Abstract
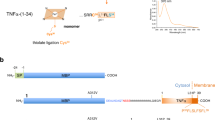
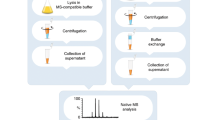
We developed a method for restricted enzymatic proteolysis using the outer membrane protease T (OmpT) to produce large peptides (>6.3 kDa on average) for mass spectrometry–based proteomics. Using this approach to analyze prefractionated high-mass HeLa proteins, we identified 3,697 unique peptides from 1,038 proteins. We demonstrated the ability of large OmpT peptides to differentiate closely related protein isoforms and to enable the detection of many post-translational modifications.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
de Godoy, L.M. et al. Nature 455, 1251–1254 (2008).
Tran, J.C. et al. Nature 480, 254–258 (2011).
Chait, B.T. Science 314, 65–66 (2006).
Nesvizhskii, A.I. & Aebersold, R. Mol. Cell. Proteomics 4, 1419–1440 (2005).
Forbes, A.J., Mazur, M.T., Patel, H.M., Walsh, C.T. & Kelleher, N.L. Proteomics 1, 927–933 (2001).
Tran, J.C. & Doucette, A.A. Anal. Chem. 80, 1568–1573 (2008).
Wu, S.L., Kim, J., Hancock, W.S. & Karger, B. J. Proteome Res. 4, 1155–1170 (2005).
Taouatas, N., Drugan, M.M., Heck, A.J. & Mohammed, S. Nat. Methods 5, 405–407 (2008).
Cannon, J. et al. J. Proteome Res. 9, 3886–3890 (2010).
Kramer, R.A., Zandwijken, D., Egmond, M.R. & Dekker, N. Eur. J. Biochem. 267, 885–893 (2000).
White, C.B., Chen, Q., Kenyon, G.L. & Babbitt, P.C. J. Biol. Chem. 270, 12990–12994 (1995).
Dekker, N., Cox, R.C., Kramer, R.A. & Egmond, M.R. Biochemistry 40, 1694–1701 (2001).
Okuno, K. et al. Biosci. Biotechnol. Biochem. 66, 127–134 (2002).
Vandeputte-Rutten, L. et al. EMBO J. 20, 5033–5039 (2001).
McCarter, J.D. et al. J. Bacteriol. 186, 5919–5925 (2004).
Olsen, J.V. et al. Mol. Cell. Proteomics 8, 2759–2769 (2009).
Dekker, N., Merck, K., Tommassen, J. & Verheij, H.M. Eur. J. Biochem. 232, 214–219 (1995).
Lee, J.E. et al. J. Am. Soc. Mass Spectrom. 20, 2183–2191 (2009).
Tran, J.C. & Doucette, A.A. Anal. Chem. 81, 6201–6209 (2009).
Kramer, R.A. et al. Eur. J. Biochem. 269, 1746–1752 (2002).
Wessel, D. & Flugge, U.I. Anal. Biochem. 138, 141–143 (1984).
Elias, J.E. & Gygi, S.P. Nat. Methods 4, 207–214 (2007).
Meng, F. et al. Nat. Biotechnol. 19, 952–957 (2001).
Benjamini, Y. & Hochberg, Y. J. R. Stat. Soc. Series B Stat. Methodol. 57, 289–300 (1995).
Storey, J.D. & Tibshirani, R. Proc. Natl. Acad. Sci. USA 100, 9440–9445 (2003).
Acknowledgements
We sincerely thank current and former members of the Kelleher and Sweedler research groups, especially B. Evans, P. Chu, P. Compton, A. Catherman, S. Sweet, I. Ntai, J. Lee, A. Vellaichamy and K. Catherman, for technical support and insightful suggestions, P. Yau and B. Imai from the Protein Sciences Facility at the University of Illinois for the synthesis of the fluorogenic substrate and M. Burke for the access to a fluorimeter. The project was supported by the US National Institutes of Health through awards R01 GM067193, P30 DA018310 and F30 DA026672, by the National Science Foundation through award DMS 0800631 and by the Chicago Biomedical Consortium with support from the Searle Funds at the Chicago Community Trust.
Author information
Authors and Affiliations
Contributions
C.W. designed research, conducted experiments, analyzed data and wrote the paper; J.C.T., L.Z., K.R.D. and B.P.E. analyzed data; M.L., K.R.D. and D.R.A. conducted cell culture; P.M.T. analyzed data and wrote the paper; J.V.S. interpreted data and wrote the paper; N.L.K. designed research, interpreted data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Note (PDF 2307 kb)
Supplementary Table 1
Identified unique OmpT peptide lists and unique protein counts from Protein Center report. Unique OmpT peptides identified from absolute mass and biomarker searches are listed separately as well as in a combined list after removing the redundant overlapped hits from both search modes. (XLS 9183 kb)
Supplementary Table 2
Pooled unique OmpT peptide identifications from the nanoLC-MS/MS injections for the CID and ETD comparisons. (XLS 2739 kb)
Rights and permissions
About this article
Cite this article
Wu, C., Tran, J., Zamdborg, L. et al. A protease for 'middle-down' proteomics. Nat Methods 9, 822–824 (2012). https://doi.org/10.1038/nmeth.2074
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2074
This article is cited by
-
Hybrid mass spectrometry approaches in glycoprotein analysis and their usage in scoring biosimilarity
Nature Communications (2016)
-
The growing landscape of lysine acetylation links metabolism and cell signalling
Nature Reviews Molecular Cell Biology (2014)
-
Quantitative proteomics reveals the kinetics of trypsin-catalyzed protein digestion
Analytical and Bioanalytical Chemistry (2014)
-
The emergence of top-down proteomics in clinical research
Genome Medicine (2013)
-
Digestion and depletion of abundant proteins improves proteomic coverage
Nature Methods (2013)