Abstract
Genes and pathways in which inactivation dampens tissue inflammation present new opportunities for understanding the pathogenesis of common human inflammatory diseases, including inflammatory bowel disease, rheumatoid arthritis and multiple sclerosis. We identified a mutation in the gene encoding the deubiquitination enzyme USP15 (Usp15L749R) that protected mice against both experimental cerebral malaria (ECM) induced by Plasmodium berghei and experimental autoimmune encephalomyelitis (EAE). Combining immunophenotyping and RNA sequencing in brain (ECM) and spinal cord (EAE) revealed that Usp15L749R-associated resistance to neuroinflammation was linked to dampened type I interferon responses in situ. In hematopoietic cells and in resident brain cells, USP15 was coexpressed with, and functionally acted together with the E3 ubiquitin ligase TRIM25 to positively regulate type I interferon responses and to promote pathogenesis during neuroinflammation. The USP15-TRIM25 dyad might be a potential target for intervention in acute or chronic states of neuroinflammation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
Change history
24 October 2016
In the version of this article initially published online, the symbol for the gene encoding granzyme B was incorrect (Gmzb) in the text in the third paragraph of the fourth subsection of Results and Figure 5d, and the symbol for the gene encoding granzyme A was incorrect (Gmza) in Figure 6h. These should be Gzmb and Gzma, respectively. The errors have been corrected for the print, PDF and HTML versions of this article.
References
Hunt, N.H. & Grau, G.E. Cytokines: accelerators and brakes in the pathogenesis of cerebral malaria. Trends Immunol. 24, 491–499 (2003).
Hansen, D.S. Inflammatory responses associated with the induction of cerebral malaria: lessons from experimental murine models. PLoS Pathog. 8, e1003045 (2012).
Brown, H. et al. Evidence of blood-brain barrier dysfunction in human cerebral malaria. Neuropathol. Appl. Neurobiol. 25, 331–340 (1999).
de Souza, J.B. & Riley, E.M. Cerebral malaria: the contribution of studies in animal models to our understanding of immunopathogenesis. Microbes Infect. 4, 291–300 (2002).
Ochiel, D.O. et al. Differential regulation of beta-chemokines in children with Plasmodium falciparum malaria. Infect. Immun. 73, 4190–4197 (2005).
Armah, H.B. et al. Cerebrospinal fluid and serum biomarkers of cerebral malaria mortality in Ghanaian children. Malar. J. 6, 147 (2007).
Kim, H. et al. Functional roles for C5a and C5aR but not C5L2 in the pathogenesis of human and experimental cerebral malaria. Infect. Immun. 82, 371–379 (2014).
Longley, R. et al. Host resistance to malaria: using mouse models to explore the host response. Mamm. Genome 22, 32–42 (2011).
Senaldi, G. et al. Protection against the mortality associated with disease models mediated by TNF and IFN-γ in mice lacking IFN regulatory factor-1. J. Immunol. 163, 6820–6826 (1999).
Berghout, J. et al. Irf8-regulated genomic responses drive pathological inflammation during cerebral malaria. PLoS Pathog. 9, e1003491 (2013).
Caignard, G. et al. Mouse ENU mutagenesis to understand immunity to infection: methods, selected examples, and perspectives. Genes (Basel) 5, 887–925 (2014).
Bongfen, S.E. et al. An N-ethyl-N-nitrosourea (ENU)-induced dominant negative mutation in the JAK3 kinase protects against cerebral malaria. PLoS One 7, e31012 (2012).
Torre, S. et al. THEMIS is required for pathogenesis of cerebral malaria and protection against pulmonary tuberculosis. Infect. Immun. 83, 759–768 (2015).
Kennedy, J.M. et al. CCDC88B is a novel regulator of maturation and effector functions of T cells during pathological inflammation. J. Exp. Med. 211, 2519–2535 (2014).
Sawcer, S. et al. Genetic risk and a primary role for cell-mediated immune mechanisms in multiple sclerosis. Nature 476, 214–219 (2011).
Dubois, P.C.A. et al. Multiple common variants for celiac disease influencing immune gene expression. Nat. Genet. 42, 295–302 (2010).
Beecham, A.H. et al. Analysis of immune-related loci identifies 48 new susceptibility variants for multiple sclerosis. Nat. Genet. 45, 1353–1360 (2013).
Jostins, L. et al. Host-microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012).
Okada, Y. et al. Genetics of rheumatoid arthritis contributes to biology and drug discovery. Nature 506, 376–381 (2014).
Cunninghame Graham, D.S. et al. Association of NCF2, IKZF1, IRF8, IFIH1, and TYK2 with systemic lupus erythematosus. PLoS Genet. 7, e1002341 (2011).
Fodil, N., Langlais, D. & Gros, P. Primary immunodeficiencies and inflammatory disease: a growing genetic intersection. Trends Immunol. 37, 126–140 (2016).
Pauli, E.-K. et al. The ubiquitin-specific protease USP15 promotes RIG-I-mediated antiviral signaling by deubiquitylating TRIM25. Sci. Signal. 7, ra3 (2014).
Gack, M.U. et al. TRIM25 RING-finger E3 ubiquitin ligase is essential for RIG-I-mediated antiviral activity. Nature 446, 916–920 (2007).
Torre, S. et al. Susceptibility to lethal cerebral malaria is regulated by epistatic interaction between chromosome 4 (Berr6) and chromosome 1 (Berr7) loci in mice. Genes Immun. 14, 249–257 (2013).
de Jong, R.N. et al. Solution structure of the human ubiquitin-specific protease 15 DUSP domain. J. Biol. Chem. 281, 5026–5031 (2006).
Hetfeld, B.K. et al. The zinc finger of the CSN-associated deubiquitinating enzyme USP15 is essential to rescue the E3 ligase Rbx1. Curr. Biol. 15, 1217–1221 (2005).
Man, S., Ubogu, E.E. & Ransohoff, R.M. Inflammatory cell migration into the central nervous system: a few new twists on an old tale. Brain Pathol. 17, 243–250 (2007).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Rusinova, I. et al. Interferome v2.0: an updated database of annotated interferon-regulated genes. Nucleic Acids Res. 41, D1040–D1046 (2013).
Schweitzer, K., Bozko, P.M., Dubiel, W. & Naumann, M. CSN controls NF-kappaB by deubiquitinylation of IkappaBalpha. EMBO J. 26, 1532–1541 (2007).
Cornelissen, T. et al. The deubiquitinase USP15 antagonizes Parkin-mediated mitochondrial ubiquitination and mitophagy. Hum. Mol. Genet. 23, 5227–5242 (2014).
Hayes, S.D. et al. Direct and indirect control of mitogen-activated protein kinase pathway-associated components, BRAP/IMP E3 ubiquitin ligase and CRAF/RAF1 kinase, by the deubiquitylating enzyme USP15. J. Biol. Chem. 287, 43007–43018 (2012).
Villeneuve, N.F. et al. USP15 negatively regulates Nrf2 through deubiquitination of Keap1. Mol. Cell 51, 68–79 (2013).
Long, L. et al. The U4/U6 recycling factor SART3 has histone chaperone activity and associates with USP15 to regulate H2B deubiquitination. J. Biol. Chem. 289, 8916–8930 (2014).
Inui, M. et al. USP15 is a deubiquitylating enzyme for receptor-activated SMADs. Nat. Cell Biol. 13, 1368–1375 (2011).
Eichhorn, P.J.A. et al. USP15 stabilizes TGF-β receptor I and promotes oncogenesis through the activation of TGF-β signaling in glioblastoma. Nat. Med. 18, 429–435 (2012).
Iyengar, P.V. et al. USP15 regulates SMURF2 kinetics through C-lobe mediated deubiquitination. Sci. Rep. 5, 14733 (2015).
Zou, Q. et al. USP15 stabilizes MDM2 to mediate cancer-cell survival and inhibit antitumor T cell responses. Nat. Immunol. 15, 562–570 (2014).
Zhang, H. et al. Ubiquitin-specific protease 15 negatively regulates virus-induced type I interferon signaling via catalytically-dependent and -independent mechanisms. Sci Rep. 5, 11220 (2015).
Inn, K.-S. et al. Linear ubiquitin assembly complex negatively regulates RIG-I- and TRIM25-mediated type I interferon induction. Mol. Cell 41, 354–365 (2011).
Liehl, P. et al. Host-cell sensors for Plasmodium activate innate immunity against liver-stage infection. Nat. Med. 20, 47–53 (2014).
Gazzinelli, R.T., Kalantari, P., Fitzgerald, K.A. & Golenbock, D.T. Innate sensing of malaria parasites. Nat. Rev. Immunol. 14, 744–757 (2014).
Miller, J.L., Sack, B.K., Baldwin, M., Vaughan, A.M. & Kappe, S.H.I. Interferon-mediated innate immune responses against malaria parasite liver stages. Cell Rep. 7, 436–447 (2014).
Ball, E.A. et al. IFNAR1 controls progression to cerebral malaria in children and CD8+ T cell brain pathology in Plasmodium berghei-infected mice. J. Immunol. 190, 5118–5127 (2013).
Palomo, J. et al. Type I interferons contribute to experimental cerebral malaria development in response to sporozoite or blood-stage Plasmodium berghei ANKA. Eur. J. Immunol. 43, 2683–2695 (2013).
Sharma, S. et al. Innate immune recognition of an AT-rich stem-loop DNA motif in the Plasmodium falciparum genome. Immunity 35, 194–207 (2011).
Coban, C. et al. Pathological role of Toll-like receptor signaling in cerebral malaria. Int. Immunol. 19, 67–79 (2007).
Togbe, D. et al. Murine cerebral malaria development is independent of toll-like receptor signaling. Am. J. Pathol. 170, 1640–1648 (2007).
Imboden, M. et al. Genome-wide association study of lung function decline in adults with and without asthma. J. Allergy Clin. Immunol. 129, 1218–1228 (2012).
Orimo, A. et al. Underdeveloped uterus and reduced estrogen responsiveness in mice with disruption of the estrogen-responsive finger protein gene, which is a direct target of estrogen receptor alpha. Proc. Natl. Acad. Sci. USA 96, 12027–12032 (1999).
Urano, T. et al. Efp targets 14-3-3 sigma for proteolysis and promotes breast tumour growth. Nature 417, 871–875 (2002).
Tanaka, K. et al. Loss of suppressor of cytokine signaling 1 in helper T cells leads to defective Th17 differentiation by enhancing antagonistic effects of IFN-gamma on STAT3 and Smads. J. Immunol. 180, 3746–3756 (2008).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Quinlan, A.R. & Hall, I.M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Gold, R., Linington, C. & Lassmann, H. Understanding pathogenesis and therapy of multiple sclerosis via animal models: 70 years of merits and culprits in experimental autoimmune encephalomyelitis research. Brain 129, 1953–1971 (2006).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Liao, Y., Smyth, G.K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Robinson, M.D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25 (2010).
Thorvaldsdóttir, H., Robinson, J.T. & Mesirov, J.P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013).
Saeed, A.I. et al. TM4: a free, open-source system for microarray data management and analysis. Biotechniques 34, 374–378 (2003).
Huang, W., Sherman, B.T. & Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Saikali, P. et al. NKG2D-mediated cytotoxicity toward oligodendrocytes suggests a mechanism for tissue injury in multiple sclerosis. J. Neurosci. 27, 1220–1228 (2007).
Durafourt, B.A. et al. Comparison of polarization properties of human adult microglia and blood-derived macrophages. Glia 60, 717–727 (2012).
Acknowledgements
We thank R. Van Bruggen, S. Gauthier, P. D'Arcy and G. Perreault for assistance in the ENU project, and C. Meunier for technical assistance. Supported by the Canadian Institutes of Health Research (MOP119342 and MOP133487 to P.G. and S.V.) and Amorchem (PT63088).
Author information
Authors and Affiliations
Contributions
S.T., M.L., J.M. and J.B. contributed to mutation identification. S.T. performed all of the PbA experiments. M.J.P. performed all of the EAE experiments. S.T. and M.J.P. performed biochemical work. I.R., J.M.K. and S.T. performed the immunophenotyping experiments. D.L. performed RNA sequencing analyses, and S.T. carried out validations by RT-qPCR. Primary cells from brain were provided and characterized by J.A., N.A., A.P., M.J.P. and L.M.H., with additional contribution from G.A.L.-T., S.I., K.M., C.L. and K.P.K. kindly provided gene knockout animals. S.T. performed Listeria experiments with guidance from C.M.K. N.F. performed BM transplant experiments. J.B., J.M. and M.L. performed analyses of exome sequences. P.G. and S.M.V. supervised the project, helped to design experiments and analyzed data. P.G., S.T., D.L., M.J.P. and N.F. wrote the first draft of the manuscript. All of the authors provided helpful comments on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Ubiquitous pattern of Usp15 mRNA expression in embryonic, post-natal and adult mice
Mouse sections were stained with cresyl violet to localize Usp15 RNA to specific organs and structures. In situ hybridization was carried out using radiolabelled antisense (as) and sense (s) probes. The results shown are from X-ray film autoradiography obtained following 5-days exposure. Non-specific localized signals (visible with sense and anti-sense probes) are indicated with an asterisk (*); in the teeth (p10) and the large intestine lumen (p10 and adult). (Magnification: Embryonic x2.4, Post-natal x3, Adult x2.4). Abbreviations: Adr–adrenal gland; At–heart atrium; Br–brain; Bro–bronchcus; Car– cartilage; Cb–cerebellum; Co–colon; Cx–cerebral cortex; Du – duodenum; E – eye; Ep – epididymis; Es – esophagus; GB – gallbladder; HV–heart ventricle; Il–ileum; Je–jejunum; Ki–kidney; Li–liver; LI–large intestine; Lu–lung; OL–olfactory lobe; Ov–ovary; Ovi–oviducts; PB–pelvis bone; Pc–pancreas; PG–pituitary gland; Pr–prostate; PTh–parathyroid gland; R–ribs; Sk–skin; Spl–spleen; St–stomach; SV–seminal vesicle; Te–testis; Th–thyroid gland; UB–urinary bladder; Ut-uterus; CA–central artery; GC-germinal center; LN–lymphatic nodule; RP–red pulp; Tr–trabeculum; V-vein; LF–lymphoid follicle; Me–medulla; MG–mammary glands; Cx–cortex.
Supplementary Figure 2 Reduced protein stability of the USP15 L720R human variant in vitro
HEK293 cells stably expressing HA-tagged WT or USP15L720R proteins were treated with cycloheximide (CHX, 20 μg/ml) for 2, 4, 8, and 16h, and equal amounts of protein were analyzed by immunoblotting. Data is from a single experiment.
Supplementary Figure 3 Immunophenotyping of Usp15L749R mutants at steady-state and following P. berghei ANKA infection
(a) The number and proportions of different spleen cell types from naïve and from day 5-PbA infected animals, were established by flow cytometry with markers for T cells (CD4, CD8), B cells (B220), NK cells (NK1.1), monocytes and neutrophils (CD11b, Ly6G). Results are pooled from 5 independent experiments. (b) The percentage of splenic CD4+ and CD8+ effector T cells (CD62L−CD44+) were also assessed. Data represents a single experiment with 5 mice per group, and are expressed as a mean ± SD. (c-d) Cells were re-stimulated in vitro with either media alone (unstimulated, US), with anti-CD3 and anti-CD28 (TCR engagement), with PMA/Ionomycin, with CpG, or with Poly:IC and cytokine production was assessed by flow cytometry (C, intracellular staining), or by ELISA (D, culture supernatants). Data is a representation of two independent experiments with 5 mice per group, and is expressed as a mean ± SD. (e) The activation state of CD4+ and CD8+ T cells were assessed by analysis of CD69 cell surface expression in response to TCR engagement (anti-CD3/anti-CD28). Data represents a single experiment with 5 mice per group, and is expressed as a mean ± SD.
Supplementary Figure 4 USP15 negatively regulates CD4+ T cell activation during Listeria monocytogenes infection
Wild type B6 mice and Usp15L749R mutants infected with 1x104 CFU of Listeria monocytogenes (strain 10403s) expressing ovalbumin (OVA) were sacrificed on day 7 post-infection, and phenotyped for the activation of the T cell response in spleen cell populations. (a, b) CD44 expression (T cell activation) on CD4+ T cells (A), or CD8+ T cells (B), expressed as percentage and total cell numbers. (c, d) Cells were re-stimulated in vitro with Listeria-specific antigens, LLO or OVA, and IFN-γ production was assessed by flow cytometry (C, intracellular staining), or by ELISA (D, culture supernatants) for CD4+ and CD8+ T cells. (e) Serum IFN-γ levels were measured by ELISA, and plotted as optical density absorbance (OD) at 450 nm. (a-e) Data is a combination of two independent experiments. All data are expressed as a mean ± SD for each group, and all statistical analyses were performed using the two-tailed unpaired Student’s t-test.
Supplementary Figure 5 Cell populations and associated molecular pathways differentially regulated in a USP15-dependent fashion
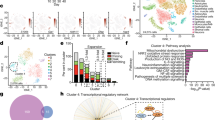
(a) LEA dendogram for genes with reduced expression in Usp15L749R mutant mice compared to B6 (day 5 post-PbA infection) and that drive significant enrichment (FDR<0.01) of immunological expression signatures (GSEA). Enriched immunological signatures and functions are highlighted by color boxes: red = signatures of IFN activation, green = myeloid signatures and responses, and purple = T cell signatures. Refer to Online Methods for details on LEA analysis. (b) LEA clustering analysis as described in (A) for immunological signatures depleted in Usp15L749R mutant mice during EAE neuroinflammation progression.
Supplementary Figure 6 Mouse mutants bearing a loss of function mutation in Irf3 are protected against neuroinflammation
Survival plots for PbA-infected (a) Irf3 mutants (Irf3-/-) (n=13) and B6 controls (n=8), and (b) Mavs mutants (Mavs-/-) (n=22) and B6 (n=11). Statistical significance for survival between groups of mice was determined by the Log-rank test (* P<0.05, **** P<0.0001).
Supplementary Figure 7 The L720R mutation affects the ability of USP15 to deubiquitinate SMURF2
HEK293T cells transiently expressing Smurf2-FLAG with or without Usp15-Xpress construct plus HA-tagged ubiquitin (Ub) were lysed 48h post-transfection, SMURF2 was immunoprecipitated using anti-FLAG antibody and immunoblotting analysis was performed as indicated. Construct expression in whole cell lysates (WCL) was confirmed by western blot
Supplementary Figure 8 Full gating strategy for flow cytometry analysis of brain cellular infiltration
Five days post-PbA infection, infiltrating cells were isolated from perfused brains and stained for flow cytometry analyses. Viable cells were selected for based on their exclusion of the Zombie Aqua Dye. Leukocytes were gated as CD45hi cells. Populations of leukocytes gated from the CD45hi gate were analyzed as follows: CD4 T cells (TCRb+CD4+CD8−), CD8 T cells (TCRb+CD4−CD8+), macrophages (Cd11b+F4/80+) and neutrophils (Cd11b+Ly6G+).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 (PDF 1128 kb)
Supplementary Table 1
List of dys-regulated genes in Usp15L749R mutant mice undergoing ECM and EAE models of neuroinflammatory diseases, related to Fig. 4b. (XLSX 120 kb)
Supplementary Table 2
Detailed matrix of leading edge analysis clustering performed on ECM and EAE depleted gene signatures in Usp15L749R mutant mice, related to Fig. 5a. (XLSX 462 kb)
Supplementary Table 3
qPCR validation primers, related to Fig. 5c-f. (XLSX 780 kb)
Rights and permissions
About this article
Cite this article
Torre, S., Polyak, M., Langlais, D. et al. USP15 regulates type I interferon response and is required for pathogenesis of neuroinflammation. Nat Immunol 18, 54–63 (2017). https://doi.org/10.1038/ni.3581
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3581
This article is cited by
-
RNA sequencing reveals differential long noncoding RNA expression profiles in bacterial and viral meningitis in children
BMC Medical Genomics (2024)
-
TRIM21/USP15 balances ACSL4 stability and the imatinib resistance of gastrointestinal stromal tumors
British Journal of Cancer (2024)
-
CCDC88B interacts with RASAL3 and ARHGEF2 and regulates dendritic cell function in neuroinflammation and colitis
Communications Biology (2024)
-
Downregulation of Ubiquitin-Specific Protease 15 (USP15) Does Not Provide Therapeutic Benefit in Experimental Mesial Temporal Lobe Epilepsy
Molecular Neurobiology (2024)
-
Association of ZC3HAV1 single nucleotide polymorphisms with the susceptibility of Vogt-Koyanagi-Harada Disease
BMC Medical Genomics (2023)