Abstract
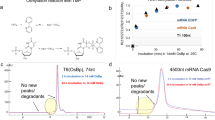
We developed a multiplex sequencing-by-synthesis method combining terminal phosphate–labeled fluorogenic nucleotides (TPLFNs) and resealable polydimethylsiloxane (PDMS) microreactors. In the presence of phosphatase, primer extension by DNA polymerase using nonfluorescent TPLFNs generates fluorophores, which are confined in the microreactors and detected. We immobilized primed DNA templates in the microreactors, then sequentially introduced one of the four identically labeled TPLFNs, sealed the microreactors and recorded a fluorescence image after template-directed primer extension. With cycle times of <10 min, we demonstrate 30 base reads with ∼99% raw accuracy. Our 'fluorogenic pyrosequencing' offers benefits of pyrosequencing, such as rapid turnaround, one-color detection and generation of native DNA, along with high detection sensitivity and simplicity of parallelization because simultaneous real-time monitoring of all microreactors is not required.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Margulies, M. et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature 437, 376–380 (2005).
Ronaghi, M., Karamohamed, S., Pettersson, B., Uhlen, M. & Nyren, P. Real-time DNA sequencing using detection of pyrophosphate release. Anal. Biochem. 242, 84–89 (1996).
Rothberg, J.M. et al. Methods and apparatus for measuring analytes using large-scale FET arrays. International patent WO 2010/008480 A2 (2010).
Bentley, D.R. et al. Accurate whole human genome sequencing using reversible terminator chemistry. Nature 456, 53–59 (2008).
Harris, T.D. et al. Single molecule DNA sequencing of a viral genome. Science 320, 106–109 (2008).
McKernan, K.J. et al. Sequence and structure variation in a human genome uncovered by short-read, massively parallel ligation sequencing using two-base encoding. Genome Res. 19, 1527–1541 (2009).
Shendure, J. et al. Accurate multiplex polony sequencing of an evolved bacterial genome. Science 309, 1728–1732 (2005).
Drmanac, R. et al. Human genome sequencing using unchained base reads on self-assembling DNA nanoarrays. Science 327, 78–81 (2010).
Eid, J. et al. Real-time DNA sequencing from single polymerase molecules. Science 323, 133–138 (2009).
Kumar, S. et al. Terminal phosphate-labeled nucleotides: synthesis, applications, and linker effect on incorporation by DNA polymerases. Nucleosides Nucleotides Nucleic Acids 24, 401–408 (2005).
Sood, A. et al. Terminal phosphate-labeled nucleotides with improved substrate properties for homogeneous nucleic acid assays. J. Am. Chem. Soc. 127, 2394–2395 (2005).
Kozlov, M., Bergendahl, V., Burgess, R., Goldfarb, A. & Mustaev, A. Homogeneous fluorescent assay for RNA polymerase. Anal. Biochem. 342, 206–213 (2005).
Rondelez, Y. et al. Microfabricated arrays of femtoliter chambers allow single molecule enzymology. Nat. Biotechnol. 23, 361–365 (2005).
Korlach, J. et al. Long, processive enzymatic DNA synthesis using 100% dye-labeled terminal phosphate-linked nucleotides. Nucleosides Nucleotides Nucleic Acids 27, 1072–1083 (2008).
Ankilova, V.N., Knorre, D.G., Kravchenko, V.V., Lavrik, O.I. & Nevinsky, G.A. Investigation of the phenylalanyl-tRNA synthetase modification with gamma-(p-azidoanilide-)-ATP. FEBS Lett. 60, 172–175 (1975).
Yarbrough, L.R. Synthesis and properties of a new fluorescent analog of ATP: adenosine-5′-triphosphoro-γ-1-(5-sulfonic acid) napthylamidate. Biochem. Biophys. Res. Commun. 81, 35–41 (1978).
Grachev, M.A. & Zaychikov, E.F. ATP gamma-anilidate: a substrate of DNA-dependent RNA-polymerase of Escherichia coli. FEBS Lett. 49, 163–166 (1974).
Takakusa, H., Kikuchi, K., Urano, Y., Kojima, H. & Negano, T. A novel design method of ratiometric fluorescent probes based on fluorescence resonance energy transfer switching by spectral overlap. Chem. Eur. J. 9, 1479–1485 (2003).
McDonald, J.C. & Whitesides, G.M. Poly(dimethylsiloxane) as a material for fabricating microfluidic devices. Acc. Chem. Res. 35, 491–499 (2002).
Piruska, A. et al. The autofluorescence of plastic materials and chips measured under laser irradiation. Lab Chip 5, 1348–1354 (2005).
Liu, D., Perdue, R.K., Sun, L. & Crooks, R.M. Immobilization of DNA onto poly(dimethylsiloxane) surfaces and application to a microelectrochemical enzyme-amplified DNA hybridization assay. Langmuir 20, 5905–5910 (2004).
Gorris, H.H., Rissin, D.M. & Walt, D.R. Stochastic inhibitor release and binding from single enzyme molecules. Proc. Natl. Acad. Sci. USA 104, 17680–17685 (2007).
Leamon, J.H. & Rothberg, J.M. Cramming more sequencing reactions onto microreactor chips. Chem. Rev. 107, 3367–3376 (2007).
Erlich, Y. et al. Alta-cyclic: a self-optimizing base caller for next-generation sequencing. Nat. Methods 5, 679–682 (2008).
Dressman, D., Yan, H., Traverso, G., Kinzler, K.W. & Vogelstein, B. Transforming single DNA molecules into fluorescent magnetic particles for detection and enumeration of genetic variations. Proc. Natl. Acad. Sci. USA 100, 8817–8822 (2003).
Hatch, A., Sano, T., Misasi, J. & Smith, C.L. Rolling circle amplification of DNA immobilized on solid surfaces and its application to multiplex mutation detection. Genet. Anal. 15, 35–40 (1999).
Marcus, J.S., Anderson, W.F. & Quake, S.R. Microfluidic single-cell mRNA isolation and analysis. Anal. Chem. 78, 3084–3089 (2006).
Fan, H.C., Wang, J., Potanina, A. & Quake, S.R. Whole-genome molecular haplotyping of single cells. Nat. Biotechnol. 29, 51–57 (2011).
Tewhey, R. et al. Microdroplet-based PCR enrichment for large-scale targeted sequencing. Nat. Biotechnol. 27, 1025–1031 (2009).
Frimat, J.P. et al. Plasma stenciling methods for cell patterning. Anal. Bioanal. Chem. 395, 601–609 (2009).
Acknowledgements
This work was supported by US National Institutes of Health National Human Genome Research Institute (HG005097-01) to X.S.X. and US National Institutes of Health National Human Genome Research Institute Recovery Act Grand Opportunities grant (1RC2HG005613-01) to X.S.X. W.J.G. is a Damon Runyon Fellow supported by the Damon Runyon Cancer Research Foundation (DRG-2000 09). We acknowledge J. Foley, S. Song, L. Song, G. Holtom and K. Fiala for valuable discussions and technical assistance. This work was performed in part at the Center for Nanoscale Systems, which is a part of Harvard University and a member of the National Nanotechnology Infrastructure Network, which is supported by the National Science Foundation award ECS-0335765.
Author information
Authors and Affiliations
Contributions
P.A.S., W.J.G. and X.S.X. conceived the fluorogenic pyrosequencing concept. P.A.S. and W.J.G. constructed the sequencing apparatus. H.D. synthesized the TPLFNs. P.A.S. collected and analyzed the data. W.J.G. carried out the microfabrication. P.A.S., W.J.G., H.D. and X.S.X. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
Harvard University has filed patent applications based on this work.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Table 1 and Supplementary Notes 1–3 (PDF 534 kb)
Rights and permissions
About this article
Cite this article
Sims, P., Greenleaf, W., Duan, H. et al. Fluorogenic DNA sequencing in PDMS microreactors. Nat Methods 8, 575–580 (2011). https://doi.org/10.1038/nmeth.1629
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1629