Abstract
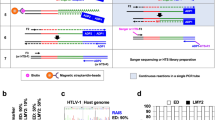
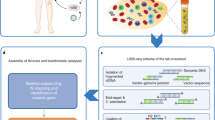
Retroviral vectors have induced subtle clonal skewing in many gene therapy patients and severe clonal proliferation and leukemia in some of them, emphasizing the need for comprehensive integration site analyses to assess the biosafety and genomic pharmacokinetics of vectors and clonal fate of gene-modified cells in vivo. Integration site analyses such as linear amplification–mediated PCR (LAM-PCR) require a restriction digest generating unevenly small fragments of the genome. Here we show that each restriction motif allows for identification of only a fraction of all genomic integrants, hampering the understanding and prediction of biological consequences after vector insertion. We developed a model to define genomic access to the viral integration site that provides optimal restriction motif combinations and minimizes the percentage of nonaccessible insertion loci. We introduce a new nonrestrictive LAM-PCR approach that has superior capabilities for comprehensive unbiased integration site retrieval in preclinical and clinical samples independent of restriction motifs and amplification inefficiency.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Aiuti, A. et al. Correction of ADA-SCID by stem cell gene therapy combined with nonmyeloablative conditioning. Science 296, 2410–2413 (2002).
Cavazzana-Calvo, M. et al. Gene therapy of human severe combined immunodeficiency (SCID)-X1 disease. Science 288, 669–672 (2000).
Flotte, T.R. Gene therapy: the first two decades and the current state-of-the-art. J. Cell. Physiol. 213, 301–305 (2007).
Gaspar, H.B. et al. Gene therapy of X-linked severe combined immunodeficiency by use of a pseudotyped gammaretroviral vector. Lancet 364, 2181–2187 (2004).
Hacein-Bey-Abina, S. et al. LMO2-associated clonal T cell proliferation in two patients after gene therapy for SCID-X1. Science 302, 415–419 (2003).
Howe, S. et al. Insertional mutagenesis in combination with acquired somatic mutations leads to leukemogenesis following gene therapy of SCID-X1. J. Clin. Invest. 118, 3143–3150 (2008).
Ott, M.G. et al. Correction of X-linked chronic granulomatous disease by gene therapy, augmented by insertional activation of MDS1-EVI1, PRDM16 or SETBP1. Nat. Med. 12, 401–409 (2006).
Aiuti, A. et al. Multilineage hematopoietic reconstitution without clonal selection in ADA-SCID patients treated with stem cell gene therapy. J. Clin. Invest. 117, 2233–2240 (2007).
Bohne, J. & Cathomen, T. Genotoxicity in gene therapy: An account of vector integration and designer nucleases. Curr. Opin. Mol. Ther. 10, 214–223 (2008).
Deichmann, A. et al. Vector integration is nonrandom and clustered and influences the fate of lymphopoiesis in SCID-X1 gene therapy. J. Clin. Invest. 117, 2225–2232 (2007).
Schwarzwaelder, K. et al. Gammaretrovirus-mediated correction of SCID-X1 is associated with skewed vector integration site distribution in vivo. J. Clin. Invest. 117, 2241–2249 (2007).
Schmidt, M. et al. High-resolution insertion-site analysis by linear amplification-mediated PCR (LAM-PCR). Nat. Methods 4, 1051–1057 (2007).
Schmidt, M. et al. Polyclonal long-term repopulating stem cell clones in a primate model. Blood 100, 2737–2743 (2002).
Mueller, P.R. & Wold, B. In vivo footprinting of a muscle-specific enhancer by ligation mediated PCR. Science 246, 780–786 (1989).
Silver, J. & Keerikatte, V. Novel use of polymerase chain reaction to amplify cellular DNA adjacent to an integrated provirus. J. Virol. 63, 1924–1928 (1989).
Harkey, M.A. et al. Multiarm high-throughput integration site detection: limitations of LAM-PCR technology and optimization for clonal analysis. Stem Cells Dev. 16, 381–392 (2007).
Wang, G.P. et al. DNA bar coding and pyrosequencing to analyze adverse events in therapeutic gene transfer. Nucleic Acids Res. 36, e49 (2008).
International Human Genome Sequencing Consortium. Finishing the euchromatic sequence of the human genome. Nature 431, 931–945 (2004).
Bovee, D. et al. Closing gaps in the human genome with fosmid resources generated from multiple individuals. Nat. Genet. 40, 96–101 (2008).
Lander, E.S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Kent, W.J. BLAT––the BLAST-like alignment tool. Genome Res. 12, 656–664 (2002).
Mikkers, H. et al. High-throughput retroviral tagging to identify components of specific signaling pathways in cancer. Nat. Genet. 32, 153–159 (2002).
Modlich, U. et al. Cell-culture assays reveal the importance of retroviral vector design for insertional genotoxicity. Blood 108, 2545–2553 (2006).
Montini, E. et al. Hematopoietic stem cell gene transfer in a tumor-prone mouse model uncovers low genotoxicity of lentiviral vector integration. Nat. Biotechnol. 24, 687–696 (2006).
Suzuki, T. et al. New genes involved in cancer identified by retroviral tagging. Nat. Genet. 32, 166–174 (2002).
Altschul, S.F., Gish, W., Miller, W., Myers, E.W. & Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Gotoh, O. An improved algorithm for matching biological sequences. J. Mol. Biol. 162, 705–708 (1982).
Smith, T.F. & Waterman, M.S. Identification of common molecular subsequences. J. Mol. Biol. 147, 195–197 (1981).
Acknowledgements
We thank I. Kutschera, N. Krenzer, C. Lulay, S. Braun and D. Glagow for technical assistance and O. Danos for fruitful discussions about the model. Funding was provided by the Deutsche Forschungsgemeinschaft (SPP1230, grant of the Tumor Center Heidelberg/Mannheim), by the Bundesministerium für Bildung und Forschung (iGene), by the VIth + VIIth Framework Programs of the European Commission (Concerted Safety & Efficiency Evaluation of Retroviral Transgenesis in Gene Therapy of Inherited Diseases (CONSERT), European Network for the Advancement of Clinical Gene Transfer and Therapy (CLINIGENE) and Persisting Transgenesis (PERSIST)) and by the Initiative and Networking Fund of the Helmholtz Association within the Helmholtz Alliance on Immunotherapy of Cancer.
Author information
Authors and Affiliations
Contributions
R.E., C.v.K., W.S. and M.S. conceived of the genome accessibility model and interpreted data. R.E. and W.S. conducted bioinformatics analyses. C.v.K. and M.S., R.G., A.P. and H.G. developed the concept of nrLAM and designed experiments. R.G., A.P., A.N., C.C.B. and C.R.B. performed experiments. W.W. provided lentiviral vectors. D.C., E.M., L.N., L.B., A. Aiuti, O.C.-H., K.S.B., R.J.Y.-M., R.R.A., A.R., C.C. and F.M. provided retroviral integration site data sets; C.C.B. and A. Arens performed LAM-PCR and generated the data on these samples. C.C.B., K.S., A. Arens, K.F. and A.D. generated LAM data on X-SCID samples provided by S.J.H., H.B.G. and A.J.T. R.G., R.E., A.P., C.C.B., R.K., W.S., C.v.K. and M.S. prepared and wrote the manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4, Supplementary Tables 1–9 and Supplementary Methods (PDF 1044 kb)
Rights and permissions
About this article
Cite this article
Gabriel, R., Eckenberg, R., Paruzynski, A. et al. Comprehensive genomic access to vector integration in clinical gene therapy. Nat Med 15, 1431–1436 (2009). https://doi.org/10.1038/nm.2057
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.2057
This article is cited by
-
TET2 guards against unchecked BATF3-induced CAR T cell expansion
Nature (2023)
-
Biodistribution of lentiviral transduced adipose-derived stem cells for “ex-vivo” regional gene therapy for bone repair
Gene Therapy (2023)
-
RAISING is a high-performance method for identifying random transgene integration sites
Communications Biology (2022)
-
A high-throughput detection method for the clonality of Human T-cell leukemia virus type-1-infected cells in vivo
International Journal of Hematology (2020)
-
Successful engraftment of gene-corrected hematopoietic stem cells in non-conditioned patients with Fanconi anemia
Nature Medicine (2019)