Abstract
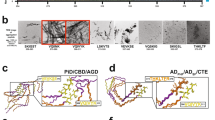
Protein aggregation results in β-sheet–like assemblies that adopt either a variety of amorphous morphologies or ordered amyloid-like structures. These differences in structure also reflect biological differences; amyloid and amorphous β-sheet aggregates have different chaperone affinities, accumulate in different cellular locations and are degraded by different mechanisms. Further, amyloid function depends entirely on a high intrinsic degree of order. Here we experimentally explored the sequence space of amyloid hexapeptides and used the derived data to build Waltz, a web-based tool that uses a position-specific scoring matrix to determine amyloid-forming sequences. Waltz allows users to identify and better distinguish between amyloid sequences and amorphous β-sheet aggregates and allowed us to identify amyloid-forming regions in functional amyloids.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Change history
29 September 2010
In the original version of this Article published online, we had inadvertently neglected to provide a proper comparison to a tool that should have been shown in Table 1. We apologize for this oversight.
29 September 2010
In the version of this article initially published, the name of and reference to the algorithm in the right column of Table 1 was incorrect. The correct reference (ref. 40) has been added in the paper. The error has been corrected in the PDF and HTML versions of the article.
References
Chiti, F. et al. Kinetic partitioning of protein folding and aggregation. Nat. Struct. Biol. 9, 137–143 (2002).
Ventura, S. et al. Short amino acid stretches can mediate amyloid formation in globular proteins: the Src homology 3 (SH3) case. Proc. Natl. Acad. Sci. USA 101, 7258–7263 (2004).
Carrio, M., Gonzalez-Montalban, N., Vera, A., Villaverde, A. & Ventura, S. Amyloid-like properties of bacterial inclusion bodies. J. Mol. Biol. 347, 1025–1037 (2005).
Marshall, K.E. & Serpell, L.C. Structural integrity of beta-sheet assembly. Biochem. Soc. Trans. 37, 671–676 (2009).
Rousseau, F., Schymkowitz, J. & Serrano, L. Protein aggregation and amyloidosis: confusion of the kinds? Curr. Opin. Struct. Biol. 16, 118–126 (2006).
Chiti, F. & Dobson, C.M. Protein misfolding, functional amyloid, and human disease. Annu. Rev. Biochem. 75, 333–366 (2006).
Matsumoto, G., Kim, S. & Morimoto, R.I. Huntingtin and mutant SOD1 form aggregate structures with distinct molecular properties in human cells. J. Biol. Chem. 281, 4477–4485 (2006).
Kopito, R.R. Aggresomes, inclusion bodies and protein aggregation. Trends Cell Biol. 10, 524–530 (2000).
Huyer, G. et al. A striking quality control subcompartment in Saccharomyces cerevisiae: the endoplasmic reticulum-associated compartment. Mol. Biol. Cell 15, 908–921 (2004).
Kaganovich, D., Kopito, R. & Frydman, J. Misfolded proteins partition between two distinct quality control compartments. Nature 454, 1088–1095 (2008).
Arrasate, M., Mitra, S., Schweitzer, E.S., Segal, M.R. & Finkbeiner, S. Inclusion body formation reduces levels of mutant huntingtin and the risk of neuronal death. Nature 431, 805–810 (2004).
McClellan, A.J., Tam, S., Kaganovich, D. & Frydman, J. Protein quality control: chaperones culling corrupt conformations. Nat. Cell Biol. 7, 736–741 (2005).
Fowler, D.M., Koulov, A.V., Balch, W.E. & Kelly, J.W. Functional amyloid–from bacteria to humans. Trends Biochem. Sci. 32, 217–224 (2007).
Wang, X. & Chapman, M.R. Sequence determinants of bacterial amyloid formation. J. Mol. Biol. 380, 570–580 (2008).
Lopez de la Paz, M. & Serrano, L. Sequence determinants of amyloid fibril formation. Proc. Natl. Acad. Sci. USA 101, 87–92 (2004).
Makin, O.S., Atkins, E., Sikorski, P., Johansson, J. & Serpell, L.C. Molecular basis for amyloid fibril formation and stability. Proc. Natl. Acad. Sci. USA 102, 315–320 (2005).
Nelson, R. et al. Structure of the cross-beta spine of amyloid-like fibrils. Nature 435, 773–778 (2005).
Chiti, F., Stefani, M., Taddei, N., Ramponi, G. & Dobson, C.M. Rationalization of the effects of mutations on peptide and protein aggregation rates. Nature 424, 805–808 (2003).
Fernandez-Escamilla, A.M., Rousseau, F., Schymkowitz, J. & Serrano, L. Prediction of sequence-dependent and mutational effects on the aggregation of peptides and proteins. Nat. Biotechnol. 22, 1302–1306 (2004).
Pawar, A.P. et al. Prediction of “aggregation-prone” and “aggregation-susceptible” regions in proteins associated with neurodegenerative diseases. J. Mol. Biol. 350, 379–392 (2005).
Sanchez de Groot, N., Pallares, I., Aviles, F.X., Vendrell, J. & Ventura, S. Prediction of “hot spots” of aggregation in disease-linked polypeptides. BMC Struct. Biol. 5, 18 (2005).
Tartaglia, G.G., Cavalli, A., Pellarin, R. & Caflisch, A. Prediction of aggregation rate and aggregation-prone segments in polypeptide sequences. Protein Sci. 14, 2723–2734 (2005).
Galzitskaya, O.V., Garbuzynskiy, S.O. & Lobanov, M.Y. Prediction of amyloidogenic and disordered regions in protein chains. PLOS Comput. Biol. 2, e177 (2006).
Saiki, M., Konakahara, T. & Morii, H. Interaction-based evaluation of the propensity for amyloid formation with cross-beta structure. Biochem. Biophys. Res. Commun. 343, 1262–1271 (2006).
Thompson, M.J. et al. The 3D profile method for identifying fibril-forming segments of proteins. Proc. Natl. Acad. Sci. USA 103, 4074–4078 (2006).
Hamodrakas, S.J., Liappa, C. & Iconomidou, V.A. Consensus prediction of amyloidogenic determinants in amyloid fibril-forming proteins. Int. J. Biol. Macromol. 41, 295–300 (2007).
Zibaee, S., Makin, O.S., Goedert, M. & Serpell, L.C. A simple algorithm locates beta-strands in the amyloid fibril core of alpha-synuclein, Abeta, and tau using the amino acid sequence alone. Protein Sci. 16, 906–918 (2007).
Sawaya, M.R. et al. Atomic structures of amyloid cross-beta spines reveal varied steric zippers. Nature 447, 453–457 (2007).
Osherovich, L.Z., Cox, B.S., Tuite, M.F. & Weissman, J.S. Dissection and design of yeast prions. PLoS Biol. 2, E86 (2004).
Tartaglia, G.G. et al. Prediction of aggregation-prone regions in structured proteins. J. Mol. Biol. 380, 425–436 (2008).
Hulo, N. et al. The PROSITE database. Nucleic Acids Res. 34, D227–D230 (2006).
Makin, O.S. & Serpell, L. X-ray diffraction studies of amyloid structure. In Amyloid Proteins: Methods and Protocols (ed. Sigurdsson, E.M.) vol. 299, 67–80 (Humana Press, 2005).
Makin, O.S., Sikorski, P. & Serpell, L. CLEARER: a new tool for the analysis of X-ray fibre diffraction patterns and diffraction simulation from atomic structural models. Appl. Cryst. 40, 966–972 (2007).
Schymkowitz, J. et al. The FoldX web server: an online force field. Nucleic Acids Res. 33, W382–388 (2005).
Maurer-Stroh, S. & Eisenhaber, F. Refinement and prediction of protein prenylation motifs. Genome Biol. 6, R55 (2005).
Mirny, L. & Shakhnovich, E. Evolutionary conservation of the folding nucleus. J. Mol. Biol. 308, 123–129 (2001).
Eisenhaber, B., Bork, P. & Eisenhaber, F. Sequence properties of GPI-anchored proteins near the omega-site: constraints for the polypeptide binding site of the putative transamidase. Protein Eng. 11, 1155–1161 (1998).
Tomii, K. & Kanehisa, M. Analysis of amino acid indices and mutation matrices for sequence comparison and structure prediction of proteins. Protein Eng. 9, 27–36 (1996).
Eisenhaber, B., Eisenhaber, F., Maurer-Stroh, S. & Neuberger, G. Prediction of sequence signals for lipid post-translational modifications: insights from case studies. Proteomics 4, 1614–1625 (2004).
Zhang, Z.Q., Chen, H. & Lai, L.H. Identification of amyloid fibril-forming segments based on structure and residue-based statistical potential. Bioinformatics 23, 2218–2225 (2007).
Acknowledgements
S.M.-S. was supported by a Marie Curie Intraeuropean fellowship. I.C.M. was supported by a doctorate scholarship from the Boerhinger Ingelheim Fonds, Foundation for Basic Research in Biomedicine and then the Fundação para a Ciência e Tecnologia. L. Serpell acknowledges the grant support from the Alzheimer's Research Trust and the Biotechnology and Biological Sciences Research Council. L. Serrano was partly supported by the EC grant Apopis. J.W.H.S. and F.R. acknowledge grant support from the Federal Office for Scientific Affairs, Belgium (Interuniversity Attraction Pole P6/43), the Fund for Scientific Research, Flanders, the Institute for Innovation by Science & Technology Flanders, the Stichting Alzheimer Onderzoek and the Alzheimer's Research Trust.
Author information
Authors and Affiliations
Contributions
S.M.-S., F.R., L. Serrano and J.S. devised the methods. S.M.-S. implemented software. J.R., S.M.-S., J.W.H.S. and F.R. performed analysis. M.D., N.K., M.L.d.I.P., I.C.M., K.L.M., A.C. and L. Serpell performed experiments. S.M.-S., J.S. and F.R. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3, Supplementary Tables 1–3 and Supplementary Notes 1–4 (PDF 1206 kb)
Rights and permissions
About this article
Cite this article
Maurer-Stroh, S., Debulpaep, M., Kuemmerer, N. et al. Exploring the sequence determinants of amyloid structure using position-specific scoring matrices. Nat Methods 7, 237–242 (2010). https://doi.org/10.1038/nmeth.1432
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1432
This article is cited by
-
Widespread amyloidogenicity potential of multiple myeloma patient-derived immunoglobulin light chains
BMC Biology (2023)
-
Exploiting the aggregation propensity of beta-lactamases to design inhibitors that induce enzyme misfolding
Nature Communications (2023)
-
Data-mining unveils structure–property–activity correlation of viral infectivity enhancing self-assembling peptides
Nature Communications (2023)
-
Mechanistic insights into the aggregation pathway of the patient-derived immunoglobulin light chain variable domain protein FOR005
Nature Communications (2023)
-
Mechanisms and pathology of protein misfolding and aggregation
Nature Reviews Molecular Cell Biology (2023)