Abstract
Fluorescent Ca2+ reporters are widely used as readouts of neuronal activities. Here we designed R-CaMP2, a high-affinity red genetically encoded calcium indicator (GECI) with a Hill coefficient near 1. Use of the calmodulin-binding sequence of CaMKK-α and CaMKK-β in lieu of an M13 sequence resulted in threefold faster rise and decay times of Ca2+ transients than R-CaMP1.07. These features allowed resolving single action potentials (APs) and recording fast AP trains up to 20–40 Hz in cortical slices. Somatic and synaptic activities of a cortical neuronal ensemble in vivo were imaged with similar efficacy as with previously reported sensitive green GECIs. Combining green and red GECIs, we successfully achieved dual-color monitoring of neuronal activities of distinct cell types, both in the mouse cortex and in freely moving Caenorhabditis elegans. Dual imaging using R-CaMP2 and green GECIs provides a powerful means to interrogate orthogonal and hierarchical neuronal ensembles in vivo.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
11 June 2015
In the version of this article initially published, GenBank/EMBL/DDBJ accession codes for R-GECO2L and R-CaMP2 were omitted. These accession codes are KR149061 and KR149062, respectively. The error has been corrected in the HTML and PDF versions of the article.
References
O'Connor, D.H., Huber, D. & Svoboda, K. Reverse engineering the mouse brain. Nature 461, 923–929 (2009).
Ziv, Y. et al. Long-term dynamics of CA1 hippocampal place codes. Nat. Neurosci. 16, 264–266 (2013).
Miyawaki, A. et al. Fluorescent indicators for Ca2+ based on green fluorescent proteins and calmodulin. Nature 388, 882–887 (1997).
Nagai, T., Yamada, S., Tominaga, T., Ichikawa, M. & Miyawaki, A. Expanded dynamic range of fluorescent indicators for Ca2+ by circularly permuted yellow. Proc. Natl. Acad. Sci. USA 101, 10554–10559 (2004).
Nagai, T., Sawano, A., Park, E.S. & Miyawaki, A. Circularly permuted green fluorescent proteins engineered to sense Ca2+. Proc. Natl. Acad. Sci. USA 98, 3197–3202 (2001).
Baird, G.S., Zacharias, D.A. & Tsien, R.Y. Circular permutation and receptor insertion within green fluorescent proteins. Proc. Natl. Acad. Sci. USA 96, 11241–11246 (1999).
Nakai, J., Ohkura, M. & Imoto, K. A high signal-to-noise Ca2+ probe composed of a single green fluorescent protein. Nat. Biotechnol. 19, 137–141 (2001).
Zhao, Y. et al. An expanded palette of genetically encoded Ca2+ indicators. Science 333, 1888–1891 (2011).
Wang, Q., Shui, B., Kotlikoff, M.I. & Sondermann, H. Structural basis for calcium sensing by GCaMP2. Structure 16, 1817–1827 (2008).
Akerboom, J. et al. Crystal structures of the GCaMP calcium sensor reveal the mechanism of fluorescence signal change and aid rational design. J. Biol. Chem. 284, 6455–6464 (2009).
Akerboom, J. et al. Optimization of a GCaMP calcium indicator for neural activity imaging. J. Neurosci. 32, 13819–13840 (2012).
Sun, X.R. et al. Fast GCaMPs for improved tracking of neuronal activity. Nat. Commun. 4, 2170 (2013).
Chen, T.-W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Wardill, T.J. et al. A neuron-based screening platform for optimizing genetically-encoded calcium indicators. PLoS ONE 8, e77728 (2013).
Ohkura, M. et al. Genetically encoded green fluorescent Ca2+ indicators with improved detectability for neuronal Ca2+ signals. PLoS ONE 7, e51286 (2012).
Ohkura, M., Sasaki, T., Kobayashi, C., Ikegaya, Y. & Nakai, J. An improved genetically encoded red fluorescent Ca2+ indicator for detecting optically evoked action potentials. PLoS ONE 7, e39933 (2012).
Wu, J. et al. Improved orange and red Ca2+ indicators and photophysical considerations for optogenetic applications. ACS Chem. Neurosci. 4, 963–972 (2013).
Akerboom, J. et al. Genetically encoded calcium indicators for multi-color neural activity imaging and combination with optogenetics. Front. Mol. Neurosci. 6, 2 (2013).
Bito, H., Deisseroth, K. & Tsien, R.W. CREB phosphorylation and dephosphorylation: a Ca2+- and stimulus duration-dependent switch for hippocampal gene expression. Cell 87, 1203–1214 (1996).
Deisseroth, K., Bito, H. & Tsien, R.W. Signaling from synapse to nucleus: postsynaptic CREB phosphorylation during multiple forms of hippocampal synaptic plasticity. Neuron 16, 89–101 (1996).
Takemoto-Kimura, S. et al. Regulation of dendritogenesis via a lipid-raft-associated Ca2+/calmodulin-dependent protein kinase CLICK-III/CaMKIγ. Neuron 54, 755–770 (2007).
Truong, K. et al. FRET-based in vivo Ca2+ imaging by a new calmodulin-GFP fusion molecule. Nat. Struct. Biol. 8, 1069–1073 (2001).
Osawa, M. et al. A novel target recognition revealed by calmodulin in complex with kinase kinase. Nat. Struct. Biol. 6, 819–824 (1999).
Hendel, T. et al. Fluorescence changes of genetic calcium indicators and OGB-1 correlated with neural activity and calcium in vivo and in vitro. J. Neurosci. 28, 7399–7411 (2008).
Fujii, H. et al. Nonlinear decoding and asymmetric representation of neuronal input information by CaMKIIα and calcineurin. Cell Reports 3, 978–987 (2013).
Ageta-Ishihara, N. et al. Control of cortical axon elongation by a GABA-driven Ca2+/calmodulin-dependent protein kinase cascade. J. Neurosci. 29, 13720–13729 (2009).
O'Connor, D.H., Peron, S.P., Huber, D. & Svoboda, K. Neural activity in barrel cortex underlying vibrissa-based object localization in mice. Neuron 67, 1048–1061 (2010).
Petersen, C.C.H. & Crochet, S. Synaptic computation and sensory processing in neocortical layer 2/3. Neuron 78, 28–48 (2013).
Tada, M., Takeuchi, A., Hashizume, M., Kitamura, K. & Kano, M. A highly sensitive fluorescent indicator dye for calcium imaging of neural activity in vitro and in vivo. Eur. J. Neurosci. 39, 1720–1728 (2014).
Taniguchi, H. et al. A resource of Cre driver lines for genetic targeting of GABAergic neurons in cerebral cortex. Neuron 71, 995–1013 (2011).
Gibson, J.R., Beierlein, M. & Connors, B.W. Two networks of electrically coupled inhibitory neurons in neocortex. Nature 402, 75–79 (1999).
Matsushita, M. & Narin, A.C. Characterization of the mechanism of regulation of Ca2+/ calmodulin-dependent protein kinase I by calmodulin and by Ca2+/calmodulin-dependent protein kinase kinase. J. Biol. Chem. 273, 21473–21481 (1998).
Callaway, E.M. Transneuronal circuit tracing with neurotropic viruses. Curr. Opin. Neurobiol. 18, 617–623 (2008).
Luo, L., Callaway, E.M. & Svoboda, K. Genetic dissection of neural circuits. Neuron 57, 634–660 (2008).
Pivetta, C., Esposito, M.S., Sigrist, M. & Arber, S. Motor-circuit communication matrix from spinal cord to brainstem neurons revealed by developmental origin. Cell 156, 537–548 (2014).
Kawashima, T. et al. Functional labeling of neurons and their projections using the synthetic activity-dependent promoter E-SARE. Nat. Methods 10, 889–895 (2013).
Kawashima, T., Okuno, H. & Bito, H. A new era for functional labeling of neurons: activity-dependent promoters have come of age. Front. Neural Circuits 8, 37 (2014).
Barth, A.L. Visualizing circuits and systems using transgenic reporters of neural activity. Curr. Opin. Neurobiol. 17, 567–571 (2007).
Xu, C. & Webb, W.W. Measurement of two-photon excitation cross sections of molecular fluorophores with data from 690 to 1050 nm. J. Opt. Soc. Am. B 13, 481–491 (1996).
Margrie, T.W. et al. Targeted whole-cell recordings in the mammalian brain in vivo. Neuron 39, 911–918 (2003).
Gengyo-Ando, K., Yoshina, S., Inoue, H. & Mitani, S. An efficient transgenic system by TA cloning vectors and RNAi for C. elegans. Biochem. Biophys. Res. Commun. 349, 1345–1350 (2006).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974).
Kawano, T. et al. An imbalancing act: gap junctions reduce the backward motor circuit activity to bias C. elegans for forward locomotion. Neuron 72, 572–586 (2011).
Acknowledgements
We thank D. Kim (Howard Hughes Medical Institute Janelia Farm Research Campus) for kindly providing access to GCaMP6f and GCaMP6s before publication (through Addgene); R. Campbell (University of Alberta) and T. Nagai (Osaka University) for R-GECO1 (through Addgene); A. Gottschalk (Frankfurt Institute for Molecular Life Sciences) and S. Ramanathan (Harvard University) for access to Psra-6::ChR2(H134R)::mCherry plasmid; N. Yamaguchi (Medical Scientist Training Program of the Faculty of Medicine, the University of Tokyo and Department of Neurochemistry, the University of Tokyo Graduate School of Medicine) for access to Tet-on vectors before publication; T. Takahashi and Y. Kita for access to fluorescence measurement stations at the Life Sciences Core Facility, Graduate School of Medicine, the University of Tokyo. We thank all of the members of H.B.'s laboratory for support and discussion. We are particularly indebted to Y. Kondo, Y. Dobashi, R. Gyobu and T. Kinbara for assistance, as well as Y. Kishida, J. Tsujimoto and D. Shuto for animal care. This work was supported in part by grants from Core Research for Evolutional Science and Technology Japan Science and Technology Agency (CREST-JST) (to H.B.), the Strategic International Research Cooperative Program Japan-Mexico (SICPME-JST) (to H.B.), Precursory Research for Embryonic Science and Technology (PRESTO)-JST (to S.T.-K. and K.K.), grants-in-aid KAKENHI from the Japanese Ministry of Education, Culture, Sports, Science and Technology (MEXT) and Japan Society for the Promotion of Science (JSPS) (to H.F., S.T.-K., M.O., K.G.-A., M.K., J.N., K.K. and H.B.), a Comprehensive Brain Science Network (CBSN) grant from MEXT (to H.B.), a Strategic Research Program for Brain Sciences (to M.K.) and grants from the Tokyo Society of Medical Sciences (to H.F.) and the Uehara Foundation (to H.B.). M.I., S.H. and S.K. were supported in part by JSPS fellowships.
Author information
Authors and Affiliations
Contributions
M.I. and H.B. conceived the original ckkap swapping experiments. M.I. designed the sensors and performed their mutagenesis screening in HEK293T. M.I., M.O. and J.N. performed in vitro characterizations. S.H. performed in utero electroporation. M.I. performed virus constructions, productions and injections. A.T., K.K. and M.K. performed electrophysiological recordings, and A.T., M.I. and K.K. performed two-photon imaging experiments in the barrel cortex. M.I. and S.H. performed histological analyses. H.F., S.K. and S.T.-K. provided critical guidance with the R-CaMP2 design and assisted with the mouse imaging experiments. K.G.-A. and J.N. performed worm imaging and data analysis. H.B. supervised the entire project. M.I. and H.B. wrote the paper. All authors discussed and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
H.B. is a scientific consultant for Chugai Pharmaceuticals Co., Ltd. M.I., A.T., M.O., J.N. and H.B. have filed a patent related to R-CaMP2.
Integrated supplementary information
Supplementary Figure 1 In vitro screening of recombinant R-CaMP2 variants.
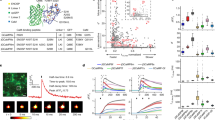
Baseline fluorescence compared to R-CaMP1.07 at nominally zero calcium plotted versus dynamic range (ΔF/F) for 150 recombinant candidate sensors (gray dots). As a reference, the values for R-GECO1 (orange), R-CaMP1.07 (blue), R-CaMP2 (black), and R-GECO2L (red) are indicated.
Supplementary Figure 2 Fluorescence spectra of R-CaMP2 and R-GECO2L.
(a,b) Normalized fluorescence excitation (dashed lines) and emission (solid lines) spectra of Ca2+-free (bottom traces) and Ca2+- saturated (upper traces) R-CaMP2 (a) and R-GECO2L (b). The fluorescence intensity of each indicator was normalized to the peaks of the Ca2+-saturated spectra. (c) Two-photon cross-section (ω) spectra of R-CaMP2 in the Ca2+-free (bottom dotted traces) or Ca2+-saturated (upper solid traces) state. Error bar, s.e.m. (n = 3).
Supplementary Figure 3 pH titration of normalized fluorescence intensity and dynamic change (ΔF/F) of red GECIs.
(a-c) pH titration of normalized fluorescence intensity of Ca2+-free (black) and Ca2+-saturated (red) states. The fluorescence intensity of each indicator was normalized to the peak of the calcium-saturated fluorescence intensity. Gray line indicates the dynamic range (ΔF/F). Experimental pKa values are summarized in Table 1. (a) R-GECO1 (b) R-CaMP1.07 (c) R-CaMP2
Supplementary Figure 4 Stopped-flow kinetic characterization of R-CaMP1.07 and R-CaMP2.
Observed relaxation rate constants (Kobs) plotted as a function of Ca2+ concentrations. The association and dissociation rate constants and the Hill coefficients (Kon, Koff and n, respectively) were determined by fitting to the equation Kobs = Kon × [Ca2+]n + Koff; these values are shown in Supplementary Table 2.
Supplementary Figure 5 Comparison of R-CaMP sensors during 1AP-triggered Ca2+ imaging of synaptic boutons.
(a) Representative image of synaptic boutons in cultured hippocampal neuron expressing R-CaMP2 (top) and R-CaMP1.07 (bottom), 50 msec after one field stimulus evoked one action potential. Left: EGFP (in monochrome). Right: Pseudocolor image of R-CaMP2 and R-CaMP1.07. Scale bar, 5 μm. (b) Single trials (gray) and trial-averaged responses of R-CaMP2 (black), R-GECO2L (red), and R-CaMP1.07 (blue) in response to a single AP. (c) Amplitude, SNR, rise times, and decay time constants for a single AP. Error bars, s.e.m. (n = 13, 8, 17 dishes sampled from 81, 52, 108 synaptic boutons for R-CaMP2, R-GECO2L, R-CaMP1.07, respectively, *P < 0.05, **P < 0.01, ***P < 0.001 in Tukey's post-hoc test following one-way ANOVA. Individual P values: Amplitude, 9.9 × 10-5 (R-CaMP2/R-GECO2L), 4.7 × 10-14 (R-CaMP2/RCaMP1.07); SNR, 1.8 × 10-8 (R-CaMP2/R-GECO2L), 3.1 × 10-13 (R-CaMP2/R-CaMP1.07); Rise time, 1.1 × 10-2 (R-CaMP2/RCaMP1.07).
Supplementary Figure 6 Comparison of R-CaMP2 and R-CaMP1.07 during a single Glu photolysis-induced somatic AP-triggered Ca2+ imaging at the soma in cultured hippocampal neurons.
(a) Single trials (n = 9, gray) and trial-averaged responses of R-CaMP2 (left, black) and R-CaMP1.07 (right, blue) in response to a single pulse of MNI-glutamate uncaging. (b) Amplitude, SNR, rise times, and decay time constants for a single pulse of MNI-glutamate uncaging. Error bars, s.e.m. (n = 9 cells each, *P < 0.05, ***P < 0.001 in Welch's t-test. Individual P values: Amplitude, 8.0 × 10-4; SNR, 4.8 × 10-2; Rise time, 2.6 × 10-2; Decay time, 2.7 × 10-7.
Supplementary Figure 7 Comparison of R-CaMP2 and R-CaMP1.07 during a 1AP-evoked Ca2+ transient in acute cortical slices.
(a) Amplitude, (b) SNR, (c) rise times, and (d) decay time constants of somatic Ca2+ transients induced by a single AP. Error bars, s.e.m. (n = 10 cells each, ***P < 0.001 in Welch's t-test. Individual P values: Amplitude, 2.0 × 10-4; SNR, 4.3 × 10-6; Rise time, 1.3 × 10-7; Decay time, 3.0 × 10-11.
Supplementary Figure 8 Trial-averaged fluorescence transients of R-CaMP2 and R-CaMP1.07 to five spike trains delivered at different frequencies.
(a,b) Trial-averaged data from experiments in Fig. 3d,e. Gray lines indicate individual sweeps (n = 10). Black and blue thick lines indicate average traces of R-CaMP2 (a) and R-CaMP1.07 (b) Right side of both figures: responses to 10, 20, and 40 Hz spike trains are shown in an expanded time scale. Stimulus time stamps are indicated by the vertical bars and dotted lines.
Supplementary Figure 9 Persistent R-CaMP2 expression does not affect passive membrane properties or Ca2+ transient kinetics.
(a) Representative traces of membrane potential deflections induced by current injection. Control (left, untransfected contralateral hemisphere), CAG-R-CaMP2 expressing neuron (right) (b) Resting membrane potential (left), membrane capacitance (middle), membrane input resistance (right) were unchanged between groups. Error bars, s.e.m. (n = 18, 17 cells for control (untransfected contralateral hemisphere) neuron, and CAG-R-CaMP2 expressing neuron, respectively, P = 0.824, 0.365, 0.588 respectively, ns (not significant), Welch's t-test). (c) Representative traces of fluorescence changes (ΔF/F) in response to a single AP in neurons expressing either a TRE or a CAG promoter-driven R-CaMP2 in acute slices. Ten-trial average response of a single sweep each (gray) and averages of 10 sweeps (black) are overlaid (n = 10, 8 cells for TRE-R-CaMP2, CAG-R-CaMP2, respectively). (d) Either TRE or CAG promoter-driven R-CaMP2-expressing neurons showed undistinguishable single AP-evoked Ca2+ transient kinetics.
Supplementary Figure 10 Ectopic R-CaMP2 expression has little effect on radial migration and does not cause toxic cell death.
Mouse embryo were electroporated at E14 with a plasmid expressing control GCaMP6f (a,d: control) or R-CaMP2 (b,e: R-CaMP2), and sacrificed at E18 (a-c) or P18 (d-f). Coronal brain slices were imaged for baseline fluorescence (white) and immunostained with activated-caspase-3 (red). White arrowheads indicate labeled activated-caspase-3 cell. Scale bars, 100 μm (a,b) and 200 μm (d,e) respectively. (c,f) Quantification of the transfected cells that are activated-caspase-3 positive (Cas3+). Error bars, s.e.m. (n = 9182, 8242 cells for control (GCaMP6f), R-CaMP2 from 7 pairs of littermate animals (E18) for each condition, P = 0.736 (c), ns (not significant), Welch's ttest). (n = 5171, 6302 cells for control (GCaMP6f), R-CaMP2 from 6 pairs of littermate animals (P18) for each condition, P = 0.978 (f), ns (not significant), Welch's t-test).
Supplementary Figure 11 Simultaneous Ca2+ imaging and electrophysiology in neocortical layer 2/3 neurons in vivo.
(a) Representative traces of simultaneous recordings of calcium transients (top) and APs (bottom) in an R-CaMP2-expressing layer 2/3 neuron in the barrel cortex. The number of spikes for each burst is indicated below the trace. Scale bar, 5 μm. (b) Amplitude, SNR, and integral of calcium transients induced by number of APs in a 200 msec bin (n = 254, 115, 45, 26, 13 events for 1, 2, 3, 4, 5 action potentials, n = 9 cells in 7 mice). Error bars, s.e.m. (c) Median fluorescence change in response to a single action potential in vivo. Shading indicates s.e.m. (n = 9 cells). (d) The rise time and decay time constants of calcium transients in vivo induced by a single AP measured with cell-attached recording. Error bars, s.e.m.
Supplementary Figure 12 All-optical interrogation of an avoidance circuit using R-CaMP2 in freely moving C. elegans.
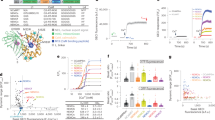
(a) Schematic diagram of the ASH neuron-mediated avoidance circuit. (b) Representative R-CaMP2 response images in a freely moving transgenic animal jqEx467, coexpressing R-CaMP2 (red) and EGFP (green) in AVA neurons (AVA::R-CaMP2), and expressing ChR2 in ASH neurons (ASH::ChR2). R-CaMP2 and EGFP signals were also weakly detected in a few other cells including AVE and AVD neurons. Arrowhead indicates AVA neuron. Bar 20 μm. (c) Behavioral responses of the jqEx467 worms with or without all-trans-retinal (ATR). The observable backward response during the blue light illumination was counted. ***P < 0.001 in Tukey's post-hoc test following one-way ANOVA. Individual P values: 1.2 × 10-4 (Wild type without ATR/ASH::ChR2; AVA::R-CaMP2 with ATR), 5.3 × 10-5 (with/without ATR in ATR/ASH::ChR2; AVA::R-CaMP2 background). (d) Causal relation between blue light-activated ASH neuronal activity (ChR2 on, light blue shade), AVA activity (R-CaMP2, blue trace) and backward movement (behavioral response). R-CaMP2 traces and backward responses of a jqEx467 worm. Gray line shows a raw data of R-CaMP2/EGFP ratio image (ΔR/R) and black line shows a moving average of across 1 sec–window.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12 and Supplementary Tables 1 and 2 (PDF 4413 kb)
Recording of population activities in layer 2/3 neurons in the barrel cortex in vivo using R-CaMP2.
This movie shows in vivo population calcium imaging of barrel cortex layer 2/3 neurons expressing R-CaMP2. Air-puff stimuli (40–45 psi, 50 msec) were delivered at 5 sec intervals during an imaging session of 140 sec. (AVI 9227 kb)
Recording of spontaneous dendritic spine calcium transients in R-CaMP2-expressing layer 2/3 barrel neuron in vivo (raw fluorescence data).
This movie shows Ca2+ transients in dendritic spines and shafts of barrel cortex layer 2/3 neurons expressing R-CaMP2 in vivo. Raw fluorescence data are shown. (AVI 1089 kb)
Recording of dendritic spine calcium transients in R-CaMP2-expressing layer 2/3 barrel neuron in vivo (ΔF/F data).
Same dataset as Movie 2, but ΔF/F signals are shown. (AVI 1015 kb)
41592_2015_BFnmeth3185_MOESM40_ESM.avi
Simultaneous recording of R-CaMP2–expressing SST-positive interneurons and GCaMP6f-expressing CaMKII α-positive excitatory neurons in virally infected layer 2/3 neurons in vivo. (AVI 6716 kb)
41592_2015_BFnmeth3185_MOESM41_ESM.avi
Simultaneous recording of GCaMP6s-expressing SST-positive interneurons and R-CaMP2–expressing CaMKII α-positive excitatory neurons in virally infected layer 2/3 neurons in vivo. (AVI 7119 kb)
Simultaneous recording of optical control and dual-color Ca2+ imaging of the locomotor circuit in freely moving C. elegans.
This video shows in vivo calcium imaging and photomanipulation of a transgenic worm coexpressing G-CaMP6 and ChR2 in GABAergic neurons and expressing R-CaMP2 in body wall muscles. Blue light was delivered at 6-11 sec. (AVI 4328 kb)
Rights and permissions
About this article
Cite this article
Inoue, M., Takeuchi, A., Horigane, Si. et al. Rational design of a high-affinity, fast, red calcium indicator R-CaMP2. Nat Methods 12, 64–70 (2015). https://doi.org/10.1038/nmeth.3185
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3185
This article is cited by
-
Large-scale recording of neuronal activity in freely-moving mice at cellular resolution
Nature Communications (2023)
-
Adaptable, turn-on maturation (ATOM) fluorescent biosensors for multiplexed detection in cells
Nature Methods (2023)
-
Fastest-ever calcium sensors broaden the potential of neuronal imaging
Nature (2023)
-
Mitochondrial Calcium Waves by Electrical Stimulation in Cultured Hippocampal Neurons
Molecular Neurobiology (2023)
-
Fast and sensitive GCaMP calcium indicators for imaging neural populations
Nature (2023)