Abstract
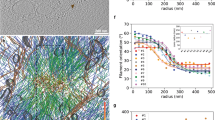
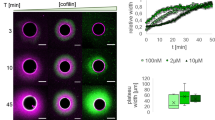
Actin filaments constitute one of the main components of cell cytoskeleton. Assembled into bundles in filopodia or in stress fibres, they play a pivotal role in eukaryotes during cell morphogenesis, adhesion and motility. The bundle emergence has been extensively related to specific actin regulators1,2,3 in vivo4,5,6,7. Such dynamic modulation was also highlighted by biochemical reconstitution of the actin-network assembly, in bulk solution or with biomimetic devices8,9,10,11,12,13,14,15,16,17,18. However, the question of how geometrical boundaries, such as those encountered in cells, affect the dynamic formation of highly ordered actin structures remains poorly studied14,19. Here we demonstrate that the nucleation geometry in itself can be the principal determinant of actin-network architecture. We developed a micropatterning method that enables the spatial control of actin nucleation sites for in vitro assays. Shape, orientation and distance between nucleation regions control filament orientation and length, filament–filament interactions and filopodium-like bundle formation. Modelling of filament growth and interactions demonstrates that basic mechanical and probabilistic laws govern actin assembly in higher-order structures.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Naumanen, P., Lappalainen, P. & Hotulainen, P. Mechanisms of actin stress fibre assembly. J. Microsc. 231, 446–454 (2008).
Welch, M. D. & Mullins, R. D. Cellular control of actin nucleation. Annu. Rev. Cell. Dev. Biol. 18, 247–288 (2002).
Chesarone, M. A., DuPage, A. G. & Goode, B. L. Unleashing formins to remodel the actin and microtubule cytoskeletons. Nature Rev. Mol. Cell. Biol. 11, 62–74 (2010).
Hotulainen, P. et al. Defining mechanisms of actin polymerization and depolymerization during dendritic spine morphogenesis. J. Cell. Biol. 185, 323–339 (2009).
Korobova, F. & Svitkina, T. Molecular architecture of synaptic actin cytoskeleton in hippocampal neurons reveals a mechanism of dendritic spine morphogenesis. Mol. Biol. Cell. 21, 165–176.
Svitkina, T. M. et al. Mechanism of filopodia initiation by reorganization of a dendritic network. J. Cell. Biol. 160, 409–421 (2003).
Vignjevic, D. et al. Role of fascin in filopodial protrusion. J. Cell. Biol. 174, 863–875 (2006).
Loisel, T. P., Boujemaa, R., Pantaloni, D. & Carlier, M. F. Reconstitution of actin-based motility of Listeria and Shigella using pure proteins. Nature 401, 613–616 (1999).
Theriot, J. A., Mitchison, T. J., Tilney, L. G. & Portnoy, D. A. The rate of actin-based motility of intracellular Listeria monocytogenes equals the rate of actin polymerization. Nature 357, 257–260 (1992).
Welch, M. D., Rosenblatt, J., Skoble, J., Portnoy, D. A. & Mitchison, T. J. Interaction of human Arp2/3 complex and the Listeria monocytogenes ActA protein in actin filament nucleation. Science 281, 105–108 (1998).
Bernheim-Groswasser, A., Wiesner, S., Golsteyn, R. M., Carlier, M. F. & Sykes, C. The dynamics of actin-based motility depend on surface parameters. Nature 417, 308–311 (2002).
Cameron, L. A., Footer, M. J., van Oudenaarden, A. & Theriot, J. A. Motility of ActA protein-coated microspheres driven by actin polymerization. Proc. Natl Acad. Sci. USA 96, 4908–4913 (1999).
Romero, S. et al. Formin is a processive motor that requires profilin to accelerate actin assembly and associated ATP hydrolysis. Cell 119, 419–429 (2004).
Liu, A. P. et al. Membrane-induced bundling of actin filaments. Nature Phys. 4, 789–793 (2008).
Achard, V. et al. A primer-based mechanism underlies branched actin filament networks formation and motility. Curr. Biol. 20, 423–428 (2010).
Brieher, W. M., Coughlin, M. & Mitchison, T. J. Fascin-mediated propulsion of Listeria monocytogenes independent of frequent nucleation by the Arp2/3 complex. J. Cell. Biol. 165, 233–242 (2004).
Haviv, L. et al. Reconstitution of the transition from lamellipodium to filopodium in a membrane-free system. Proc. Natl Acad. Sci. USA 103, 4906–4911 (2006).
Vignjevic, D. et al. Formation of filopodia-like bundles in vitro from a dendritic network. J. Cell. Biol. 160, 951–962 (2003).
Mogilner, A. & Rubinstein, B. The physics of filopodial protrusion. Biophys. J. 89, 782–795 (2005).
Azioune, A., Storch, M., Bornens, M., Thery, M. & Piel, M. Simple and rapid process for single cell micro-patterning. Lab Chip 9, 1640–1642 (2009).
Blanchoin, L. et al. Direct observation of dendritic actin filament networks nucleated by Arp2/3 complex and WASP/Scar proteins. Nature 404, 1007–1011 (2000).
Machesky, L. M. et al. Scar, a WASp-related protein, activates nucleation of actin filaments by the Arp2/3 complex. Proc. Natl Acad. Sci. USA 96, 3739–3744 (1999).
Mullins, R. D., Heuser, J. A. & Pollard, T. D. The interaction of Arp2/3 complex with actin: Nucleation, high affinity pointed end capping, and formation of branching networks of filaments. Proc. Natl Acad. Sci. USA 95, 6181–6186 (1998).
Courson, D. S. & Rock, R. S. Actin crosslink assembly and disassembly mechanics for alpha-actinin and fascin. J. Biol. Chem. 285, 26350–26357 (2010).
Medalia, O. et al. Organization of actin networks in intact filopodia. Curr. Biol. 17, 79–84 (2007).
Iwasa, J. H. & Mullins, R. D. Spatial and temporal relationships between actin-filament nucleation, capping, and disassembly. Curr. Biol. 17, 395–406 (2007).
Parker, K. K. et al. Directional control of lamellipodia extension by constraining cell shape and orienting cell tractional forces. Faseb. J. 16, 1195–1204 (2002).
Thery, M., Pepin, A., Dressaire, E., Chen, Y. & Bornens, M. Cell distribution of stress fibres in response to adhesive environment geometry. Cell. Motil. Cytoskeleton 63, 341–355 (2006).
Acknowledgements
We are grateful to C. J. Staiger, J. Plastino and D. Pellman for critical reading of the manuscript and suggestions. This work was supported by grants from Agence National pour la Recherche to L.B. (ANR-06-PCV1-0022 and ANR-08-BLAN-0012) and M.T. (ANR-08-JC-0103 and ANR-PCV08-322457); A-C.R. is supported by an IRTELIS fellowship from CEA.
Author information
Authors and Affiliations
Contributions
A-C.R. and T.C. carried out the experiments. J-L.M. carried out the physical modelling. L.B., R.B-P. and M.T. directed the project and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1385 kb)
Supplementary Information
Supplementary Movie 1 (MOV 5887 kb)
Supplementary Information
Supplementary Movie 2 (MOV 2729 kb)
Supplementary Information
Supplementary Movie 3 (MOV 1006 kb)
Supplementary Information
Supplementary Movie 4 (MOV 5138 kb)
Supplementary Information
Supplementary Movie 5 (MOV 7066 kb)
Rights and permissions
About this article
Cite this article
Reymann, AC., Martiel, JL., Cambier, T. et al. Nucleation geometry governs ordered actin networks structures. Nature Mater 9, 827–832 (2010). https://doi.org/10.1038/nmat2855
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmat2855
This article is cited by
-
Flagella-like beating of actin bundles driven by self-organized myosin waves
Nature Physics (2022)
-
Talin dissociates from RIAM and associates to vinculin sequentially in response to the actomyosin force
Nature Communications (2020)
-
Synthetic asters as elastic and radial skeletons
Nature Communications (2019)
-
Network heterogeneity regulates steering in actin-based motility
Nature Communications (2017)
-
A new method to measure mechanics and dynamic assembly of branched actin networks
Scientific Reports (2017)