Abstract
Regulatory T (Treg) cells respond to immune and inflammatory signals to mediate immunosuppression, but how the functional integrity of Treg cells is maintained under activating environments is unclear. Here we show that autophagy is active in Treg cells and supports their lineage stability and survival fitness. Treg cell–specific deletion of Atg7 or Atg5, two essential genes in autophagy, leads to loss of Treg cells, greater tumor resistance and development of inflammatory disorders. Atg7-deficient Treg cells show increased apoptosis and readily lose expression of the transcription factor Foxp3, especially after activation. Mechanistically, autophagy deficiency upregulates metabolic regulators mTORC1 and c-Myc and glycolysis, which contribute to defective Treg function. Therefore, autophagy couples environmental signals and metabolic homeostasis to protect lineage and survival integrity of Treg cells in activating contexts.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Josefowicz, S.Z., Lu, L.F., Rudensky, A.Y. & Regulatory, T. Cells: mechanisms of differentiation and function. Annu. Rev. Immunol. 30, 531–564 (2012).
Liston, A. & Gray, D.H.D. Homeostatic control of regulatory T cell diversity. Nat. Rev. Immunol. 14, 154–165 (2014).
Smigiel, K.S. et al. CCR7 provides localized access to IL-2 and defines homeostatically distinct regulatory T cell subsets. J. Exp. Med. 211, 121–136 (2014).
Fisson, S. et al. Continuous activation of autoreactive CD4+CD25+ regulatory T cells in the steady state. J. Exp. Med. 198, 737–746 (2003).
Arvey, A. et al. Inflammation-induced repression of chromatin bound by the transcription factor Foxp3 in regulatory T cells. Nat. Immunol. 15, 580–587 (2014).
Rubtsov, Y.P. et al. Stability of the regulatory T cell lineage in vivo. Science 329, 1667–1671 (2010).
Zhou, X.Y. et al. Instability of the transcription factor Foxp3 leads to the generation of pathogenic memory T cells in vivo. Nat. Immunol. 10, 1000–1007 (2009).
Bailey-Bucktrout, S.L. et al. Self-antigen-driven activation induces instability of regulatory T cells during an inflammatory autoimmune response. Immunity 39, 949–962 (2013).
Komatsu, N. et al. Pathogenic conversion of Foxp3+ T cells into TH17 cells in autoimmune arthritis. Nat. Med. 20, 62–68 (2014).
Tsuji, M. et al. Preferential generation of follicular B helper T cells from Foxp3+ T cells in gut Peyer's patches. Science 323, 1488–1492 (2009).
Pierson, W. et al. Antiapoptotic Mcl-1 is critical for the survival and niche-filling capacity of Foxp3+ regulatory T cells. Nat. Immunol. 14, 959–965 (2013).
Levine, B., Mizushima, N. & Virgin, H.W. Autophagy in immunity and inflammation. Nature 469, 323–335 (2011).
Jiang, X., Overholtzer, M. & Thompson, C.B. Autophagy in cellular metabolism and cancer. J. Clin. Invest. 125, 47–54 (2015).
Galluzzi, L., Pietrocola, F., Levine, B. & Kroemer, G. Metabolic control of autophagy. Cell 159, 1263–1276 (2014).
Pua, H.H., Dzhagalov, I., Chuck, M., Mizushima, N. & He, Y.W. A critical role for the autophagy gene Atg5 in T cell survival and proliferation. J. Exp. Med. 204, 25–31 (2007).
Xu, X. et al. Autophagy is essential for effector CD8+ T cell survival and memory formation. Nat. Immunol. 15, 1152–1161 (2014).
Hubbard, V.M. et al. Macroautophagy regulates energy metabolism during effector T cell activation. J. Immunol. 185, 7349–7357 (2010).
Li, C. et al. Autophagy is induced in CD4+ T cells and important for the growth factor–withdrawal cell death. J. Immunol. 177, 5163–5168 (2006).
Pengo, N. et al. Plasma cells require autophagy for sustainable immunoglobulin production. Nat. Immunol. 14, 298–305 (2013).
MacIver, N.J., Michalek, R.D. & Rathmell, J.C. Metabolic regulation of T lymphocytes. Annu. Rev. Immunol. 31, 259–283 (2013).
Mizushima, N., Yamamoto, A., Matsui, M., Yoshimori, T. & Ohsumi, Y. In vivo analysis of autophagy in response to nutrient starvation using transgenic mice expressing a fluorescent autophagosome marker. Mol. Biol. Cell 15, 1101–1111 (2004).
Li, X., Liang, Y., LeBlanc, M., Benner, C. & Zheng, Y. Function of a Foxp3 cis-element in protecting regulatory T cell identity. Cell 158, 734–748 (2014).
Feng, Y. et al. Control of the inheritance of regulatory T cell identity by a cis element in the Foxp3 locus. Cell 158, 749–763 (2014).
Liu, G., Yang, K., Burns, S., Shrestha, S. & Chi, H. The S1P1-mTOR axis directs the reciprocal differentiation of TH1 and Treg cells. Nat. Immunol. 11, 1047–1056 (2010).
Yang, K. et al. T cell exit from quiescence and differentiation into Th2 cells depend on Raptor-mTORC1-mediated metabolic reprogramming. Immunity 39, 1043–1056 (2013).
Chi, H. Regulation and function of mTOR signalling in T cell fate decisions. Nat. Rev. Immunol. 12, 325–338 (2012).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Roychoudhuri, R. et al. BACH2 represses effector programs to stabilize Treg-mediated immune homeostasis. Nature 498, 506–510 (2013).
Ouyang, W. et al. Foxo proteins cooperatively control the differentiation of Foxp3+ regulatory T cells. Nat. Immunol. 11, 618–627 (2010).
Gerriets, V.A. et al. Metabolic programming and PDHK1 control CD4+ T cell subsets and inflammation. J. Clin. Invest. 125, 194–207 (2015).
Michalek, R.D. et al. Cutting edge: distinct glycolytic and lipid oxidative metabolic programs are essential for effector and regulatory CD4+ T cell subsets. J. Immunol. 186, 3299–3303 (2011).
Shi, L.Z. et al. HIF1α-dependent glycolytic pathway orchestrates a metabolic checkpoint for the differentiation of TH17 and Treg cells. J. Exp. Med. 208, 1367–1376 (2011).
Polansky, J.K. et al. DNA methylation controls Foxp3 gene expression. Eur. J. Immunol. 38, 1654–1663 (2008).
Finlay, D.K. et al. PDK1 regulation of mTOR and hypoxia-inducible factor 1 integrate metabolism and migration of CD8+ T cells. J. Exp. Med. 209, 2441–2453 (2012).
Wang, R. et al. The transcription factor Myc controls metabolic reprogramming upon T lymphocyte activation. Immunity 35, 871–882 (2011).
Delmore, J.E. et al. BET bromodomain inhibition as a therapeutic strategy to target c-Myc. Cell 146, 904–917 (2011).
Bandukwala, H.S. et al. Selective inhibition of CD4+ T-cell cytokine production and autoimmunity by BET protein and c-Myc inhibitors. Proc. Natl. Acad. Sci. USA 109, 14532–14537 (2012).
Boursalian, T.E., Golob, J., Soper, D.M., Cooper, C.J. & Fink, P.J. Continued maturation of thymic emigrants in the periphery. Nat. Immunol. 5, 418–425 (2004).
Yu, L. et al. Termination of autophagy and reformation of lysosomes regulated by mTOR. Nature 465, 942–946 (2010).
Zeng, H. et al. mTORC1 couples immune signals and metabolic programming to establish Treg-cell function. Nature 499, 485–490 (2013).
Park, Y. et al. TSC1 regulates the balance between effector and regulatory T cells. J. Clin. Invest. 123, 5165–5178 (2013).
Paul, S., Kashyap, A.K., Jia, W., He, Y.W. & Schaefer, B.C. Selective autophagy of the adaptor protein Bcl10 modulates T cell receptor activation of NF-κB. Immunity 36, 947–958 (2012).
Shrestha, S. et al. Treg cells require the phosphatase PTEN to restrain TH1 and TFH cell responses. Nat. Immunol. 16, 178–187 (2015).
Huynh, A. et al. Control of PI(3) kinase in Treg cells maintains homeostasis and lineage stability. Nat. Immunol. 16, 188–196 (2015).
De Rosa, V. et al. Glycolysis controls the induction of human regulatory T cells by modulating the expression of FOXP3 exon 2 splicing variants. Nat. Immunol. 16, 1174–1184 (2015).
Komatsu, M. et al. Impairment of starvation-induced and constitutive autophagy in Atg7-deficient mice. J. Cell Biol. 169, 425–435 (2005).
Hara, T. et al. Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 441, 885–889 (2006).
Rubtsov, Y.P. et al. Reaulatory T cell-derived interleukin-10 limits inflammation at environmental interfaces. Immunity 28, 546–558 (2008).
Wang, Y. et al. Tuberous sclerosis 1 (Tsc1)-dependent metabolic checkpoint controls development of dendritic cells. Proc. Natl. Acad. Sci. USA 110, E4894–E4903 (2013).
Acknowledgements
The authors acknowledge B. Rhode, C. Cloer and S. Rankin for animal colony management, X. Du for help with immunoblot analysis, N. Kalupahana for initial analysis of Atg7-deficient mice, N. Mizushima for GFP-LC3 mice, A. Rudensky for Foxp3YFP-Cre mice, J.E. Bradner for providing the JQ-1 compound, St. Jude Immunology FACS core facility for cell sorting and Y. Wang for editing. Supported by US National Institutes of Health (NIH AI105887, AI101407, CA176624 and NS064599 to H.C.), and the Crohn's and Colitis Foundation of America (to H.C.).
Author information
Authors and Affiliations
Contributions
J.W. designed and performed cellular, molecular and biochemical experiments and wrote the manuscript; L.L. performed immunoblot analysis; K.Y. performed Seahorse assays and helped early development of the project; C.G. performed imaging assays; S.S., Z.C. and C.W. performed the TSDR methylation analysis; P.V. performed histopathology analysis; G.N. performed bioinformatic analysis; D.R.G. contributed genetic models; and H.C. designed experiments, wrote the manuscript, and provided overall direction.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
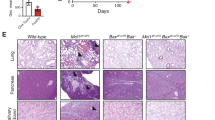
Supplementary Figure 1 Ablation of autophagy in Foxp3CreAtg7fl/fl mice and analysis of homeostasis of Foxp3CreAtg7fl/fl mice, Foxp3CreAtg5fl/fl mice and mixed BM chimeras.
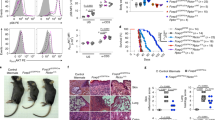
(a) Immunoblot analysis of LC3-I and LC3-II in naïve CD4 and Treg cells from Foxp3Cre mice and in Treg cells from Foxp3CreAtg7fl/fl mice. Cells were rested in complete medium for 1 h in the presence or absence of 100 nM Baf1A. (b) Foxp3CreAtg7+/fl and Foxp3CreAtg7fl/fl mice (n=4 per genotype) were inoculated with MC38 colon adenocarcinoma cells. Flow cytometry analyzing CD4 and CD8 expression (left), and frequency of CD4+ and CD8+ T cells (right) in tumor-infiltrating lymphocytes. Numbers adjacent to outlined areas indicate percent CD4+ and CD8+ cells (left). (c) Number of CD4+ and CD8+ T cells in the spleen and PLNs of Foxp3CreAtg7+/fl and Foxp3CreAtg7fl/fl mice (n=5 mice per genotype). (d) Flow cytometry analyzing IL-4 expression in Foxp3−CD4+ and CD8+ T cells from the spleen of Foxp3CreAtg7+/fl and Foxp3CreAtg7fl/fl mice. Numbers adjacent to outlined areas indicate percent IL-4+ cells. (e) Flow cytometry analyzing Foxp3 expression in CD4+ T cells in the colon lamina propria of Foxp3CreAtg7+/fl and Foxp3CreAtg7fl/fl mice (3-4 weeks old). Numbers adjacent to outlined areas indicate percent Foxp3+ cells. (f) Flow cytometry analyzing YFP-Foxp3 expression in CD45.2+CD4+ T cells in the spleen of Foxp3CreAtg7+/fl and Foxp3CreAtg7fl/fl chimeras. Numbers adjacent to outlined areas indicate percent YFP-Foxp3+ cells. (g,h) Flow cytometry analyzing the expression of CD62L and CD44 (g), and IFN-γ (h) in Foxp3−CD4+ and CD8+ T cells and in the spleen of Foxp3CreAtg5+/fl and Foxp3CreAtg5fl/fl mice. Numbers in quadrants indicate percent cells in each (g), and numbers adjacent to outlined areas indicate percent IFN-γ+ cells (h). (i) Flow cytometry analyzing Foxp3 expression in CD4+ T cells in the spleen and PLNs of Foxp3CreAtg5+/fl and Foxp3CreAtg5fl/fl mice. Numbers adjacent to outlined areas indicate percent Foxp3+ cells. NS, not significant (P > 0.05); * P < 0.05 and ** P < 0.001 (two-tail unpaired Student's t-test in b,c). Data are representative of three (a, g-i), two (b) or five (d-f) experiments, or pooled from three out of six (c) experiments (mean ± s.e.m in b,c).
Supplementary Figure 2 Proliferation, survival and lineage stability of Treg cells from Foxp3CreAtg7fl/fl and Foxp3CreAtg7fl/flBcl2-TG mice.
(a,b) Flow cytometry analyzing Ki67 expression in Treg cells from the spleen of Foxp3CreAtg7+/fl and Foxp3CreAtg7fl/fl mice (a) and Foxp3CreAtg7+/fl and Foxp3CreAtg7fl/fl chimeras (b). Numbers above bracketed lines indicate percent Ki67+ cells (a,b). (c) Flow cytometry analyzing CellTrace dilution in Treg cells stimulated with anti-CD3, anti-CD28, and IL-2 for 96 h. Numbers above bracketed lines indicate percent CellTracelo cells. (d) Foxp3CreAtg7+/fl or Foxp3CreAtg7fl/fl Treg cells were transferred into Rag1−/− mice, followed by flow cytometry analysis of CellTrace dilution in donor cells in the spleen and PLNs of recipients 7 days later. Numbers above bracketed lines indicate percent CellTracelo cells. (e,f) Flow cytometry analyzing the expression of active caspase-3 (e) and Bim (f) in CD45.2+ Treg cells in the spleen of Foxp3CreAtg7+/fl and Foxp3CreAtg7fl/fl chimeras. Numbers adjacent to outlined areas indicate percent caspase-3+ cells (e), and numbers above graph indicate MFI of Bim (f). (g) Flow cytometry analyzing the expression of IFN-γ and IL-17 (left), and frequency of IFN-γ+ cells and IL-17+ cells (right) in MC38 tumor-infiltrating Foxp3CreAtg7+/fl and Foxp3CreAtg7fl/fl Treg cells (n=8 mice per genotype). Numbers in quadrants indicate percent cells in each throughout (left). (h) Flow cytometry analyzing the expression of IFN-γ and IL-17 in Foxp3CreAtg7+/fl and Foxp3CreAtg7fl/fl Treg cells stimulated in vitro with anti-CD3, anti-CD28 and IL-2 for 96 h. (i) Flow cytometry analyzing active caspase-3 expression in Treg cells (from Foxp3CreAtg7+/fl, Foxp3CreAtg7fl/fl Foxp3CreAtg7+/flBcl2-TG and Foxp3CreAtg7fl/flBcl2-TG mice) stimulated with anti-CD3, anti-CD28, and IL-2 for overnight. Numbers above bracketed lines indicate percent caspase-3+ cells. (j,k) Flow cytometry analyzing the expression of Foxp3 in CD4+ T cells (j) and CD62L and CD44 in Foxp3−CD4+ T cells (k) in the spleen of Foxp3CreAtg7+/fl, Foxp3CreAtg7fl/fl Foxp3CreAtg7+/flBcl2-TG and Foxp3CreAtg7fl/flBcl2-TG mice. Numbers adjacent to outlined areas indicate percent Foxp3+ cells (j). (l) Flow cytometry analyzing the expression of IFN-γ and IL-17 in Treg cells in the spleen of Foxp3CreAtg7+/flBcl2-TG and Foxp3CreAtg7fl/flBcl2-TG mice. (m) Flow cytometry analyzing YFP-Foxp3 expression in divided Treg cells (from Foxp3CreAtg7+/flBcl2-TG and Foxp3CreAtg7fl/flBcl2-TG mice) stimulated in vitro with anti-CD3, anti-CD28 and IL-2 for 96 h. Numbers above graphs indicate MFI of YFP-Foxp3. NS, not significant (P > 0.05); * P < 0.05 (two-tail unpaired Student's t-test in g). Data are representative of four (a-f), two (h,m) or three (i-l) experiments, or pooled from two out of two (g) experiments (mean ± s.e.m in g).
Supplementary Figure 3 Control of mTORC1 activity by autophagy in Treg cells.
(a) Flow cytometry analyzing cell size and the expression of CD71, CD98 and p-S6 in Treg cells from the spleen of Foxp3CreAtg5+/fl and Foxp3CreAtg5fl/fl mice. Numbers above graphs indicate MFI of FSC-A, CD71, CD98 or p-S6. (b) Immunoblot analysis of Lck, p110δ, p85 and PDK1 in Treg cells from Foxp3CreAtg7+/fl and Foxp3CreAtg7fl/fl mice. Data are representative of three (a,b) experiments.
Supplementary Figure 4 Autophagy-dependent transcriptional profiles in Treg cells are responsive to rapamycin.
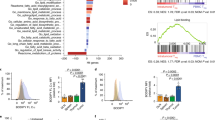
Foxp3CreAtg7+/fl and Foxp3CreAtg7fl/fl mice were treated with or without rapamycin (n=4 mice per group). Treg cells were sorted and activated with anti-CD3 and anti-CD28 for 4 h for gene expression profiling. (a) The top 10 canonical pathways upregulated in Foxp3CreAtg7fl/fl Treg cells as compared to Foxp3CreAtg7+/fl Treg cells by ingenuity pathway analysis (IPA) of the differentially expressed genes at the 0.5 log2 cut-offs. (b) Upstream regulators (kinases and transcription regulators) in Foxp3CreAtg7fl/fl Treg cells analyzed by IPA. Positive and negative activation z-scores (at the ± 2 cut-offs) indicate the activation and suppression of gene function in Foxp3CreAtg7fl/fl Treg cells respectively. (c,d) Heat maps of the expression changes of top hit genes in caspase (c) and cytokine pathways (d) in non-treated Foxp3CreAtg7fl/fl versus Foxp3CreAtg7+/fl Treg cells and rapamycin-treated Foxp3CreAtg7fl/fl versus Foxp3CreAtg7+/fl Treg cells. Red color denotes upregulated genes in Foxp3CreAtg7fl/fl Treg cells, and blue color denotes downregulated genes in Foxp3CreAtg7fl/fl Treg cells. Data are from one experiment (a-d).
Supplementary Figure 5 Metabolic and epigenetic analyses of Atg7-deficient Treg cells.
(a) Measurement of OCR in Treg cells (from Foxp3CreAtg7+/fl and Foxp3CreAtg7fl/fl mice) stimulated with anti-CD3 and anti-CD28 for 4 h. (b) Methylation status of CpG motifs of the TSDR at Foxp3 locus, assessed by bisulfite sequencing of divided Treg cells (from Foxp3CreAtg7+/fl and Foxp3CreAtg7fl/fl mice) activated with anti-CD3, anti-CD28 and IL-2 for 96 h in the presence of DMSO or DCA. Numbers above boxes (1-14) indicate the 14 CpG islands from 5′ to 3′ in the intron 1 of the Foxp3 locus. (c) Immunoblot analysis of HK2 in Treg cells (from Foxp3CreAtg7+/fl and Foxp3CreAtg7fl/fl mice) stimulated with anti-CD3 and anti-CD28 for 4 h. (d) Foxp3CreAtg7+/fl and Foxp3CreAtg7fl/fl mice received mock or rapamycin treatment (n=4 mice per group). Treg cells were sorted and stimulated with anti-CD3 and anti-CD28 for 4 h, followed by analysis of mRNA expression of Hk2. (e,f) mRNA expression (e) and immunoblot analysis (f) of HIF1α in Foxp3CreAtg7+/fl and Foxp3CreAtg7fl/fl Treg cells stimulated with anti-CD3 and anti-CD28 for 4 h. NS, not significant (P > 0.05); * P < 0.05 (two-tail unpaired Student's t-test in a,e and one-way ANOVA in d). Data are representative of two (a) or three (b,c,f) experiments, or pooled from four out of four (d,e) experiments (mean ± s.e.m in a,d,e).
Supplementary Figure 6 Autophagy regulates c-Myc expression in an mTORC1-dependent manner.
(a) Expression of c-Myc in Treg cells (from Cd4CreAtg7+/flRptor+/fl, Cd4CreAtg7fl/flRptor+/fl, Cd4CreAtg7+/flRptorfl/fl and Cd4CreAtg7fl/flRptorfl/fl mice) stimulated with anti-CD3 and anti-CD28 for 4 h. Numbers above bracketed lines indicate percent c-Myc+ cells. (b) Flow cytometry analyzing YFP-Foxp3 expression in divided Treg cells (from Foxp3CreAtg7+/fl and Foxp3CreAtg7fl/fl mice) that were activated in vitro with anti-CD3 and anti-CD28, and IL-2 for 96 h in the presence of DMSO or i-BET-762. Numbers above graphs indicate MFI of YFP-Foxp3. Data are representative of three independent experiments (a ,b).
Supplementary Figure 7 Schematics of autophagy in Treg cell functional integrity and immune tolerance.
Environmental cues drive the activation and functional maturation of Treg cells, but also potently activate mTORC1 which could be detrimental to Treg cell maintenance. Consequently, activated Treg cells upregulate autophagy in response to immune and inflammatory signals to protect their functional integrity by coordinately regulating lineage stability and survival. Specifically, autophagy downregulates the mTORC1-c-Myc pathway to maintain the metabolic homeostasis to support lineage stability of Treg cells. Additionally, autophagy is crucial for inhibiting the apoptotic pathway, which is partially mediated by restraining mTORC1 activity. In the absence of autophagy, the impaired lineage stability and increased apoptosis of Treg cells lead to the loss of Treg cell-mediated immune tolerance.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 (PDF 1104 kb)
Rights and permissions
About this article
Cite this article
Wei, J., Long, L., Yang, K. et al. Autophagy enforces functional integrity of regulatory T cells by coupling environmental cues and metabolic homeostasis. Nat Immunol 17, 277–285 (2016). https://doi.org/10.1038/ni.3365
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3365
This article is cited by
-
IL-23 stabilizes an effector Treg cell program in the tumor microenvironment
Nature Immunology (2024)
-
Exploiting autophagy balance in T and NK cells as a new strategy to implement adoptive cell therapies
Molecular Cancer (2023)
-
Impact of context-dependent autophagy states on tumor progression
Nature Cancer (2023)
-
Artesunate targets cellular metabolism to regulate the Th17/Treg cell balance
Inflammation Research (2023)
-
Dietary restriction to optimize T cell immunity is an ancient survival strategy conserved in vertebrate evolution
Cellular and Molecular Life Sciences (2023)