Abstract
Compartments for the spatially and temporally controlled assembly of biological processes are essential towards cellular life. Synthetic mimics of cellular compartments based on lipid-based protocells lack the mechanical and chemical stability to allow their manipulation into a complex and fully functional synthetic cell. Here, we present a high-throughput microfluidic method to generate stable, defined sized liposomes termed ‘droplet-stabilized giant unilamellar vesicles (dsGUVs)’. The enhanced stability of dsGUVs enables the sequential loading of these compartments with biomolecules, namely purified transmembrane and cytoskeleton proteins by microfluidic pico-injection technology. This constitutes an experimental demonstration of a successful bottom-up assembly of a compartment with contents that would not self-assemble to full functionality when simply mixed together. Following assembly, the stabilizing oil phase and droplet shells are removed to release functional self-supporting protocells to an aqueous phase, enabling them to interact with physiologically relevant matrices.
Similar content being viewed by others
Main
The formation of lipid membrane-based compartments is one of the distinguishing features of eukaryotic cells. Compartments provide physical and chemical barriers that prevent the uncontrolled diffusion of molecular components to and from the surrounding environment, thereby allowing independent and self-contained metabolic, signalling, or synthesizing activities1,2,3,4. Moreover, biological membranes allow for chemically selective intra- and intercellular material transport and signal transduction by various transmembrane proteins, such as ion channels and receptors5,6. In the context of synthetic biology, protocells are synthetic, biomolecule-containing, lipid-based compartments. These compartments can either be small, large or giant unilamellar vesicles (SUVs, LUVs or GUVs)7,8. The chemical and mechanical instabilities of phospholipids under high ionic strength conditions, especially multivalent cations, and their sensitivity to pH changes are considered to be the main challenges in utilizing protocells for synthetic biology9,10,11. In addition, inserting molecules into protocells represents a particular challenge given their impermeability and mechanical instability12.
Polymersomes made from amphiphilic block copolymers constitute alternative compartments to protocells. They enclose and are surrounded by an aqueous solution13. Some polymersomes can be engineered with transmembrane proteins or synthetic channel molecules that enable certain chemicals to pass the polymer membrane14,15. They are both chemically and mechanically more stable than protocells and are adjustable to certain environments and functionalities13. In contrast to GUVs, where the manipulation of chemical and physical properties bears limitations, the thickness, bending and stretching moduli of the polymeric membrane is tuned by changing the block-copolymer molecular properties. However, the encapsulation of biomolecules and further manipulation of traditional water-in-water polymersomes still represent challenges. The uncontrolled permeability of the polymersomes16,17 and a lack of technologies which allow for the precise and efficient delivery of different biological components are the main drawbacks.
In contrast to polymersomes, copolymer-stabilized water-in-oil droplet compartments can be manipulated at high throughput using automated microfluidic technologies18,19. The biophysical properties of droplet-based compartments, their ultra-fast generation (up to 1 MHz), and the high degree to which this micro-environment is controllable add up to a system that combines the necessary requirements of various synthetic biology applications20,21,22. Moreover, it was shown that the continuous oil phase and its additives play an important role not only in the stable separation of the droplets and in preserving the bio-content23 but also in the combinatorial delivery of biological materials exclusively into the droplets by means of microfluidic pico-injection technology24. However, much like in the case of polymersomes, the ability of the droplets to serve as optimal cell-like compartments is mainly hindered by their inability to imitate the biophysical properties of cellular lipid membranes17,25.
In this study, we present an approach that merges lipid vesicles and copolymer-stabilized droplets to generate mechanically and chemically stable cell-like compartments, which we have called droplet-stabilized GUVs (dsGUVs). These compartments can be sequentially loaded with biomolecules by means of pico-injection technology and offer a well-defined micro-environment for a real step-by-step bottom-up assembly of intracellular modules with desired features. Moreover, the model systems were released from the oil phase and stabilizing droplets after assembly, thus paving the way towards interactions of these synthetic cells with physiologically relevant environments such as extracellular matrices, cells or signalling proteins.
Formation of dsGUVs and their analysis
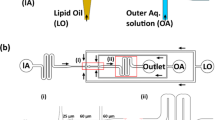
Droplet-based microfluidics allows for high-throughput generation of monodisperse droplets, thus enabling the precise estimation of the lipid concentration required for the formation of a continuous lipid bilayer at the droplets’ inner interface of the polymer shell (Supplementary Note 1 and Supplementary Fig. 1). The determined optimal amount (800 μM) of various lipid compositions in the form of LUVs or GUVs was dissolved in Milli-Q water and encapsulated into copolymer-stabilized droplets of 40 μm diameter (Supplementary Methods). The lipid-containing droplets are shown in Fig. 1a, b. To transform the encapsulated vesicles into a continuous supported lipid bilayer at the droplets’ inner interface, a solution with optimized MgCl2 concentration of 10 mM was introduced during droplet production or by means of pico-injection (Supplementary Methods). In the case of droplets containing LUVs, the formation of the continuous lipid bilayer was detected within a few seconds. The formation of a dsGUV out of the GUVs was considerably slower (in the range of 30 min) due to the slow diffusion of the GUVs towards the droplets’ interface (Supplementary Note 2 and Supplementary Movie 1). Mg2+ ions are considered to be the most efficient mediators of lipid vesicle rupture, because they promote adhesion to a supporting surface10,26. It is worth mentioning that neither the addition of Ca2+ (10 mM) nor lower concentrations of Mg2+ (1 mM) as tested using the same lipid concentration could obtain comprehensive vesicle fusion to the droplets’ inner surface (Supplementary Fig. 2).
a, Lipid vesicles, either LUVs or GUVs, were encapsulated in water-in-oil copolymer-stabilized droplets by means of microfluidics. To transform the encapsulated vesicles into a supported lipid bilayer at the copolymer-stabilized droplets’ inner interface, 10 mM Mg2+ was applied during droplet production or by means of pico-injection. b, Representative combined images of green fluorescence from lipids (ATTO 488-labelled DOPE) and bright-field microscopy of the encapsulated LUVs and GUVs, and of the dsGUVs. The latter two images show superimposed the fluorescence intensity profile. Scale bar, 10 μm. c, The microfluidic device designed to release the assembled lipid compartments from the surrounding stabilizing polymer droplets into the aqueous phase. The left image shows the fluorescence images of the monodisperse dsGUVs in the oil phase prior to release. After injection, droplets are separated from each other at a T-junction where a tributary oil flow containing 20 vol% destabilizing surfactants merges with the droplet flow. Passive trapping structures within the microfluidic channel enable the draining of the continuous oil phase into adjacent oil outlets and decelerate the droplets before they enter the aqueous phase. A comparison of dsGUVs and released lipid compartments shows that they are comparable in size. Scale bars, 20 μm.
To assess the formation process and the dynamics of the assembled dsGUV lipid bilayer, fluorescence intensity analyses, fluorescence recovery after photobleaching (FRAP) measurements, and the release of GUVs from the stabilizing droplet shell were performed. The insets in Fig. 1b show representative fluorescence intensity profiles of the encapsulated GUVs and the dsGUVs (both with identical lipid compositions), evaluated with a laser scanning confocal microscope using the same acquisition conditions (Methods). A fit to the profile revealed similar integrated intensity values of 42 ± 8 and 44 ± 4 (arbitrary intensity units) for dsGUVs and encapsulated GUVs, respectively. These findings may indicate a similar lipid density for dsGUVs and GUVs.
FRAP measurements (Table 1) revealed slightly lower lipid mobility in dsGUVs compared to encapsulated free-standing GUVs (Supplementary Methods and Supplementary Note 3). This can be attributed to the fact that the supported lipid membrane is subject to perturbations from the inner interface of the copolymer-stabilized droplet. Previous reports found a similar or an even lower diffusion coefficient for planar supported lipid membranes in comparison to free-standing GUVs27,28.
Importantly, a microfluidic release device was developed to transfer the assembled lipid compartments into a physiologically relevant, aqueous phase (Fig. 1c, Methods). The release of the assembled lipid compartments (Methods) from the stabilizing polymer droplets into the aqueous phase is shown in Fig. 1c. As can be observed from the fluorescence images, the diameter of the released GUVs (26.2 ± 1.2 μm) in the aqueous phase is slightly smaller compared to that of the dsGUV prior to release (28.3 ± 0.6 μm). A slight shrinkage of the released GUVs can be explained by the change of the osmotic balance. FRAP measurements performed on the released GUVs and on those formed by traditional electroformation (consisting of the same lipid composition) revealed similar lipid mobility (Table 1).
Biofunctionalization of dsGUVs
To test the ability of the dsGUVs to serve as anchoring points for the immobilization of (His-tag) proteins, a lipid solution consisting of 9:1 DOPC:DGS-NTA(Ni) (220 μM) and (His6)-GFP was encapsulated into copolymer-stabilized droplet shells (Methods). The analysis of the fluorescence images of the (His6)-GFP inside the dsGUVs revealed the localization of the fluorescence signal at the periphery of the dsGUV—in other words, the same place at which the anchor DGS-NTA(Ni2+) lipids are situated (Supplementary Fig. 3a). In contrast, a homogeneously distributed fluorescence signal was observed inside non-functionalized dsGUVs (Supplementary Fig. 3b).
FRAP measurements were performed to assess the kinetics of diffusion within the GFP-linked copolymer-stabilized droplets21 and the GFP-linked dsGUVs (Supplementary Fig. 3c). Data analysis revealed diffusion coefficients of 1.22 ± 0.03 and 0.20 ± 0.05 μm2 s−1 for proteins linked to the lipid bilayer or to the polymer membrane, respectively.
Sequential assembly of sub-cellular units into the dsGUVs
To construct cellular modules by means of bottom-up assembly, an automated microfluidic approach was adapted to allow for the sequential loading of various sub-cellular functional units into the dsGUVs. Towards this end, we incorporated transmembrane proteins (integrin and ATP synthase) and cytoskeletal proteins (G-actin and tubulin) into the dsGUVs (Fig. 2).
Following the formation of the dsGUVs, we used a pico-injection system24 to inject and to fuse transmembrane protein-containing proteoliposomes, that is, liposomes containing TAMRA-labelled αIIbβ3 integrin or Alexa 488-labelled FoF1-ATP synthase (Methods), with the dsGUVs composed of 8:1:1 DOPC:DOPE:DOPS. Lipid labelling was obtained by adding either 1% ATTO 488-labelled DOPE or 1% Rhodamine B (RhB)-labelled DOPE. The co-localization of the integrin or the FoF1-ATP synthase with the lipid fluorescence signals is shown in Fig. 3a, indicating the successful fusion of the proteoliposomes with the dsGUVs. As a control, we injected FoF1-ATP synthase into copolymer-stabilized droplets without any dsGUVs, and mixed these droplets with dsGUVs containing FoF1-ATP synthase. It should be noted that only within the dsGUV was the FoF1-ATP synthase fused with the periphery. In the case of droplets without dsGUVs, FoF1-ATP synthases were distributed homogeneously within the entire droplet. Similar results were obtained for integrin. This clearly indicates that transmembrane proteins such as FoF1-ATP synthase or integrin need the formation of a dsGUV to fuse with the lipid bilayer on the droplet periphery.
a, The left and right pair of images show representative fluorescence images of the same areas of observation after pico-injection of integrin-containing liposomes (ATTO 488-labelled DOPE lipids in green and TAMRA-labelled αIIbβ3 integrin in red) and of FoF1-ATP synthase-containing liposomes into the dsGUVs (RhB-labelled DOPE lipids in yellow and Alexa 488-labelled FoF1-ATP synthase in green), respectively. Superimposed on the right pair of images is the fluorescence intensity profile. Lipids and FoF1-ATP synthases clearly co-localize in dsGUVs. Without an inner dsGUV (white dashed circle), the signal from the Alexa 488-labelled FoF1-ATP synthase is equally distributed within the droplet. Scale bars, 20 μm. b, Schematic illustration and representative results of FRAP measurement of labelled integrin with (top row) and without (bottom row) immobilization to the gold-linked surfactants via RGD peptides. The focus area of the light source is encircled in the pre-bleach frame and the timeline on top indicates when follow-up images were taken. Scale bar, 10 μm. c, Schematic illustration of FoF1-ATP synthase-reconstituted dsGUVs and of the transmembrane pH gradient—the driving force of ATP synthesis—as achieved by the addition of acidic FC-40 oil. The right graph shows the bioluminescence intensity response to the ATP content as a function of time. The insert shows the representative bioluminescence intensity curves obtained from the aqueous content of dsGUVs activated (red) by a pH gradient and dsGUVs without a pH gradient (black). The bioluminescence curve was calibrated by the addition of 100 nM ATP solution. d, Representative fluorescence images of microtubules (10% ATTO 488-labelled tubulin, left panel) or the actin cytoskeleton (1% Alexa 488-labelled actin, right panel) in droplets containing RhB-labelled DOPE lipids, as obtained by either the premixed (top) or the pico-injection approach (bottom). Scale bar, 20 μm.
FRAP measurements of transmembrane proteins reconstituted into dsGUVs revealed diffusion coefficients of 1.20 ± 0.7 and 0.70 ± 0.1 μm2 s−1 for FoF1-ATP synthase and integrin, respectively. This value is in good agreement with previously published studies on integrin αIIbβ3 mobility in planar supported lipid bilayers and in cellular membranes29,30,31. Moreover, to test the functionality of the reconstituted integrin, RGD peptides anchored to gold-linked surfactants 21,32 (Methods) were used to provide binding sites for integrin adhesion (Fig. 3b). In this case, the diffusion coefficient of integrin dropped to 0.13 ± 0.03 μm2 s−1, consistent with the mobility of the copolymer surfactant layer that stabilizes the droplet. Successful binding between the integrin and the RGD on the droplet interface indicates the functional incorporation of integrin into the lipid bilayer of the dsGUVs. It also reveals that at least some of the integrin proteins are oriented correctly, with their extracellular parts pointing towards the inner interface of the copolymer-stabilized droplet. It should be mentioned that the establishment of a methodology for integrating membrane proteins into lipid bilayers with a specific orientation is not in the scope of this work, but is an important task for future studies.
For the activity assessment of the reconstituted FoF1-ATP synthase in dsGUVs, an acidic FC-40 oil was added to the oil phase. This addition resulted in the establishment of a transmembrane pH gradient (ΔpH) as a driving force for ATP synthesis inside the droplets (Methods and Fig. 3c). Following 2 min incubation time, the aqueous content of the dsGUVs was released and the synthesized ATP concentration was determined by means of bioluminescence luciferase in a plate reader (Methods). A total amount of 5 nM ATP was measured in the released solution (Fig. 3c and Methods), which indicates the functional reconstitution of the FoF1-ATP synthases in the dsGUVs. It also reveals that at least some of the FoF1-ATP synthases are oriented correctly—that is, the F1 portion is pointing towards the inside of the dsGUV. The low concentration of synthesized ATP can be attributed to the lack of a sufficient transmembrane electric potential gradient33. Moreover, the time delay between generating the proton motive force for ATP synthesis and the determination of the produced ATP was long so that enzyme-catalysed and pH-dependent ATP hydrolysis could reduce the amount of the remaining ATP. The drop of ATP over time as measured in the plate reader is attributed to the kinetic instability of ATP (Fig. 3c).
The same pico-injection approach was applied to obtain a bottom-up reconstitution of the actin cytoskeleton or microtubules within the dsGUVs (Methods). Towards this end, we proceeded by the following two-step process: the formation of dsGUVs (90% DOPC, 9% DOPS and 1% RhB-DOPE) in the presence of an actin or tubulin polymerization buffer; the pico-injection of G-actin (10 μM final concentration, including 1% Alexa 488-labelled G-actin) or tubulin (10 μM final concentration, including 10% ATTO 488-labelled tubulin) solution into the dsGUVs. Moreover, we compared this pico-injection approach to the premixed (one-step) approach, in which G-actin or tubulin proteins were mixed with LUVs prior to droplet formation (Methods).
The successful reconstitution of the actin filaments and microtubules within the dsGUVs could be obtained by sequential pico-injection only (Fig. 3d, middle panel). When using the one-step premixed approach, vesicle fusion to the droplets’ inner surface was suppressed. In the case of microtubules none, and in the case of F-actin only partial fusion, was observed (Fig. 3d, top panel). The inability to form dsGUVs in the presence of microtubules is related to the fact that LUVs are subject to perturbations stemming from the amphiphilic nature of tubulin34. The sequential pico-injection approach enables the prior formation of stable dsGUVs with appropriate ionic conditions followed by the injection of proteins without perturbing the lipid bilayer of the dsGUVs.
Release of GUVs into physiological environments
To pave the way towards investigations of protocell interactions with physiological environments, we developed both a bulk and a microfluidic method to release assembled GUVs with or without proteins from the oil phase and stabilizing droplet shell to the water phase (Methods). By dissolving distinctive fluorophores in the oil phase as well as the enclosed and continuous aqueous phase, we could validate that the lipid membrane remains fully intact and impervious during the release process and that no oil residue could be detected in the water phase or the released GUVs (Supplementary Fig. 6). Moreover, to assess the purity of the GUVs, Raman spectra were collected on released GUVs (Supplementary Methods and Supplementary Fig. 7). As can be observed, no oil/surfactants were detected in the Raman spectra of the released GUVs. Additionally, by means of fluctuation analysis we calculated the bending rigidity values (21.5 ± 3.4kBT) of released GUVs (Supplementary Methods). This value lies in the range of typical values reported for the bending rigidity of PC membranes35, suggesting that the bilayers are clean of impurities. Moreover, these values correlate strongly with the bending rigidity values (25.3 ± 3.0kBT) of the GUVs, consisting of the same lipid composition and formed by a standard electroformation method (Supplementary Methods).
To demonstrate the interactions of the released protocells with physiological environments, Fig. 4 shows optical micrographs of integrin αIIbβ3- or F-actin-containing dsGUVs and their corresponding protocells in the aqueous phase after release. To validate the functionality of the reconstituted integrin αIIbβ3, the spreading behaviour of the released integrin αIIbβ3-functionalized GUVs is investigated. While these integrin αIIbβ3-protocells do not spread on BSA-coated interfaces, they spread well on fibrinogen but less on fibronectin or collagen matrices as is expected from the platelet adhesion receptor integrin αIIbβ3 (Fig. 4b and Supplementary Methods and Supplementary Fig. 8; refs 32,36). The differential adhesion on the various matrices further demonstrates that the protein reconstitution and release process does not affect the biological functionality of the synthetic cells.
a,b, Representative 3D reconstitution of confocal images of dsGUVs with reconstituted integrin (a) and released lipid compartments with reconstituted integrin in contact with BSA- (left image) and fibrinogen-coated (right image) glass surfaces (b). The merged fluorescence signals include both ATTO 488-labelled DOPE lipids and TAMRA-labelled αIIbβ3 integrin. c,d, Representative fluorescence images of the actin cytoskeleton-containing dsGUVs (c) and of the released actin cytoskeleton-containing lipid compartments (d). The fluorescence signals correspond to Alexa 488-labelled actin (red) and RhB-labelled DOPE lipids (green). Scale bar, 20 μm.
In conclusion, the developed technology overcomes fundamental limitations associated with the formation and manipulation of currently existing protocells or polymersomes for the design of complex synthetic cells. Recently developed microfluidic methods showed remarkable success with the high-throughput formation of monodisperse GUVs37,38,39. Moreover, a control over the GUV lipid composition and its asymmetry was achieved by microfluidic methods such as layer-by-layer lipids assembly40, pulse-jet41 and droplet shooting42. Despite the progress made in recent years in designing protocells, the major drawback of all of these methods is a lack of ability to sufficiently manipulate the protocell content sequentially. The assembly of a cell-like compartment with distinct functionality requires the combination of different proteins, molecules and buffer conditions that are incompatible when applied to the same spatial confinement all at once. The formation of dsGUVs enables the generation of mechanically and chemically stable compartments which can be loaded sequentially with different proteins and molecules using pico-injection technology. This sequential procedure allows for a combination of molecules and organized molecular structures that would not occur spontaneously. These system properties lay the groundwork for realizing a real bottom-up assembly of rather complex combinations and functions in synthetic compartments. Importantly, the ability to release functional protocells from the stabilizing droplets into water phase paves the way towards investigations of interactions of synthetic cells with physiologically relevant environments such as given by extracellular matrices, cells, viruses or signalling molecules. Finally, it is worthwhile mentioning that the formation of synthetic cells by this technology is a high-throughput method enabling the generation of up to 103 functional compartments per second. As a result, it generates a vast number of synthetic cells of unique and precise composition, which can be applied for follow-up scientific and technological applications.
Methods
Fluorescence intensity analysis.
To compare the lipid membrane fluorescence intensity of encapsulated GUVs to that of dsGUVs with the same lipid composition (1:1 Egg PC:Egg PG, including 0.5% ATTO488-DOPE), the droplets were evaluated back-to-back, preserving identical settings with a Leica SP5 confocal microscope (Supplementary Methods). More than twenty intensity profiles were extracted for each droplet type using Fiji/ImageJ. Fluorescence analysis of the encapsulated GUVs revealed a reduction in the amplitude and a broadening of the fluorescence intensity profile peak in proximity to the copolymer-stabilized droplet interface (Fig. 1b). This is due to refraction and diffraction at the water–oil phase barrier. To compare the fluorescence intensities, a Gaussian function with a background correction was fitted to the intensity profile peaks using a nonlinear least-squares fit (Matlab 2015 SP1).
Microfluidic release device.
A high-throughput microfluidic device was developed to release assembled lipid compartments from the stabilizing polymer droplet shells into the aqueous phase (Fig. 1c). All flows inside the device were controlled by a microfluidic flow control system (MFCS-EZ, Fluigent). To minimize shear forces, the height of the channels was designed to exceed the droplet diameter, and the pressure in the inlet channels was adjusted to a maximum of 20 mbar with minor corrections for individual devices and experimental conditions. The dsGUVs were injected into the inlet channel of the release device and isolated at the T-junction where a tributary oil flow containing 20 vol% perfluoro-1-octanol destabilizing surfactants (Sigma-Aldrich) joins. The total flow was adjusted to allow efficient time for the destabilizing surfactants to replace and displace stabilizing surfactants prior to reaching the release unit. In this unit, the dsGUVs encounter the aqueous phase in a wide perpendicular channel. To minimize the mechanical impact on the droplets at the oil/water junction, the droplets were decelerated using passive trapping structures within the microfluidic channels (that is, rows of pillars separated by distances smaller than the representative droplet dimensions), which we designed for this purpose (Supplementary Movie 2)43.
To avoid oil penetrating into the aqueous channel whenever there were no droplets in the trapping structures, the aqueous flow was adjusted to achieve a zero-pressure gradient at the oil/water junction. As a result, the oil flows into the adjacent oil outlet channels without droplets blocking the slits. Whenever a droplet enters, it blocks the first slits on both sides, thereby increasing the pressure. As the droplet flows along the passive trapping structures, it passes pairs of slits, opening these up for oil flow to the oil outlet channels. With each pair of slits that opens up, the channel cross section for the oil flow to the adjacent oil channels increases, subsequently decreasing the pressure that is pushing the droplet along the channel. The droplet decelerates as it approaches the oil–water interface. Upon contact with the aqueous phase, the residual surfactant layer peels off the droplet’s polymer shell, which flows to the oil outlet channel. This releases the droplet’s aqueous content (including the lipid compartments) into the aqueous phase with the appropriate ionic conditions.
Lipid composition for release.
The release efficiency of GUVs was found to be dependent on the lipid composition. A few general key factors for the stability of lipid bilayers have been identified. First, for better stability of self-supporting released GUVs 10–20 mol% cholesterol was included (C8667, Sigma-Aldrich) for the release experiments. The lipid composition used for the results presented in Fig. 1c, consisted of a molar ratio of 4:4:2 of DOPC, POPC and cholesterol, respectively and included 0.5% Atto488-DOPE for observation. Second, in the case of negatively charged GUVs, release efficiency was affected when lipid compositions exceeded a net concentration of 10 mol% of negatively charged lipids. Therefore, the optimized lipid composition for the release experiments of negatively charged GUVs consisted of a molar ratio of 1:8:1 of DOPG, DOPC and cholesterol, respectively.
RGD-peptide immobilization to the gold-linked surfactants.
To provide integrin adhesion sites inside the gold-nanostructured droplets, a two-step method to immobilize SN528 RGD peptides32 (kindly provided by Kessler’s group from TU Munich) on gold nanoparticles via its thiol linker was followed21. The freeze-dried Au-PEG(436 g mol−1)-PFPE(7,000 g mol−1) diblock-copolymer surfactant was dissolved in 100 μl fluorinated FC-40 oil. Next, the aqueous RGD-peptide solution (50 μM, 100 μl) was added and everything was stirred for 1 h. To remove the unbound RGD peptides, the crude product solution was centrifuged. The supernatant solution was removed and the precipitant was freeze-dried for 24 h to completely remove the remaining water. Finally, the product was dissolved in 1 ml of FC-40 oil and filtered with a hydrophobic filter (PTFE 0.2 μm) to remove traces of unreacted peptide.
GFP immobilization.
(His6)-GFP was a gift from S. Gardia, Addgene, plasmid #29663. The protein was expressed in Escherichia coli using standard protocols and purified by Ni-NTA chromatography. The linkage of (His6)-GFP to gold-linked surfactants via Ni-NTA-thiol was performed as previously described21, with the exception that purified Milli-Q water (Millipore filtered) was used as the aqueous phase instead of PBS buffer. To link (His6)-GFP to the lipid bilayer, the following procedure was applied. NiCl2 (9 μl, 100 mM, Fluka) was mixed with a water solution of DGS-NTA lipids (300 μl, 1 mM) and stirred for 20 min. LUVs produced from 9:1 DOPC:DGS-NTA(Ni) (220 μM) were encapsulated into copolymer-stabilized droplet shells (100 μm diameter). Following encapsulation, a water solution containing (His6)-GFP (10 μM) and MgCl2 (10 mM) was pico-injected into the droplets to form GFP-linked dsGUVs.
Integrin purification labelling and reconstitution.
Integrin αIIbβ3 was purified from outdated human blood platelets (Katharinenhospital Stuttgart) using TBS and Triton X-100 according to a previously reported protocol32. Affinity chromatography over Concanavalin A and Heparin columns was followed by gel filtration over a Superdex 200 Prep Grade column (GE Healthcare). The biological activity of the purified integrin was analysed by an enzyme-linked immunosorbent assay (ELISA) using AB1967 anti-integrin αIIb antibodies (Merck Millipore). Following purification, integrin was stored at −80 °C in TRIS storage buffer, consisting of 20 mM TRIS/HCl pH 7.4, 150 mM NaCl, 1 mM CaCl2, 1 mM MgCl2, 0.1% (w/v) Triton X-100, 0.02% (w/v) NaN3 and 2 mg ml−1 Aprotinin. Integrin αIIbβ3 was labelled with 5-(and-6)-carboxytetramethylrhodamine, succinimidyl ester, #C1171 TAMRA (Life Technologies) according to a previously published protocol44. A dye/protein ratio of 1.0 to 1.2 was measured by molecular dye extinction (ɛ555 = 80,000 mol−1cm−1). Functionality was measured with an ELISA binding assay.
Integrin reconstitution into the liposomes followed a previously published protocol30. In brief, 50 mol% of Egg PC and Egg PG were dried under a gentle stream of nitrogen and then placed under vacuum conditions overnight. Dried lipids were dissolved in 1 ml of TRIS buffer, consisting of 20 mM TRIS/HCl, pH 7.4, 50 mM NaCl, 0.5 mM CaCl2 and 0.1% (w/v) Triton X-100, and integrin was added to a final 1:1,000 integrin–lipid ratio and incubated in the solution at 37 °C for 2 h. Triton X-100 was removed in two subsequent washing steps of 3.5 h each using 50 mg ml−1 BT Bio-Beads SM-2 (#152-8920, BIO-RAD). The size distribution of the liposomes and integrin-reconstituted proteoliposomes was measured by dynamic light scattering (DLS) in a Malvern Zetasizer (Nano ZS). Pico-injection technology was used to inject the integrin liposomes into the dsGUVs, which contained integrin activation buffer (20 mM TRIS/HCl, pH 7.4, 50 mM NaCl, 0.5 mM CaCl2, 1 mM MnCl2 and 1 mM MgCl2). Theoretical estimation of the number of integrin proteins that might be reconstituted into dsGUVs is presented in Supplementary Note 4.
For the release of integrin-reconstituted GUVs from the stabilizing droplet shells, integrin was reconstituted into liposomes consisting of 100% EggPC. These proteoliposomes were mixed at a ratio of 1:9 with liposomes containing 4:3:2 DOPC:POPC:cholesterol including 1% ATTO 488-labelled DOPE and used for dsGUV formation.
FoF1-ATP synthase purification, labelling, reconstitution and activity assessment.
The purification of FoF1-ATP synthase from E. coli and subsequent cysteine labelling at the C terminus of subunit a with Alexa 488 maleimide were performed according to the protocols described in Zimmermann et al. 45 and Heitkamp and colleagues46. Labelled (or unlabelled) FoF1-ATP synthase was reconstituted into preformed liposomes (∼120 nm diameter) as described in Fischer and Gräber33 and stored at −80 °C in tricine buffer, consisting of 20 mM tricine–NaOH (pH 8.0), 20 mM succinic acid, 0.6 mM KCl, 50 mM NaCl and 2.5 mM MgCl2. Pico-injection technology was used to inject the FoF1-ATP synthase liposomes into the dsGUVs, which contained FoF1-ATP activity buffer, consisting of 20 mM tricine–NaOH (pH 7.5), 20 mM succinic acid, 10 mM MgCl2, 5 mM NaH2PO4 and 50 μM ultra-pure ADP (Cell Technology).
For the activity assessment of the reconstituted FoF1-ATP synthase in dsGUVs the ATP synthase has to be energized by a transmembrane pH gradient established between the FoF1-ATP synthase-containing dsGUVs and the surrounding oil. To generate a pH gradient (ΔpH ≈ 3), 1μl of trifluoroacetic acid (TFA, 99%, Sigma-Aldrich) was dissolved in 1 ml FC-40 oil47 and an oil exchange was performed. Following the application of the acidic oil, the change in the droplets’ internal pH through proton diffusion was analysed by pyranine intensity detection (Supplementary Movie 3 and Supplementary Fig. 10)48.
Following the reconstitution of the FoF1-ATP synthases in dsGUVs, 100 μl oil/dsGUVs solution was transferred to a 500 μl Eppendorf tube and 20 μl of acidic FC-40 oil was added by pipetting. The Eppendorf tube was carefully tilted and slowly rotated for 2 min. Then, 5 μl of perfluoro-1-octanol 20 vol% destabilizing surfactants (Sigma-Aldrich) was added to release the content of the droplets. To analyse the ATP content, 5 μl of the released aqueous solution was transferred to a well on a non-transparent 96-well plate with a flat bottom, containing 180 μl tricine buffer and 20 μl of 10-fold concentrated luciferase reagent (ATP Bioluminescence Kit CLS II, Sigma-Aldrich). A plate reader (Infinite M200, Tecan) was used to detect the bioluminescence intensity corresponding to the synthesized ATP in the aqueous solution. As a control, the same amount of aqueous solution was released from the FoF1-ATP synthase-containing dsGUVs that were not energized by a transmembrane pH gradient and analysed. To assess the amount of synthesized ATP, a bioluminescence calibration curve was produced by the addition of 100 nM ATP solution.
Actin polymerization within dsGUVs.
Actin was purchased from Cytoskeleton. Actin was stored in TRIS storage buffer, consisting of 2.0 mM TRIS/HCl pH 8, 0.2 mM CaCl2, 0.2 mM ATP, 0.005% NaN3 and 0.2 mM DTT, at −80 °C. Human recombinant fascin was purchased from Novubio (Bio-Techne), aliquoted and stored at −80 °C. 1% of Alexa 488-labelled actin (Life Technologies) was added to the non-labelled actin in all experiments to detect actin filament formation. To polymerize actin inside the dsGUVs, two methods were tested:
Premixed approach: Lipid and G-actin solutions were introduced via a microfluidic aqueous two-phase system (Supplementary Fig. 4C). To avoid polymerization prior to droplet formation, one aqueous phase contained 10 μM actin (1% Alexa 488-labelled actin) in storage buffer and the other aqueous phase contained the polymerization buffer (2.0 mM TRIS/HCl pH 8, 20 mM MgCl2 0.2 mM CaCl2, 0.5 mM ATP, 0.005% NaN3 and 0.2 mM DTT), 0.5 μM fascin and LUV solution consisting of 90% DOPC, 9% DOPS and 1% RhB-DOPE. The two aqueous phases were mixed during droplet generation at a flow-focusing junction (Supplementary Fig. 4).
Pico-injection approach: Two experimental steps were required for stable F-actin creation in dsGUVs. The first step entailed the creation of dsGUVs (90% DOPC, 9% DOPS and 1% RhB-DOPE) that contain actin polymerization buffer. The second step was the pico-injection of storage buffer containing G-actin (10 μM final concentration) and fascin (0.5 μM final concentration) into these droplets.
For the release of F-actin-containing GUVs a lipid composition of DOPC:cholesterol:DOPG (76 mol% DOPC, 20 mol% cholesterol, 3 mol% DOPG including 1 mol% RhB-labelled DOPE) was used when creating the dsGUVs.
Microtubule formation within dsGUVs.
Tubulin (kindly provided by Surrey’s group, the Francis Crick Institute) was purified from pig brain according to previously described protocols49. It was then labelled with ATTO 488-SE (Life Technologies) as described earlier50. Tubulin concentrations were measured using UV spectroscopy (ɛ280 = 115,000 M−1cm−1). The labelling ratio of ATTO 488-labelled tubulin was 0.65 dye molecules per tubulin dimer. Labelled and unlabelled tubulin were stored at −80 °C in PIPES storage buffer (20 mM PIPES pH 6.8, 7.25 mM MgCl2, 1 mM EGTA, 1 mM 2-mercaptoethanol, 50 mM KCl, 31 mM glucose, 1 mg ml−1 glucose oxidase and 0.5 mg ml−1 catalase and 0.25 mg ml−1 beta-casein). To form microtubule networks inside the dsGUVs, two methods were tested:
Premixed approach: Tubulin 14.5 μM (10% labelled with ATTO 488-SE) dissolved in polymerization buffer (20 mM PIPES pH 6.8, 7.25 mM MgCl2, 1 mM EGTA, 3 mM GTP, 1 mM 2-mercaptoethanol, 50 mM KCl, 31 mM glucose, 1 mg ml−1 glucose oxidase and 0.5 mg ml−1 catalase, 0.25 mg ml−1 beta-casein) was mixed with the LUV solution (90% DOPC, 9% DOPS and 1% RhB-DOPE) and introduced into the microfluidic device via an aqueous channel. To avoid tubulin polymerization prior to droplet formation, droplets were produced at 4 °C. After droplet production, tubulin polymerization was triggered by transferring the droplets to a 37 °C observation chamber.
Pico-injection approach: Two experimental steps were required for producing stable microtubules in dsGUVs. The first step entailed the creation of dsGUVs (90% DOPC, 9% DOPS and 1% RhB-DOPE) that contain tubulin polymerization buffer. The second step was the pico-injection of the storage buffer containing tubulin (14.5 μM) into these droplets to get a 10 μM final concentration.
Bulk release approach.
For the successful release of integrin αIIbβ3- or F-actin-containing GUVs (Fig. 4a, b, respectively), the lipid compositions of the dsGUVs were optimized in each case. More details are given in the corresponding methods.
Following the formation of protein-containing dsGUVs, 100 μl oil/dsGUV-containing solution was transferred into a 2 ml Eppendorf tube containing 1 ml FC-40 oil/surfactant solution (identical to the FC-40 oil/surfactant solution used for dsGUV production). Next, 100 μl of the appropriate buffer (actin polymerization buffer or integrin activation buffer) was pipetted on to the droplet emulsion. To destabilize the polymer shell of the droplets, 100 μl of 20 vol% perfluoro-1-octanol destabilizing surfactants (Sigma-Aldrich) dissolved in FC-40 oil was added. The Eppendorf tube was carefully tilted and slowly rotated until the emulsion was broken. The released GUVs were studied in an observation chamber made of BSA-coated glass slides and cover slips. The observation chambers were prepared by incubating the glass with 10 mg ml−1 BSA in PBS for 2 h at room temperature, followed by two 5 min washing steps—one with PBS and one with water.
Code availability.
Matlab 2015 SP1 codes used for FRAP and intensity analyses are available in the Supplementary Notes 5 and 6, respectively.
Data availability.
Data supporting the findings of this study are available within the article and its Supplementary Information files and from the corresponding authors on reasonable request.
Additional Information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Szostak, J. W., Bartel, D. P. & Luisi, P. L. Synthesizing life. Nature 409, 387–390 (2001).
Tawfik, D. S. & Griffiths, A. D. Man-made cell-like compartments for molecular evolution. Nat. Biotechnol. 16, 652–656 (1998).
Diekmann, Y. & Pereira-Leal, J. B. Evolution of intracellular compartmentalization. Biochem. J. 449, 319–331 (2013).
Agapakis, C. M., Boyle, P. M. & Silver, P. A. Natural strategies for the spatial optimization of metabolism in synthetic biology. Nat. Chem. Biol. 8, 527–535 (2012).
Yoshida, M., Muneyuki, E. & Hisabori, T. ATP synthase—A marvellous rotary engine of the cell. Nat. Rev. Mol. Cell Biol. 2, 669–677 (2001).
Geiger, B., Spatz, J. P. & Bershadsky, A. D. Environmental sensing through focal adhesions. Nat. Rev. Mol. Cell Biol. 10, 21–33 (2009).
Nomura, S. M. et al. Gene expression within cell-sized lipid vesicles. Chembiochem 4, 1172–1175 (2003).
Merkle, D., Kahya, N. & Schwille, P. Reconstitution and anchoring of cytoskeleton inside giant unilamellar vesicles. Chembiochem 9, 2673–2681 (2008).
Hardy, G. J., Nayak, R. & Zauscher, S. Model cell membranes: techniques to form complex biomimetic supported lipid bilayers via vesicle fusion. Curr. Opin. Colloid Interface Sci. 18, 448–458 (2013).
Seantier, B. & Kasemo, B. Influence of mono- and divalent ions on the formation of supported phospholipid bilayers via vesicle adsorption. Langmuir 25, 5767–5772 (2009).
Shigematsu, T., Koshiyama, K. & Wada, S. Effects of stretching speed on mechanical rupture of phospholipid/cholesterol bilayers: molecular dynamics simulation. Sci. Rep. 5, 15369 (2015).
Jorgensen, I. L., Kemmer, G. C. & Pomorski, T. G. Membrane protein reconstitution into giant unilamellar vesicles: a review on current techniques. Eur. Biophys. J. 46, 103–119 (2016).
Discher, B. M. et al. Polymersomes: tough vesicles made from diblock copolymers. Science 284, 1143–1146 (1999).
Palivan, C. G. et al. Bioinspired polymer vesicles and membranes for biological and medical applications. Chem. Soc. Rev. 45, 377–411 (2016).
Onaca, O., Nallani, M., Ihle, S., Schenk, A. & Schwaneberg, U. Functionalized nanocompartments (Synthosomes): limitations and prospective applications in industrial biotechnology. Biotechnol. J. 1, 795–805 (2006).
Picker, A., Nuss, H., Guenoun, P. & Chevallard, C. Polymer vesicles as microreactors for bioinspired calcium carbonate precipitation. Langmuir 27, 3213–3218 (2011).
Lee, J. C. M., Santore, M., Bates, F. S. & Discher, D. E. From membranes to melts, rouse to reptation: diffusion in polymersome versus lipid bilayers. Macromolecules 35, 323–326 (2002).
Duncombe, T. A., Tentori, A. M. & Herr, A. E. Microfluidics: reframing biological enquiry. Nat. Rev. Mol. Cell Biol. 16, 554–567 (2015).
Martino, C. & deMello, A. J. Droplet-based microfluidics for artificial cell generation: a brief review. Interface Focus 6, 20160011 (2016).
Schaerli, Y. et al. Continuous-flow polymerase chain reaction of single-copy DNA in microfluidic microdroplets. Anal. Chem. 81, 302–306 (2009).
Platzman, I., Janiesch, J.-W. & Spatz, J. P. Synthesis of nanostructured and biofunctionalized water-in-oil droplets as tools for homing T cells. J. Am. Chem. Soc. 135, 3339–3342 (2013).
Huebner, A. et al. Quantitative detection of protein expression in single cells using droplet microfluidics. Chem. Commun. 12, 1218–1220 (2007).
Janiesch, J. W. et al. Key factors for stable retention of fluorophores and labeled biomolecules in droplet-based microfluidics. Anal. Chem. 87, 2063–2067 (2015).
Abate, A. R., Hung, T., Mary, P., Agresti, J. J. & Weitz, D. A. High-throughput injection with microfluidics using picoinjectors. Proc. Natl Acad. Sci. USA 107, 19163–19166 (2010).
Itel, F. et al. Molecular organization and dynamics in polymersome membranes: a lateral diffusion study. Macromolecules 47, 7588–7596 (2014).
Bhatia, T., Husen, P., Ipsen, J. H., Bagatolli, L. A. & Simonsen, A. C. Fluid domain patterns in free-standing membranes captured on a solid support. Biochim. Biophys. Acta Biomembr. 1838, 2503–2510 (2014).
Machan, R. & Hof, M. Lipid diffusion in planar membranes investigated by fluorescence correlation spectroscopy. Biochim. Biophys. Acta 1798, 1377–1391 (2010).
Przybylo, M. et al. Lipid diffusion in giant unilamellar vesicles is more than 2 times faster than in supported phospholipid bilayers under identical conditions. Langmuir 22, 9096–9099 (2006).
Goennenwein, S., Tanaka, M., Hu, B., Moroder, L. & Sackmann, E. Functional incorporation of integrins into solid supported membranes on ultrathin films of cellulose: impact on adhesion. Biophys. J. 85, 646–655 (2003).
Erb, E. M., Tangemann, K., Bohrmann, B., Muller, B. & Engel, J. Integrin alpha IIb beta 3 reconstituted into lipid bilayers is nonclustered in its activated state but clusters after fibrinogen binding. Biochemistry 36, 7395–7402 (1997).
Edel, J. B., Wu, M., Baird, B. & Craighead, H. G. High spatial resolution observation of single-molecule dynamics in living cell membranes. Biophys. J. 88, L43–L45 (2005).
Frohnmayer, J. P. et al. Minimal synthetic cells to study integrin-mediated adhesion. Angew. Chem. Int. Ed. 54, 12472–12478 (2015).
Fischer, S. & Graber, P. Comparison of Delta pH- and Delta phi-driven ATP synthesis catalyzed by the H + -ATPases from Escherichia coli or chloroplasts reconstituted into liposomes. FEBS Lett. 457, 327–332 (1999).
Wolff, J. Plasma membrane tubulin. Biochim. Biophys. Acta Biomembr. 1788, 1415–1433 (2009).
Dimova, R. Recent developments in the field of bending rigidity measurements on membranes. Adv. Colloid Interface Sci. 208, 225–234 (2014).
Streicher, P. et al. Integrin reconstituted in GUVs: a biomimetic system to study initial steps of cell spreading. Biochim. Biophys. Acta Biomembr. 1788, 2291–2300 (2009).
Karamdad, K., Law, R. V., Seddon, J. M., Brooks, N. J. & Ces, O. Preparation and mechanical characterisation of giant unilamellar vesicles by a microfluidic method. Lab Chip 15, 557–562 (2015).
Matosevic, S. & Paegel, B. M. Stepwise synthesis of giant unilamellar vesicles on a microfluidic assembly line. J. Am. Chem. Soc. 133, 2798–2800 (2011).
Deng, N.-N., Yelleswarapu, M. & Huck, W. T. S. Monodisperse uni- and multicompartment liposomes. J. Am. Chem. Soc. 138, 7584–7591 (2016).
Matosevic, S. & Paegel, B. M. Layer-by-layer cell membrane assembly. Nat. Chem. 5, 958–963 (2013).
Kamiya, K., Kawano, R., Osaki, T., Akiyoshi, K. & Takeuchi, S. Cell-sized asymmetric lipid vesicles facilitate the investigation of asymmetric membranes. Nat. Chem. 8, 881–889 (2016).
Morita, M. et al. Droplet-shooting and size-filtration (DSSF) method for synthesis of cell-sized liposomes with controlled lipid compositions. ChemBioChem 16, 2029–2035 (2015).
Niu, X., Gulati, S., Edel, J. B. & deMello, A. J. Pillar-induced droplet merging in microfluidic circuits. Lab Chip 8, 1837–1841 (2008).
Eberhard, C. Online-Ressource (Heidelberg Univ., 2012).
Zimmermann, B., Diez, M., Zarrabi, N., Graber, P. & Borsch, M. Movements of the epsilon-subunit during catalysis and activation in single membrane-bound H + -ATP synthase. EMBO J. 24, 2053–2063 (2005).
Heitkamp, T., Deckers-Hebestreit, G. & Borsch, M. in Single Molecule Spectroscopy and Superresolution Imaging IX Vol. 9714 (eds Enderlein, J., Gregor, I., Gryczynski, Z. K., Erdmann, R. & Koberling, F.) (Spie-Int Soc Optical Engineering, 2016).
Mashaghi, S. & van Oijen, A. M. External control of reactions in microdroplets. Sci. Rep. 5, 11837 (2015).
Gan, B. S., Krump, E., Shrode, L. D. & Grinstein, S. Loading pyranine via purinergic receptors or hypotonic stress for measurement of cytosolic pH by imaging. Am. J. Physiol. 275, C1158–C1166 (1998).
Castoldi, M. & Popov, A. V. Purification of brain tubulin through two cycles of polymerization-depolymerization in a high-molarity buffer. Protein Expr. Purif. 32, 83–88 (2003).
Hyman, A. et al. Preparation of modified tubulins. Methods Enzymol. 196, 478–485 (1991).
Acknowledgements
Parts of the research leading to these results have received funding from the European Research Council/ERC Grant Agreement no. 294852, SynAd. This work is also part of the MaxSynBio consortium, which is jointly funded by the Federal Ministry of Education and Research of Germany and the Max Planck Society. The work was also partly supported by the SFB 1129 of the German Science Foundation and the VolkswagenStiftung (priority call ‘Life?’). J.P.S. is the Weston Visiting Professor at the Weizmann Institute of Science and part of the excellence cluster CellNetworks at the University of Heidelberg. J.-C.B. acknowledges financial support by the ERC (FP7/2007-2013/ERC Grant agreement 306385-SofI). I.P. acknowledges the support of the Alexander von Humboldt Foundation. The authors acknowledge the help of P. Gruner and B. Riechers for their technical assistance with preliminary microfluidic experiments and A. Richter (WITec GmbH, Germany) for her technical assistance with Raman microscopy. The support of N. Grunze for editing the manuscript as well as of J. Ricken and Ch. Mollenhauer for their general support in protein purification and chemical synthesis is highly acknowledged. The Max Planck Society is appreciated for its general support in all aspects of our research.
Author information
Authors and Affiliations
Contributions
M.W. and J.P.F. realized experimentally the general concept of dsGUVs by microfluidic technology, established pico-injection technology and its application for the design of synthetic cells—this includes the formation of dsGUVs, release of GUVs from oil to water phase, FRAP measurements—and wrote parts of the manuscript. M.W. performed microtubule assembly and integration of functional ATP synthase and its analysis; J.P.F. reconstituted functional Integrin in liposomes and dsGUV and performed release of Integrin GUVs from oil to water phase; L.T.B. optimized the release of GUVs from oil to water phase, in particular the release of integrin functionalized GUVs, performed adhesion experiments of integrin functionalized GUVs to different matrices, and wrote parts of the manuscript; B.H. performed the experiments of dsGUVs with F-actin and its release from oil to water phase and performed Raman spectroscopy analysis; J.-W.J. performed the experiments of dsGUVs with F-actin and synthesized polymer-based surfactants; T.H. and M.B. prepared the labelled FoF1-ATP synthase, R.B.L., R.D. and R.L. developed and discussed lipid bilayer formation using droplets, E.B. and J.-C.B. helped install the pico-injection technology, T.V.-K. and K.S. supported the reconstitution of FoF1-ATP synthase, I.P. designed and supervised the experiments, and wrote the manuscript; J.P.S. invented the concept of synthetic cell formation by sequential bottom-up assembly in droplet-stabilized compartments, designed, supervised and managed the experiments, and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1353 kb)
Supplementary Information
Supplementary movie 1 (MOV 19932 kb)
Supplementary Information
Supplementary movie 2 (MOV 3953 kb)
Supplementary Information
Supplementary movie 3 (MOV 5871 kb)
Rights and permissions
About this article
Cite this article
Weiss, M., Frohnmayer, J., Benk, L. et al. Sequential bottom-up assembly of mechanically stabilized synthetic cells by microfluidics. Nature Mater 17, 89–96 (2018). https://doi.org/10.1038/nmat5005
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmat5005
This article is cited by
-
Mesoporous carbon spheres with programmable interiors as efficient nanoreactors for H2O2 electrosynthesis
Nature Communications (2024)
-
Artificial cells for in vivo biomedical applications through red blood cell biomimicry
Nature Communications (2024)
-
Hydrophobic mismatch drives self-organization of designer proteins into synthetic membranes
Nature Communications (2024)
-
Lipid droplets: a cellular organelle vital in cancer cells
Cell Death Discovery (2023)
-
Miniaturizing chemistry and biology using droplets in open systems
Nature Reviews Chemistry (2023)