Abstract
The worldwide diversity of HIV-1 presents an unprecedented challenge for vaccine development1,2. Antigens derived from natural HIV-1 sequences have elicited only a limited breadth of cellular immune responses in nonhuman primate studies and clinical trials to date. Polyvalent 'mosaic' antigens, in contrast, are designed to optimize cellular immunologic coverage of global HIV-1 sequence diversity3. Here we show that mosaic HIV-1 Gag, Pol and Env antigens expressed by recombinant, replication-incompetent adenovirus serotype 26 vectors markedly augmented both the breadth and depth without compromising the magnitude of antigen-specific T lymphocyte responses as compared with consensus or natural sequence HIV-1 antigens in rhesus monkeys. Polyvalent mosaic antigens therefore represent a promising strategy to expand cellular immunologic vaccine coverage for genetically diverse pathogens such as HIV-1.
Similar content being viewed by others
Main
The development of vaccine strategies that expand cellular immune breadth will be crucial for achieving immunologic coverage of the enormous global genetic diversity of HIV-11,2. Moreover, the breadth of Gag-specific cellular immune responses has been shown to correlate with control of HIV-1 replication in humans4 and control of simian immunodeficiency virus (SIV) challenges in vaccinated rhesus monkeys5. Polyvalent mosaic proteins are assembled from natural sequences by in silico recombination and are optimized to provide maximal coverage of potential T cell epitopes (PTEs) for a given valency3. Mosaic antigens are full-length proteins that are designed to preserve natural antigen expression and processing. We used a bivalent mosaic strategy consisting of two HIV-1 Gag, Pol and Env antigens to balance the competing issues of theoretical coverage and practical utility. Here we report the breadth and magnitude of epitope-specific CD8+ and CD4+ T lymphocyte responses elicited by mosaic, consensus and natural sequence HIV-1 antigens in rhesus monkeys.
We immunized 27 outbred rhesus monkeys with a single injection of recombinant adenovirus serotype 26 (rAd26) vectors6 expressing the following antigens: bivalent mosaic (n = 7), M consensus7 (n =7), bivalent combined clade B + clade C (n = 7) or optimal natural clade C (n = 6) HIV-1 Gag, Pol and Env antigens. We administered a total dose of 3 × 1010 viral particles of rAd26 vectors expressing these antigens once intramuscularly (i.m.) to each monkey. The optimal clade C antigens were the natural strain sequences selected to provide maximal PTE coverage of clade C sequences (Methods). We assessed the breadth and magnitude of vaccine-elicited HIV-1–specific T lymphocyte responses by interferon-γ (IFN-γ) enzyme-linked immunospot (ELISPOT) assays at week 4 after immunization using pools and subpools of peptides that included all global PTEs found in at least 15% of HIV-1 M group sequences8. We resolved all individual peptide responses and performed cell-depleted IFN-γ ELISPOT assays to determine whether the reactive peptides represented CD8+ or CD4+ T lymphocyte epitopes.
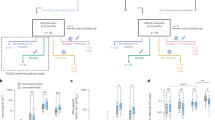
The total number of Gag-, Pol- and Env-specific cellular immune responses to PTE peptides elicited by the mosaic antigens was 3.8-fold higher than the number of responses induced by the consensus or natural sequence antigens (Fig. 1a; P = 1 × 10−11, comparing the mosaic with the consensus antigens, the next highest group, on the basis of a Poisson regression model9; http://cran.r-project.org/web/packages/lme4/ and http://www.R-project.org/). There were 4.4-fold more CD8+ than CD4+ T lymphocyte responses (P < 10−11) and fewer responses to Env than to Gag or Pol (Fig. 1a; P < 0.0007). The median number of CD8+ T lymphocyte responses was highest for the mosaic vaccine, followed by the consensus, the combined B + C and the natural clade C vaccines (Fig. 1a; medians of 16, 5, 3 and 2 responses per monkey in each group, respectively). Although there were fewer CD4+ T lymphocyte responses overall, the same relative pattern emerged with the highest number of CD4+ T lymphocyte responses to the mosaic vaccine, followed by the consensus, the combined B + C and the natural clade C vaccines (Fig. 1a; medians of 4, 1, 1 and 0.5 responses per monkey in each group, respectively). The best fitting Poisson regression model indicated similar relative degrees of augmentation for CD8+ and CD4+ T lymphocyte responses induced by the mosaic vaccine as compared with the other vaccines but greater absolute numbers of CD8+ T lymphocyte epitopes. T lymphocyte responses elicited by the consensus, the combined B + C and the natural clade C vaccines were not statistically distinguishable.
(a) Numbers of epitope-specific CD4+ (top) and CD8+ (bottom) T lymphocyte responses to individual PTE peptides are shown after a single immunization of rAd26 vectors expressing mosaic, M consensus, clade B + clade C or optimal natural clade C HIV-1 Gag, Pol and Env antigens. Individual monkeys are depicted on the x axis. The various shades of each color reflect responses to the different antigens (Gag, Pol and Env), as indicated. (b) Numbers of CD4+ (top) and CD8+ (bottom) T lymphocyte response regions. (c) Magnitude of all Gag-, Pol- and Env-specific CD8+ (top and middle) and CD4+ (bottom) T lymphocyte responses arranged from lowest to highest. Spot-forming cells (SFCs) per 1 × 106 PBMCs are shown for each epitope-specific response.
PTE peptides include multiple overlapping sequences that reflect naturally occurring HIV-1 polymorphisms8, and thus PTE peptide responses encompass both the recognition of a particular epitope (breadth) and the cross-recognition of variants of that epitope (depth). We performed a conservative analysis of breadth by assessing the number of reactive epitopic regions per monkey in which all reactive PTE peptides that overlapped by eight or more amino acids were counted as one event. In this conservative analysis, we observed that the mosaic antigens elicited 3.1-fold higher numbers of Gag-, Pol- and Env-specific epitopic regions as compared with the consensus antigens or natural sequence antigens (Fig. 1b; P = 1.6 × 10−7, Poisson regression). The median number of CD8+ T lymphocyte epitopic regions was higher for the mosaic vaccine (median of 8 response regions, range 7–14) than the other vaccines (median of 2 response regions, range 0–6). Epitopes showed some clustering across monkeys, as evidenced by regions of high epitope density (Supplementary Figs. 1 and 2). Complete alignments of all responses are also available (Supplementary Fig. 3).
These data show that the mosaic antigens substantially increased cellular immune breadth as compared with the consensus or the natural sequence antigens. In addition to evaluating breadth, we also assessed the magnitude of all individual CD8+ and CD4+ T lymphocyte responses. The magnitude of these responses proved comparable among all groups (Fig. 1c; P = 0.58 and P = 0.99, respectively, two-sided Kolmogorov-Smirnov tests), demonstrating that the mosaic antigens expanded cellular immune breadth without compromising the magnitude of individual epitope-specific responses.
We next characterized the depth of the cellular immune responses elicited by the various vaccine regimens. We defined depth as the number of simultaneously elicited variant PTE peptides for a particular epitopic region. Inducing responses to multiple epitope variants may increase immunologic coverage of infecting virus sequences and may also block common escape routes in vivo or force the virus down suboptimal escape routes that incur high fitness costs3. The consensus and natural sequence antigens elicited responses that were characterized by a high degree of sequence identity between the vaccine sequences and the reactive PTE peptides, as exemplified by the responses in monkey 366 (Fig. 2a and Supplementary Fig. 3). In contrast, the mosaic antigens elicited responses that were characterized by multiple reactive PTE peptides in particular epitopic regions. These peptides represented common variants and typically reflected the polymorphisms contained in the mosaic vaccine sequences, as exemplified by the responses in monkey 361 (Fig. 2b and Supplementary Fig. 3). A summary analysis shows that the mosaic antigens elicited a higher frequency of cellular immune responses to peptides with two or more targeted variants as compared with the consensus or natural sequence antigens (Fig. 2c; P = 0.001, Wilcoxon rank-sum test comparing the mosaic with the other groups).
(a) Example of mapped T lymphocyte responses in monkey 366, which received the optimal natural clade C antigens. (b) Example of mapped T lymphocyte responses in monkey 361, which received the bivalent mosaic antigens. In a and b, vaccine sequences are shown on the top and are designated OptC (optimal natural clade C), Mos1 (mosaic sequence 1) and Mos2 (mosaic sequence 2), and reactive PTE peptides are shown beneath the vaccine sequences denoted by the antigen (G, Gag; P, Pol; E, Env) and the PTE peptide number. Sequence polymorphisms between the two mosaic antigens are shown in blue. Differences between the vaccine sequences and the reactive PTE peptides are shown in red. Complete alignments of all positive peptides organized by response regions are shown in Supplementary Figure 3. (c) Depth of CD4+ (top) and CD8+ (bottom) T lymphocyte responses after immunization with rAd26 vectors expressing mosaic, M consensus, clade B + clade C or optimal natural clade C antigens. Individual monkeys are depicted on the x axis. One response variant or more than one response variants shown for each epitopic region.
To complement the analysis with PTE peptides, we also assessed the breadth of cellular immune responses in the vaccinated monkeys using overlapping peptides covering five HIV-1 Gag sequences: clade C DU422, clade C ZM651, consensus C, consensus A and consensus B. We determined cellular immune breadth by assessing reactivity to subpools of ten overlapping peptides spanning each Gag sequence. The mosaic antigens elicited a greater breadth of T lymphocyte responses as compared with the consensus or natural sequence antigens against all five Gag sequences (Fig. 3; P = 1 × 10−7, binomial regression). In particular, the mosaic antigens proved superior to the optimal natural clade C antigens for inducing responses against clade C Gag (Fig. 3). The mosaic antigens also elicited comparable responses to Gag from multiple clades, whereas the natural clade C antigens showed diminished responses to clade A Gag and clade B Gag. The breadth of Gag-specific responses elicited by the consensus, the combined B + C and the natural clade C vaccines were not statistically distinguishable, consistent with the results with PTE peptides (Fig. 1).
Breadth of cellular immune responses was assessed with subpools of overlapping peptides spanning the following strains of HIV-1 Gag: clade C DU422, clade C ZM651, consensus C, consensus A and consensus B. Numbers of positive subpools are shown after a single immunization of rAd26 vectors expressing mosaic, M consensus, clade B + clade C or optimal natural clade C HIV-1 Gag, Pol and Env antigens. Individual monkeys are depicted on the x axis.
To assess the durability of these observations, we boosted the monkeys that received the mosaic, consensus and optimal natural clade C antigens at week 40 with a total dose of 3 × 1010 viral particles of the heterologous vector rAd5HVR48 (ref. 10) expressing HIV-1 Gag, Pol and Env antigens that matched the initial immunization sequences. We determined cellular immune breadth by assessing reactivity to subpools of ten PTE peptides at week 4 after the prime and at week 44 after the boost. The majority of CD8+ and CD4+ T lymphocyte responses that we observed after the priming immunization expanded after the boost, although we also detected new responses (Fig. 4a). At week 44, the frequency of high-magnitude, epitope-specific cellular immune responses was comparable among groups (Fig. 4a), but the number of subpool responses elicited by the mosaic antigens (median 27 responses per monkey) remained substantially higher than the number induced by the consensus antigens (median 11 responses per monkey) or the optimal natural clade C antigens (median 10 responses per monkey) (Fig. 4b; P < 0.001, Wilcoxon rank-sum tests for all pair-wise comparisons).
(a,b) Magnitude (a) and breadth (b) of T lymphocyte subpool responses at week 4 after prime (left data point) and at week 44 after boost (right data point) for each monkey. Monkeys were primed at week 0 with rAd26 vectors and boosted at week 40 with rAd5HVR48 vectors expressing mosaic, M consensus or optimal natural clade C HIV-1 Gag, Pol and Env antigens. Individual monkeys are depicted on the x axis. In a, red denotes epitope-specific CD8+ T lymphocyte responses, blue denotes epitope-specific CD4+ T lymphocyte responses, lines depict responses observed at both time points and dots depict responses observed at only one time point. (c) Env-specific ELISA end-point titers are shown at weeks 0, 4 and 44. (d) NAb titers to the tier 1 clade A (DJ263.8), clade B (SF162.LS) and clade C (MW965.26) viruses at week 44. NAb titers to murine leukemia virus as a negative control were <20 for all samples (data not shown).
We also measured Env-specific humoral immune responses after the boost immunization by ELISAs (Fig. 4c) and luciferase-based pseudovirus neutralization assays11 (Fig. 4d). All groups showed comparable ELISA titers to clade C gp140 and comparable neutralizing antibody (NAb) responses to the tier 1 clade C virus MW965.26 (Fig.4d). The mosaic antigens elicited slightly higher NAb responses to the tier 1 clade B virus SF162.LS as compared with the consensus or natural clade C antigens (Fig. 4d; P = 0.02, Wilcoxon rank-sum test). NAb responses to the tier 1 clade A virus DJ263.8 were low in all groups (Fig. 4d), and we did not detect any NAb responses to tier 2 viruses (data not shown). Thus, the mosaic Env immunogens elicited noninferior antibody responses as compared with the consensus or natural sequence Env antigens.
Our data demonstrate that mosaic HIV-1 Gag, Pol and Env antigens augment both the breadth and depth of epitope-specific cellular immune responses as compared with consensus or natural sequence antigens in rhesus monkeys, in agreement with theoretical predictions (Supplementary Fig. 4). In particular, a bivalent set of mosaic antigens proved superior to a bivalent set of clade B + clade C antigens, indicating that the augmented cellular immune breadth and depth was dependent on the mosaic antigen design3,12, which not only maximized inclusion of common epitope variants but also excluded rare epitope variants that are unlikely to be useful in a vaccine. The results with the mosaic antigens in this study probably reflected the efficiency of rAd26 vectors in eliciting CD8+ T lymphocyte responses5,6, together with the capacity of the mosaic antigens to augment CD8+ T lymphocyte breadth. The mosaic antigens also substantially enhanced CD4+ T lymphocyte breadth, but these effects were less marked and were characterized by lower total numbers of CD4+ T lymphocyte responses.
The importance of expanding cellular immune breadth has been demonstrated by recent studies showing that the breadth of Gag-specific T lymphocyte responses correlates with control of HIV-1 replication in chronically infected humans4 and control of SIV replication after challenge of vaccinated rhesus monkeys5. In the phase 2b STEP study, the rAd5-based HIV-1 vaccine candidate expressing natural clade B Gag, Pol and Nef antigens elicited only a limited breadth of cellular immune responses, and no vaccine benefit was observed13,14. This narrow breadth of cellular immune responses probably provided insufficient immunologic coverage of the diversity of infecting viruses. Moreover, viral escape from CD8+ T lymphocytes has been reported to occur during acute HIV-1 infection15, and cellular immune responses against variant epitopes have been shown to block SIV mutational evolution in rhesus monkeys in vivo16, thus suggesting a key role of expanding cellular immune depth. We were unable to assess the protective efficacy of mosaic HIV-1 vaccines in the present study as a result of the lack of an appropriate preclinical HIV-1 challenge model. Modeling the protective efficacy of mosaic vaccines against SIV challenges in nonhuman primates also has intrinsic limitations, as the observed diversity of SIV in natural hosts and HIV-1 in humans differs substantially and is influenced by different underlying biology17,18.
In summary, this study shows that bivalent mosaic HIV-1 Gag, Pol and Env antigens expand cellular immune breadth and depth in rhesus monkeys as compared with consensus or natural sequence antigens. These findings have major implications for HIV-1 vaccine development, as global virus diversity and viral escape from cellular immune responses represent major hurdles in the development of a T cell–based HIV-1 vaccine. A bivalent cocktail of mosaic antigens is also practical and potentially feasible for clinical development. Moreover, the mosaic antigen strategy is generalizable and should be applicable for other genetically diverse pathogens in addition to HIV-1.
Methods
Antigen design.
We constructed bivalent mosaic Gag, Pol and Env antigens to provide optimal coverage of HIV-1 M group sequences in the Los Alamos HIV-1 Sequence Database essentially as previously described3,19, using the coverage of epitope length peptides in the global database as the selection criterion. We used the global sequence data set for each antigen, restricted to include only full-length proteins and one sequence per individual. We then fractured this set of proteins into all possible fragments of nine contiguous amino acids (nonamers). We tallied the nonamer frequencies and used a genetic algorithm to design and select in silico recombinant sequences that resembled real proteins and that maximized the number of nonamer sequence matches between the vaccine candidates and the global database. We selected the optimal clade C antigens with a similar selection strategy, but we used the clade C data set as the input sequences. We selected the single clade C Gag, Pol and Env natural sequences that provided optimal coverage of all nonamers in the global clade C data set (C.IN.-.70177 Gag, C.ZA.04.04ZASK208B1 Pol, C.SN.90.90SE_364 Env). The clade B antigens were previously selected to be near-consensus or consensus sequences with good coverage of the clade B data set (B.CAM-1 Gag, B.IIIB Pol, B.Con Env)20, and we used them to complement the optimal clade C antigens for the bivalent clade B + C vaccine approach. Pol antigens contained reverse transcriptase and integrase without protease and included point mutations to eliminate catalytic activity, as previously described20. Env gp140 antigens contained point mutations to eliminate cleavage and fusion activity. Vaccine sequences are depicted in Supplementary Figure 4.
Vector production.
We produced recombinant, replication-incompetent, E1/E3-deleted adenovirus serotype 26 (rAd26) and hexon-chimeric rAd5HVR48 vectors expressing these antigens and purified them essentially as previously described6,10. We expressed the HIV-1 antigens in the adenovirus E1 region under control of a human cytomegalovirus promoter.
Animals and immunizations.
We housed 27 outbred rhesus monkeys that did not express the major histocompatibility complex class I alleles Mamu-A*01, Mamu-B*08 and Mamu-B*17 at New England Primate Research Center. Immunizations involved 3 × 1010 viral particles of rAd26 or rAd5HVR48 vectors expressing mosaic, M consensus, clade B + clade C or optimal natural clade C HIV-1 Gag, Pol and Env antigens delivered as 1 ml injections i.m. in both quadriceps muscles at weeks 0 and 40. All monkey studies were approved by the Harvard Medical School and Beth Israel Deaconess Medical Center Institutional Animal Care and Use Committees.
Interferon-γ enzyme-linked immunosorbent spot assays.
We assessed HIV-1–specific cellular immune responses in vaccinated monkeys by IFN-γ ELISPOT assays, essentially as previously described5,10. We obtained HIV-1 Gag, Pol and Env potential T cell epitope (PTE) peptides, which included all PTEs found in at least 15% of HIV-1 M group sequences8, as well as HIV-1 Gag peptides from clade C DU422, clade C ZM651, consensus C, consensus A and consensus B strains, from the NIH AIDS Research and Reference Reagent Program. We calculated SFCs per 1 × 106 peripheral blood mononuclear cells. Media backgrounds were typically <15 SFCs per 1 × 106 peripheral blood mononuclear cells. We defined positive responses as >55 SFCs per 1 × 106 PBMCs and >4-fold background.
Epitope mapping.
We performed comprehensive CD8+ and CD4+ T lymphocyte epitope mapping with Gag, Pol and Env PTE peptides. We conducted IFN-γ ELISPOT assays at week 4 after immunization with complete peptide pools as well as with subpools containing ten PTE peptides that were pooled sequentially by peptide number. We then deconvoluted all peptide subpools with positive responses and confirmed epitopes with individual 15 amino acid PTE peptides. We performed cell-depleted IFN-γ ELISPOT assays to determine whether reactive peptides represented CD8+ or CD4+ T lymphocyte epitopes. We performed cell depletions by negative bead selection that was >98% efficient. We also performed partial epitope mapping of PTE subpools 4 weeks after the boost immunization at week 44. We retested all borderline responses and only considered them positive if confirmed. We also performed partial epitope mapping of subpools containing ten overlapping Gag peptides to assess breadth to HIV-1 Gag from various clades.
Humoral immune assays.
We evaluated Env-specific humoral immune responses by direct ELISAs with HIV-1 clade C Env gp140 and luciferase-based pseudovirus neutralization assays essentially as previously described11.
Statistical analyses.
We did all statistical analyses with the package R (http://www.R-project.org/). To analyze the breadth of cellular immune responses to mapped PTE peptides (Fig. 1a), we fit Poisson regression models that predicted the number of reactive peptides as a function of vaccine group, antigen (Gag, Pol or Env) and lymphocyte subpopulation (CD4 or CD8). Our models included random effects to accommodate monkey-to-monkey variation and fit them with the lme4 library (http://cran.r-project.org/web/packages/lme4/) of the package R. The data fit the models well (dispersion parameter 1.0), and there were no significant interactions among the three explanatory factors. For example, the 3.8-fold enhancement in the number of PTE peptides recognized by monkeys that received the mosaic antigens as compared to those that received the consensus or natural sequence antigens (Fig. 1a) applied equally to PTEs from Gag, Pol and Env and held for responses by CD8+ as well as CD4+ T lymphocytes. The analysis of the number of reactive epitopic regions (Fig. 1b) also included Poisson regression models with random effects and again fit well (dispersion parameter 0.87) without any significant interactions. We compared the magnitude of CD8+ and CD4+ T lymphocyte responses (Fig. 1c) with two-sided Kolmogorov-Smirnov tests. We also performed nonparametric tests to compare the breadth and depth of responses per monkey between the vaccines (Figs. 1a and 2c). We initially used Kruskal-Wallis tests to determine whether there was a difference among the four vaccine groups, and we then assessed all pair-wise comparisons between the four vaccine groups with Wilcoxon rank-sum tests. To analyze the breadth of responses to HIV-1 Gag from various sequences (Fig. 3), we fit the data to binomial regression models. These models used the vaccine group as an explanatory variable and included random effects to account for monkey-to-monkey and strain-to-strain variation.
References
Barouch, D.H. Challenges in the development of an HIV-1 vaccine. Nature 455, 613–619 (2008).
Korber, B.T., Letvin, N.L. & Haynes, B.F. T-cell vaccine strategies for human immunodeficiency virus, the virus with a thousand faces. J. Virol. 83, 8300–8314 (2009).
Fischer, W. et al. Polyvalent vaccines for optimal coverage of potential T-cell epitopes in global HIV-1 variants. Nat. Med. 13, 100–106 (2007).
Kiepiela, P. et al. CD8+ T-cell responses to different HIV proteins have discordant associations with viral load. Nat. Med. 13, 46–53 (2007).
Liu, J. et al. Immune control of an SIV challenge by a T-cell–based vaccine in rhesus monkeys. Nature 457, 87–91 (2009).
Abbink, P. et al. Comparative seroprevalence and immunogenicity of six rare serotype recombinant adenovirus vaccine vectors from subgroups B and D. J. Virol. 81, 4654–4663 (2007).
Santra, S. et al. A centralized gene-based HIV-1 vaccine elicits broad cross-clade cellular immune responses in rhesus monkeys. Proc. Natl. Acad. Sci. USA 105, 10489–10494 (2008).
Li, F. et al. Peptide selection for human immunodeficiency virus type 1 CTL-based vaccine evaluation. Vaccine 24, 6893–6904 (2006).
Pinheiro, J.C. & Bates, D.M. Mixed-Effects Models in S and S-plus (Springer, New York, 2000).
Roberts, D.M. et al. Hexon-chimaeric adenovirus serotype 5 vectors circumvent pre-existing anti-vector immunity. Nature 441, 239–243 (2006).
Montefiori, D. Evaluating neutralizing antibodies against HIV, SIV and SHIV in luciferase reporter gene assays. Curr. Protoc. Immunol. 64, 12.1.1–12.1.17 (2004).
Kong, W.P. et al. Expanded breadth of the T-cell response to mosaic human immunodeficiency virus type 1 envelope DNA vaccination. J. Virol. 83, 2201–2215 (2009).
Buchbinder, S.P. et al. Efficacy assessment of a cell-mediated immunity HIV-1 vaccine (the STEP Study): a double-blind, randomised, placebo-controlled, test-of-concept trial. Lancet 372, 1881–1893 (2008).
McElrath, M.J. et al. HIV-1 vaccine–induced immunity in the test-of-concept STEP Study: a case-cohort analysis. Lancet 372, 1894–1905 (2008).
Goonetilleke, N. et al. The first T cell response to transmitted/founder virus contributes to the control of acute viremia in HIV-1 infection. J. Exp. Med. 206, 1253–1272 (2009).
Barouch, D.H. et al. Dynamic immune responses maintain cytotoxic T lymphocyte epitope mutations in transmitted simian immunodeficiency virus variants. Nat. Immunol. 6, 247–252 (2005).
Barry, A.P. et al. Depletion of CD8+ cells in sooty mangabey monkeys naturally infected with simian immunodeficiency virus reveals limited role for immune control of virus replication in a natural host species. J. Immunol. 178, 8002–8012 (2007).
Sodora, D.L. et al. Toward an AIDS vaccine: lessons from natural simian immunodeficiency virus infections of African nonhuman primate hosts. Nat. Med. 15, 861–865 (2009).
Thurmond, J. et al. Web-based design and evaluation of T-cell vaccine candidates. Bioinformatics 24, 1639–1640 (2008).
Priddy, F.H. et al. Safety and immunogenicity of a replication-incompetent adenovirus type 5 HIV-1 clade B gag/pol/nef vaccine in healthy adults. Clin. Infect. Dis. 46, 1769–1781 (2008).
Acknowledgements
We thank B. Haynes, N. Letvin, P. Swanson, F. Stephens, B. Hahn, J. McElrath, J. Goudsmit, M. Pau, N. Michael, M. Marovich and M. Pensiero for generous advice and assistance. We obtained peptides from the US National Institutes of Health (NIH) AIDS Research and Reference Reagent Program. We acknowledge support from NIH grants AI058727 (D.H.B.), AI066305 (D.H.B.), AI066924 (D.H.B.), AI078526 (D.H.B.), AI084794 (D.H.B.), RR000168, AI067854 (B.K.), AI061734 (B.K.) and directed research funding from Los Alamos National Laboratory (B.K.).
Author information
Authors and Affiliations
Contributions
W.F., B.K. and D.H.B. designed the antigens. S.L.K., P.A., L.F.M., Y.-H.S. and D.H.B. generated the vaccine vectors. K.L.O., N.L.S., A.L.P., A.M.R., D.M.L., S.L.C. and D.H.B. designed and conducted the cellular immunologic assays. A.L.P., K.B., J.R.P. and M.S.S. designed and conducted the humoral immunologic assays. A.C. and K.G.M. led the monkey work. M.M. and B.K. led the data analysis and J.J.S. wrote the response mapping software. D.H.B. and B.K. designed the study, and D.H.B. led the study. All authors contributed to the writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 (PDF 1086 kb)
Rights and permissions
About this article
Cite this article
Barouch, D., O'Brien, K., Simmons, N. et al. Mosaic HIV-1 vaccines expand the breadth and depth of cellular immune responses in rhesus monkeys. Nat Med 16, 319–323 (2010). https://doi.org/10.1038/nm.2089
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.2089
This article is cited by
-
Safety and immunogenicity of a mosaic vaccine booster against Omicron and other SARS-CoV-2 variants: a randomized phase 2 trial
Signal Transduction and Targeted Therapy (2023)
-
HIV envelope antigen valency on peptide nanofibers modulates antibody magnitude and binding breadth
Scientific Reports (2021)
-
New insights into pathogenesis point to HIV-1 Tat as a key vaccine target
Archives of Virology (2021)
-
Recombinant MVA-prime elicits neutralizing antibody responses by inducing antigen-specific B cells in the germinal center
npj Vaccines (2021)
-
Mathematical model of broadly reactive plasma cell production
Scientific Reports (2020)