Abstract
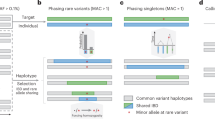
We report a targeted, cost-effective method to quantify rare single-nucleotide polymorphisms from pooled human genomic DNA using second-generation sequencing. We pooled DNA from 1,111 individuals and targeted four genes to identify rare germline variants. Our base-calling algorithm, SNPSeeker, derived from large deviation theory, detected single-nucleotide polymorphisms present at frequencies below the raw error rate of the sequencing platform.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Cohen, J.C. et al. Science 305, 869–872 (2004).
Ji, W. et al. Nat. Genet. 40, 592–599 (2008).
Hayden, E.C. Nature 451, 378–379 (2008).
Hamvas, A. et al. Pediatr. Res. 50, 666–668 (2001).
Barbazuk, W.B., Emrich, S.J., Chen, H.D., Li, L. & Schnable, P.S. Plant J. 51, 910–918 (2007).
Van Tassell, C.P. et al. Nat. Methods 5, 247–252 (2008).
Albert, T.J. et al. Nat. Methods 4, 903–905 (2007).
Hodges, E. et al. Nat. Genet. 39, 1522–1527 (2007).
Porreca, G.J. et al. Nat. Methods 4, 931–936 (2007).
Acknowledgements
This work was supported in part by the US National Institutes of Health under the Ruth L. Kirschstein National Research Service Award T32 HD 007499 from the National Institute of Child Health and Human Development (T.E.D.), the National Heart, Lung and Blood Institute (RO1HL065174, RO1HL082747, F.S.C.), the Children's Discovery Institute Fellowship Award MC-F-2006-1 (T.E.D.), the Children's Discovery Institute grant MC-II-2006-1 (R.D.M.) and the Saigh Foundation (F.S.C., R.D.M. and T.E.D).
Author information
Authors and Affiliations
Contributions
T.E.D., R.D.M. and K.E.V. designed the experiments. T.E.D. performed the sequencing experiments. D.J.W. and S.W.R. performed the TaqMan assays. O.L.K. and J.A.B. punched the bloodspots from filter paper. F.L.M.V designed and wrote SNPSeeker. F.L.M.V. and R.D.M. analyzed data. J.C.F. and S.W.D. designed and executed the comparative genomic analysis. D.J.W., A.H. and F.S.C. provided reagents and advice. T.E.D., R.D.M., F.L.M.V. and K.E.V. wrote the manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3, Supplementary Tables 1–5, Supplementary Methods, Supplementary Results (PDF 398 kb)
Supplementary Software
SNPSeeker README file and binaries for Linux 32-bit, 64-bit, and Mac OSX (ZIP 69 kb)
Rights and permissions
About this article
Cite this article
Druley, T., Vallania, F., Wegner, D. et al. Quantification of rare allelic variants from pooled genomic DNA. Nat Methods 6, 263–265 (2009). https://doi.org/10.1038/nmeth.1307
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1307
This article is cited by
-
A new primer construction technique that effectively increases amplification of rare mutant templates in samples
BMC Biotechnology (2019)
-
Analysis of genome-wide SNPs based on 2b-RAD sequencing of pooled samples reveals signature of selection in different populations of Haemonchus contortus
Journal of Biosciences (2019)
-
Solid-phase enzyme catalysis of DNA end repair and 3′ A-tailing reduces GC-bias in next-generation sequencing of human genomic DNA
Scientific Reports (2018)
-
Targeted sequencing of both DNA strands barcoded and captured individually by RNA probes to identify genome-wide ultra-rare mutations
Scientific Reports (2017)
-
Population-based frequency of surfactant dysfunction mutations in a native Chinese cohort
World Journal of Pediatrics (2016)