Abstract
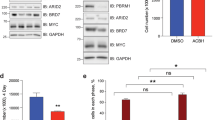
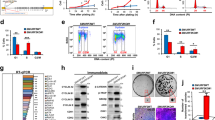
In addition to allelic mutations, cancers are known to harbor alterations in their chromatin landscape. Here we show that genomic ablation of Smad ubiquitin regulatory factor 2 (Smurf2), a HECT-domain E3 ubiquitin ligase, results in dysregulation of both the DNA damage response and genomic stability, culminating in increased susceptibility to various types of cancers in aged mice. We show that Smurf2 regulates the monoubiquitination of histone H2B as well as the trimethylation of histone H3 at Lys4 and Lys79 by targeting ring finger protein 20 (RNF20) for proteasomal degradation in both mouse and human cells. We also show that Smurf2 and RNF20 are colocalized at the γ-H2AX foci of double-stranded DNA breaks in the nucleus. Thus, Smurf2 has a tumor suppression function that normally maintains genomic stability by controlling the epigenetic landscape of histone modifications through RNF20.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Hershko, A. & Ciechanover, A. The ubiquitin system. Annu. Rev. Biochem. 67, 425–479 10.1146/annurev.biochem.67.1.425 (1998).
Izzi, L. & Attisano, L. Regulation of the TGF-β signalling pathway by ubiquitin-mediated degradation. Oncogene 23, 2071–2078 (2004).
Lönn, P., Moren, A., Raja, E., Dahl, M. & Moustakas, A. Regulating the stability of TGF-β receptors and Smads. Cell Res. 19, 21–35 (2009).
Zhang, Y., Chang, C., Gehling, D.J., Hemmati-Brivanlou, A. & Derynck, R. Regulation of Smad degradation and activity by Smurf2, an E3 ubiquitin ligase. Proc. Natl. Acad. Sci. USA 98, 974–979 (2001).
Lin, X., Liang, M. & Feng, X.H. Smurf2 is a ubiquitin E3 ligase mediating proteasome-dependent degradation of Smad2 in transforming growth factor-β signaling. J. Biol. Chem. 275, 36818–36822 (2000).
Kavsak, P. et al. Smad7 binds to Smurf2 to form an E3 ubiquitin ligase that targets the TGF-β receptor for degradation. Mol. Cell 6, 1365–1375 (2000).
Li, H. & Seth, A. An RNF11: Smurf2 complex mediates ubiquitination of the AMSH protein. Oncogene 23, 1801–1808 (2004).
Schwamborn, J.C., Muller, M., Becker, A.H. & Puschel, A.W. Ubiquitination of the GTPase Rap1B by the ubiquitin ligase Smurf2 is required for the establishment of neuronal polarity. EMBO J. 26, 1410–1422 (2007).
Narimatsu, M. et al. Regulation of planar cell polarity by Smurf ubiquitin ligases. Cell 137, 295–307 (2009).
Zhang, H. & Cohen, S.N. Smurf2 up-regulation activates telomere-dependent senescence. Genes Dev. 18, 3028–3040 (2004).
Fukuchi, M. et al. High-level expression of the Smad ubiquitin ligase Smurf2 correlates with poor prognosis in patients with esophageal squamous cell carcinoma. Cancer Res. 62, 7162–7165 (2002).
Jin, C. et al. Smad ubiquitination regulatory factor 2 promotes metastasis of breast cancer cells by enhancing migration and invasiveness. Cancer Res. 69, 735–740 (2009).
Johnstone, S.E. & Baylin, S.B. Stress and the epigenetic landscape: a link to the pathobiology of human diseases? Nat. Rev. Genet. 11, 806–812 (2010).
Kim, J., Hake, S.B. & Roeder, R.G. The human homolog of yeast BRE1 functions as a transcriptional coactivator through direct activator interactions. Mol. Cell 20, 759–770 (2005).
Zhu, B. et al. Monoubiquitination of human histone H2B: the factors involved and their roles in HOX gene regulation. Mol. Cell 20, 601–611 (2005).
Minsky, N. et al. Monoubiquitinated H2B is associated with the transcribed region of highly expressed genes in human cells. Nat. Cell Biol. 10, 483–488 (2008).
Moyal, L. et al. Requirement of ATM-dependent monoubiquitylation of histone H2B for timely repair of DNA double-strand breaks. Mol. Cell 41, 529–542 (2011).
Nakamura, K. et al. Regulation of homologous recombination by RNF20-dependent H2B ubiquitination. Mol. Cell 41, 515–528 (2011).
Tang, L.Y. et al. Ablation of Smurf2 reveals an inhibition in TGF-β signalling through multiple mono-ubiquitination of Smad3. EMBO J. 30, 4777–4789 (2011).
Todaro, G.J. & Green, H. Quantitative studies of the growth of mouse embryo cells in culture and their development into established lines. J. Cell Biol. 17, 299–313 (1963).
Harvey, D.M. & Levine, A.J. p53 alteration is a common event in the spontaneous immortalization of primary BALB/c murine embryo fibroblasts. Genes Dev. 5, 2375–2385 (1991).
Sherr, C.J. Tumor surveillance via the ARF-p53 pathway. Genes Dev. 12, 2984–2991 (1998).
Franken, N.A., Rodermond, H.M., Stap, J., Haveman, J. & van Bree, C. Clonogenic assay of cells in vitro. Nat. Protoc. 1, 2315–2319 (2006).
Montecucco, A. & Biamonti, G. Cellular response to etoposide treatment. Cancer Lett. 252, 9–18 (2007).
Bonner, W.M. et al. γH2AX and cancer. Nat. Rev. Cancer 8, 957–967 (2008).
Falk, M., Lukasova, E. & Kozubek, S. Chromatin structure influences the sensitivity of DNA to gamma-radiation. Biochim. Biophys. Acta 1783, 2398–2414 (2008).
Murga, M. et al. Global chromatin compaction limits the strength of the DNA damage response. J. Cell Biol. 178, 1101–1108 (2007).
Shilatifard, A. Chromatin modifications by methylation and ubiquitination: implications in the regulation of gene expression. Annu. Rev. Biochem. 75, 243–269 (2006).
Weake, V.M. & Workman, J.L. Histone ubiquitination: triggering gene activity. Mol. Cell 29, 653–663 (2008).
Sun, Z.W. & Allis, C.D. Ubiquitination of histone H2B regulates H3 methylation and gene silencing in yeast. Nature 418, 104–108 (2002).
Laribee, R.N. et al. BUR kinase selectively regulates H3 K4 trimethylation and H2B ubiquitylation through recruitment of the PAF elongation complex. Curr. Biol. 15, 1487–1493 (2005).
Lee, J.S. et al. Histone crosstalk between H2B monoubiquitination and H3 methylation mediated by COMPASS. Cell 131, 1084–1096 (2007).
Kim, J. et al. RAD6-Mediated transcription-coupled H2B ubiquitylation directly stimulates H3K4 methylation in human cells. Cell 137, 459–471 (2009).
Kizer, K.O. et al. A novel domain in Set2 mediates RNA polymerase II interaction and couples histone H3 K36 methylation with transcript elongation. Mol. Cell. Biol. 25, 3305–3316 (2005).
Berglund, L. et al. A genecentric Human Protein Atlas for expression profiles based on antibodies. Mol. Cell. Proteomics 7, 2019–2027 (2008).
Haber, D. & Harlow, E. Tumour-suppressor genes: evolving definitions in the genomic age. Nat. Genet. 16, 320–322 (1997).
Fukunaga, E. et al. Smurf2 induces ubiquitin-dependent degradation of Smurf1 to prevent migration of breast cancer cells. J. Biol. Chem. 283, 35660–35667 (2008).
Fierz, B. et al. Histone H2B ubiquitylation disrupts local and higher-order chromatin compaction. Nat. Chem. Biol. 7, 113–119 (2011).
Hogan, B., Beddington, R., Constantini, F. & Lacy, E. Manipulating the Mouse Embryo: A Laboratory Manual. 2 edn. (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, USA, 1994).
Blank, M., Lerenthal, Y., Mittelman, L. & Shiloh, Y. Condensin I recruitment and uneven chromatin condensation precede mitotic cell death in response to DNA damage. J. Cell Biol. 174, 195–206 (2006).
Galanty, Y. et al. Mammalian SUMO E3-ligases PIAS1 and PIAS4 promote responses to DNA double-strand breaks. Nature 462, 935–939 (2009).
Rudolph, C., Adam, G. & Simm, A. Determination of copy number of c-Myc protein per cell by quantitative Western blotting. Anal. Biochem. 269, 66–71 (1999).
Shechter, D., Dormann, H.L., Allis, C.D. & Hake, S.B. Extraction, purification and analysis of histones. Nat. Protoc. 2, 1445–1457 (2007).
Ziv, Y. et al. Chromatin relaxation in response to DNA double-strand breaks is modulated by a novel ATM- and KAP-1 dependent pathway. Nat. Cell Biol. 8, 870–876 (2006).
Zaret, K. Micrococcal nuclease analysis of chromatin structure. Curr Protoc Mol. Biol. Chapter 21, Unit 21 (2005).
Yamashita, M. et al. Ubiquitin ligase Smurf1 controls osteoblast activity and bone homeostasis by targeting MEKK2 for degradation. Cell 121, 101–113 (2005).
Acknowledgements
We thank M. Anver for pathology services, V. Barr for assistance with microscope, X. Wu for assistance with the microarray experiments, N. Morris for the animal husbandry and N. Teja for assistance with cell culture. We also thank K. Sixt for comments on the manuscript. This research is supported by the Intramural Research Program of the US National Cancer Institute, US National Institutes of Health, Center for Cancer Research. M.Y. was partially supported by the Japan Society for the Promotion of Science grant 21689053.
Author information
Authors and Affiliations
Contributions
M.Y. and Y.T. maintained mouse colonies and generated primary MEFs and mouse dermal fibroblasts. M.Y., S.Y.C. and Y.E.Z. observed and analyzed the spontaneous tumor formation in mice. S.S.B. performed karyotyping analyses. Y.E.Z. analyzed the microarray data. M.B. performed all other experiments described in the manuscript. M.B. and Y.E.Z. conceived of the study, analyzed the data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9, Supplementary Tables 1–5 and Supplementary Methods (PDF 2152 kb)
Rights and permissions
About this article
Cite this article
Blank, M., Tang, Y., Yamashita, M. et al. A tumor suppressor function of Smurf2 associated with controlling chromatin landscape and genome stability through RNF20. Nat Med 18, 227–234 (2012). https://doi.org/10.1038/nm.2596
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.2596
This article is cited by
-
Smurf1 and Smurf2 mediated polyubiquitination and degradation of RNF220 suppresses Shh-group medulloblastoma
Cell Death & Disease (2023)
-
SMURF2 phosphorylation at Thr249 modifies glioma stemness and tumorigenicity by regulating TGF-β receptor stability
Communications Biology (2022)
-
Smurf2 inhibition enhances chemotherapy and radiation sensitivity in non-small-cell lung cancer
Scientific Reports (2022)
-
The E3 ubiquitin ligase SMURF2 stabilizes RNA editase ADAR1p110 and promotes its adenosine-to-inosine (A-to-I) editing function
Cellular and Molecular Life Sciences (2022)
-
SMURF1, a promoter of tumor cell progression?
Cancer Gene Therapy (2021)