. Data are from one experiment representative of two (a–c) or three (d) independent experiments. (mean and s.d. in b,c).
Supplementary Figure 3 HIF-1α protein is increased by CD28-mediated costimulation and HIF-1α binds the Mir210 promoter region.
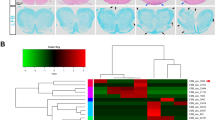
(a) Immunoblot analysis with a monoclonal HIF-1α-specific or GAPDH-specific antibody in total protein extracts of resting or anti-CD3–anti-CD28 stimulated primary CD8+ T cells. Cells were stimulated for 24 h under normoxic (21% O2) or hypoxic (1% O2) conditions and protein was harvested at the indicated time points. (b) Naive CD4+ T cells from wild-type (WT) or CD28-deficient (KO) animals were stimulated with anti-CD3 and anti-CD28 for 3 d or kept unstimulated. The cells were harvested and an immunoblot analysis with a monoclonal HIF-1α-specific or GAPDH-specific antibody in whole cell lysates was performed. (c) ChIP-seq analysis of the interaction of HIF-1α with the Mir210 promoter region in polarized TH17 cells. ChIP-seq primary data was from http://th17.bio.nyu.edu/pages/igv.php and ref.60. Naive T cells were polarized towards the TH17 lineage for 48 h. The results of two independent experiments are shown. The blue bar indicates the location of the Mir210 sequence. Data are representative of two (a,b) independent experiments.
Supplementary Figure 4 Analysis of miR-210 abundance during T cell development and identification of miR-210 target genes.
(a) Quantitative RT-PCR analysis of miR-210 abundance in flow cytometry sorted thymocyte populations and in resting or anti-CD3–anti-CD28 stimulated primary CD4+ T cells derived from wild-type or Mir210–/– animals. Relative expression is normalized to sno202. Data are from one experiment representative of three independent experiments. (mean and s.d.; abbreviations: DN, double negative; DP, double positive). (b) miR-210 target gene identification flow chart. miR-210 target genes in mouse T cells were selected by performing a two-step selection process. First, four algorithms, Target scan, PicTar, miRDB and miRanda, were used to predict miR-210 target genes. All targets predicted by miRanda were scored for an empirical probability of target inhibition using mirSVR scores and a stringent mirSVR score cutoff of -1.1. This list was combined with previously reported miR-210 targets, resulting in 69 potential miR-210 target genes (Supplementary Table 1). Next, T cell-expressed target genes were selected according to the Immgen data-base (www.immgen.org), resulting in 21 genes. Their expression was compared by RT-PCR in wild-type or Mir210–/– CD4+ T cells, which were activated for 4 d. Assuming that miR-210–deficiency resulted in a higher expression of direct miR-210 targets, five candidate genes were identified, that exhibited a more than two-fold increased expression in Mir210–/– CD4+ T cells
Supplementary Figure 5 Hif1a 3' UTR harbors a conserved miR-210 target sequence in the arognaute-binding region and hypoxic conditions inhibit T cell proliferation.
(a) AGO binds the miR-210 targeting site in Hif1a 3' UTR in activated T cells. AGO binding site information in primary T cells came from CLIP Base (http://bit.ly/12z7yMe) and ref.61. High-throughput sequencing of RNA isolated by crosslinking immunoprecipitation (HITSCLIP) was performed on T cells that were activated by anti-CD3 and anti-CD28 for 4 d at 37°C. Blue shade represents WT replicates, and yellow shade is miR-155-deficient replicates. The black bar indicates the location of the miR-210 seed region in the binding site. Coordinates along the x-axis indicate nucleotide position relative to the start of the Hif1a 3' UTR. The y-axis indicates read counts. (b) Sequence alignment of miR-210 target sequences from multiple species. (c) Naive mouse CD4+ T cells were anti-CD3–anti-CD28 stimulated for up to 4 d under normoxic (21% O2) or hypoxic (1% O2) conditions and cell numbers were determined at the indicated timepoints by using a Vi-Cell® counter. Data are representative of three (c) independent experiments.
Supplementary Figure 6 miR-210 deficiency along with reoxygenation conditions markedly increases TH17 differentiation but not TH1, TH2 or iTreg differentiation
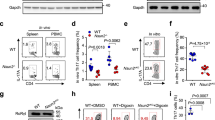
(a) The reoxygenation culture conditions, involving the differentiation of TH17 cells by a priming step under a low O2 concentration (5% O2) followed by transfer to normoxic conditions. (b–c) RT-PCR analysis of miR-210 abundance and immunoblot analysis of HIF8 1α and GAPDH in CD4+ T cells stimulated in nonpolarizing conditions under normoxic or reoxygenation conditions. Relative miR-210 expression is normalized to its expression in naïve T cells. (d) Naive CD4+ T cells from wild-type or Mir210–/– mice were differentiated under TH1, TH2, TH17 (0.2 ng/ml of IL-6) or iTreg skewing conditions under normoxic or reoxygenetion conditions (see scheme in a), followed by intracellular staining of IL-4, IL-17A, IFN-γ or Foxp3. For TH1 conditions, 10 μg/ml anti–IL-4, 100 U/ml IL-2, 3.5 ng/ml IL-12 were added to the cultures; For TH2 conditions, 10 μg/ml anti–IFN-g, 100 U/ml IL-2, 50 ng/ml IL-4 were added to the cultures; For iTreg assays, naive T cells were cultured in the presence of 0.5 ng/ml TGF-β. (e) Naive CD4+ T cells were polarized towards iTregs under normoxic or reoxygenation conditions with varying doses of TGF-β for 3 d, followed by intracellular Foxp3 staining. (f) Wild-type or Mir210–/– CD4+ T cells were differentiated as described in Fig. 7a. Foxp3 expression was assessed by intracellular staining. Data are from one experiment representative of two (b,c,e,f) or three (d) independent experiments. (mean and s.d. in b).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Tables 1 and 2 (PDF 5024 kb)
Rights and permissions
About this article
Cite this article
Wang, H., Flach, H., Onizawa, M. et al. Negative regulation of Hif1a expression and TH17 differentiation by the hypoxia-regulated microRNA miR-210. Nat Immunol 15, 393–401 (2014). https://doi.org/10.1038/ni.2846
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.2846
This article is cited by
-
The roles of microRNAs in mouse development
Nature Reviews Genetics (2021)
-
Hypoxia-challenged MSC-derived exosomes deliver miR-210 to attenuate post-infarction cardiac apoptosis
Stem Cell Research & Therapy (2020)
-
IFN-γ promoted exosomes from mesenchymal stem cells to attenuate colitis via miR-125a and miR-125b
Cell Death & Disease (2020)
-
MiRNA-210 induces microglial activation and regulates microglia-mediated neuroinflammation in neonatal hypoxic-ischemic encephalopathy
Cellular & Molecular Immunology (2020)
-
microRNAs as therapeutic targets in intestinal diseases
ExRNA (2019)