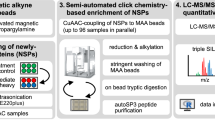
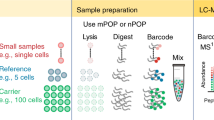
Abstract
Here we demonstrate quantitation of stimuli-induced proteome dynamics in primary cells by combining the power of bio-orthogonal noncanonical amino acid tagging (BONCAT) and stable-isotope labeling of amino acids in cell culture (SILAC). In conjunction with nanoscale liquid chromatography–tandem mass spectrometry (nanoLC-MS/MS), quantitative noncanonical amino acid tagging (QuaNCAT) allowed us to monitor the early expression changes of >600 proteins in primary resting T cells subjected to activation stimuli.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Ghazalpour, A. et al. PLoS Genet. 7, e1001393 (2011).
Foss, E.J. et al. Nat. Genet. 39, 1369–1375 (2007).
Rogers, S. et al. Bioinformatics 24, 2894–2900 (2008).
Schwanhausser, B. et al. Nature 473, 337–342 (2011).
Mann, M. Nat. Rev. Mol. Cell Biol. 7, 952–958 (2006).
Geiger, T. et al. Nat. Protoc. 6, 147–157 (2011).
Schwanhausser, B., Gossen, M., Dittmar, G. & Selbach, M. Proteomics 9, 205–209 (2009).
Dieterich, D.C., Link, A.J., Graumann, J., Tirrell, D.A. & Schuman, E.M. Proc. Natl. Acad. Sci. USA 103, 9482–9487 (2006).
Dieterich, D.C. et al. Nat. Protoc. 2, 532–540 (2007).
Van Kasteren, S.I., Kramer, H.B., Gamblin, D.P. & Davis, B.G. Nat. Protoc. 2, 3185–3194 (2007).
van Kasteren, S.I. et al. Nature 446, 1105–1109 (2007).
Szychowski, J. et al. J. Am. Chem. Soc. 132, 18351–18360 (2010).
Cox, J. & Mann, M. Nat. Biotechnol. 26, 1367–1372 (2008).
Smyth, G.K. in Bioinformatics and Computational Biology Solutions using R and Bioconductor (eds., Gentleman, R., Dudoit, S., Irizarry, R. & Huber, W.) 397–420 (Springer, New York, 2005).
Diehn, M. et al. Proc. Natl. Acad. Sci. USA 99, 11796–11801 (2002).
Eichelbaum, K., Winter, M., Diaz, M.B., Herzig, S. & Krijgsveld, J. Nat. Biotechnol. 30, 984–990 (2012).
The UniProt Consortium. Nucleic Acids Res. 40, D71–D75 (2012).
Cox, J. et al. J. Proteome Res. 11, 1794–1805 (2011).
Perkins, D.N., Pappin, D.J., Creasy, D.M. & Cottrell, J.S. Electrophoresis 20, 3551–3567 (1999).
Ihaka, R. & Gentleman, R.R. J. Comput. Graph. Statist. 5, 299–314 (1996).
Carrillo, B., Yanofsky, C., Laboissiere, S., Nadon, R. & Kearney, R.E. Bioinformatics 26, 98–103 (2010).
Smyth, G.K. & Speed, T.P. Methods 31, 265–273 (2003).
Smyth, G.K. Stat. Appl. Genet. Mol. Biol. 3, 3 (2004).
Benjamini, Y. & Hochberg, Y. J. R. Stat. Soc., B 57, 289–300 (1995).
Acknowledgements
This work was supported by Wellcome Trust grant WT094296MA and EU-FP7 'Sybilla' number 201106 to O.A.; D.C.D. was supported by a Deutsche Forschungsgemeinschaft Emmy Noether grant DI1512/1-1. B.G.D. and O.A. are supported by Royal Society Wolfson Research Merit awards. We thank P. Charles, S. Taylor and E. Giannoulatou for advice on the use of bioinformatics and statistics software, W. Paster and K. Nika for helpful suggestions and for critical feedback on the manuscript, and M. Selbach for helpful suggestions. V.G. was supported by a Ph.D. fellowship from the Biotechnology and Biological Sciences Research Council, O.B. by a Marie Curie Intra European Fellowship, B.B. by a Rhodes scholarship. This paper is dedicated to Jamie.
Author information
Authors and Affiliations
Contributions
A.J.M.H., B.T., D.C.T., V.G. and O.A. initially conceived the QuaNCAT strategy. A.J.M.H., V.G., K.K., G.E., B.T., D.C.T., O.B., B.B., B.G.D. and O.A. designed and optimized the QuaNCAT procedure. B.B. and O.B. synthesized reagents for the CuAAC reaction and associated cell labeling; D.C.D. provided the cleavable tag and an improved protocol for affinity purification; A.J.M.H., V.G., K.K. and G.E. performed cell stimulation, metabolic labeling, CuAAC reactions in T cell extracts, protein affinity purification; optimized CuAAC protocol was initially performed by B.B. and O.B. on A.J.M.H.'s initial cell extracts; K.K., G.E., B.M.K. and O.A. performed radioactive labeling; K.K. performed flow cytometry analysis. A.J.M.H., V.G., K.K., G.E. and B.T. carried out mass spectrometry experiments and data analysis; A.J.M.H., V.G., K.K., G.E., B.T., B.B., B.G.D. and O.A. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Table 1, Supplementary Note (PDF 2760 kb)
Supplementary Table 1
List of data set 1. (XLSX 125 kb)
Rights and permissions
About this article
Cite this article
Howden, A., Geoghegan, V., Katsch, K. et al. QuaNCAT: quantitating proteome dynamics in primary cells. Nat Methods 10, 343–346 (2013). https://doi.org/10.1038/nmeth.2401
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2401
This article is cited by
-
An integrated workflow for quantitative analysis of the newly synthesized proteome
Nature Communications (2023)
-
Modulation of cellular transcriptome and proteome composition by azidohomoalanine—implications on click chemistry–based secretome analysis
Journal of Molecular Medicine (2023)
-
Extracellular matrix dynamics: tracking in biological systems and their implications
Journal of Biological Engineering (2022)
-
A new strategy to uncover fragile X proteomic biomarkers using the nascent proteome of peripheral blood mononuclear cells (PBMCs)
Scientific Reports (2021)
-
CHCHD4 confers metabolic vulnerabilities to tumour cells through its control of the mitochondrial respiratory chain
Cancer & Metabolism (2019)