Abstract
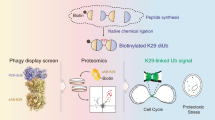
Ubiquitination controls a plethora of cellular processes. Modifications by linear polyubiquitin have so far been linked with acquired and innate immunity, lymphocyte development and genotoxic stress response. Until now, a single E3 ligase complex (LUBAC), one specific deubiquitinase (OTULIN) and a very few linear polyubiquitinated substrates have been identified. Current methods for studying lysine-based polyubiquitination are not suitable for the detection of linear polyubiquitin-modified proteins. Here, we present an approach to discovering linear polyubiquitin-modified substrates by combining a lysine-less internally tagged ubiquitin (INT-Ub.7KR) with SILAC-based mass spectrometry. We applied our approach in TNFα-stimulated T-REx HEK293T cells and validated several newly identified linear polyubiquitin targets. We demonstrated that linear polyubiquitination of the novel LUBAC substrate TRAF6 is essential for NFκB signaling.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Varshavsky, A. Regulated protein degradation. Trends Biochem. Sci. 30, 283–286 (2005).
Hershko, A. & Ciechanover, A. The ubiquitin system. Annu. Rev. Biochem. 67, 425–479 (1998).
Kulathu, Y. & Komander, D. Atypical ubiquitylation — the unexplored world of polyubiquitin beyond Lys48 and Lys63 linkages. Nat. Rev. Mol. Cell Biol. 13, 508–523 (2012).
Kirisako, T. et al. A ubiquitin ligase complex assembles linear polyubiquitin chains. EMBO J. 25, 4877–4887 (2006).
Matsumoto, M.L. et al. Engineering and structural characterization of a linear polyubiquitin-specific antibody. J. Mol. Biol. 418, 134–144 (2012).
Phu, L. et al. Improved quantitative mass spectrometry methods for characterizing complex ubiquitin signals. Mol. Cell. Proteomics 10, M110.003756 (2011).
Tokunaga, F. et al. Involvement of linear polyubiquitylation of NEMO in NF-kappaB activation. Nat. Cell Biol. 11, 123–132 (2009).
Asaoka, T. et al. Linear ubiquitination by LUBEL has a role in Drosophila heat stress response. EMBO Rep. 17, 1624–1640 (2016).
Damgaard, R.B. et al. The ubiquitin ligase XIAP recruits LUBAC for NOD2 signaling in inflammation and innate immunity. Mol. Cell 46, 746–758 (2012).
Shimizu, Y., Taraborrelli, L. & Walczak, H. Linear ubiquitination in immunity. Immunol. Rev. 266, 190–207 (2015).
Tokunaga, F. Linear ubiquitination-mediated NF-κB regulation and its related disorders. J. Biochem. 154, 313–323 (2013).
Okamura, K. et al. Survival of mature T cells depends on signaling through HOIP. Sci. Rep. 6, 36135 (2016).
Teh, C.E. et al. Linear ubiquitin chain assembly complex coordinates late thymic T-cell differentiation and regulatory T-cell homeostasis. Nat. Commun. 7, 13353 (2016).
Sasaki, Y. et al. Defective immune responses in mice lacking LUBAC-mediated linear ubiquitination in B cells. EMBO J. 32, 2463–2476 (2013).
Smit, J.J. et al. The E3 ligase HOIP specifies linear ubiquitin chain assembly through its RING-IBR-RING domain and the unique LDD extension. EMBO J. 31, 3833–3844 (2012).
Stieglitz, B. et al. Structural basis for ligase-specific conjugation of linear ubiquitin chains by HOIP. Nature 503, 422–426 (2013).
Ikeda, F. et al. SHARPIN forms a linear ubiquitin ligase complex regulating NF-κB activity and apoptosis. Nature 471, 637–641 (2011).
Gerlach, B. et al. Linear ubiquitination prevents inflammation and regulates immune signalling. Nature 471, 591–596 (2011).
Tokunaga, F. et al. SHARPIN is a component of the NF-κB-activating linear ubiquitin chain assembly complex. Nature 471, 633–636 (2011).
Keusekotten, K. et al. OTULIN antagonizes LUBAC signaling by specifically hydrolyzing Met1-linked polyubiquitin. Cell 153, 1312–1326 (2013).
Rivkin, E. et al. The linear ubiquitin-specific deubiquitinase gumby regulates angiogenesis. Nature 498, 318–324 (2013).
Fiil, B.K. et al. OTULIN restricts Met1-linked ubiquitination to control innate immune signaling. Mol. Cell 50, 818–830 (2013).
Rodgers, M.A. et al. The linear ubiquitin assembly complex (LUBAC) is essential for NLRP3 inflammasome activation. J. Exp. Med. 211, 1333–1347 (2014).
Satpathy, S. et al. Systems-wide analysis of BCR signalosomes and downstream phosphorylation and ubiquitylation. Mol. Syst. Biol. 11, 810 (2015).
Emmerich, C.H. et al. Activation of the canonical IKK complex by K63/M1-linked hybrid ubiquitin chains. Proc. Natl. Acad. Sci. USA 110, 15247–15252 (2013).
Smahi, A. et al. Genomic rearrangement in NEMO impairs NF-kappaB activation and is a cause of incontinentia pigmenti. Nature 405, 466–472 (2000).
Döffinger, R. et al. X-linked anhidrotic ectodermal dysplasia with immunodeficiency is caused by impaired NF-kappaB signaling. Nat. Genet. 27, 277–285 (2001).
Filipe-Santos, O. et al. X-linked susceptibility to mycobacteria is caused by mutations in NEMO impairing CD40-dependent IL-12 production. J. Exp. Med. 203, 1745–1759 (2006).
Xu, G., Paige, J.S. & Jaffrey, S.R. Global analysis of lysine ubiquitination by ubiquitin remnant immunoaffinity profiling. Nat. Biotechnol. 28, 868–873 (2010).
Wagner, S.A. et al. A proteome-wide, quantitative survey of in vivo ubiquitylation sites reveals widespread regulatory roles. Mol. Cell. Proteomics 10, M111.013284 (2011).
Kim, W. et al. Systematic and quantitative assessment of the ubiquitin-modified proteome. Mol. Cell 44, 325–340 (2011).
Danielsen, J.M.R. et al. Mass spectrometric analysis of lysine ubiquitylation reveals promiscuity at site level. Mol. Cell. Proteomics 10, M110.003590 (2011).
Meierhofer, D., Wang, X., Huang, L. & Kaiser, P. Quantitative analysis of global ubiquitination in HeLa cells by mass spectrometry. J. Proteome Res. 7, 4566–4576 (2008).
Peng, J. et al. A proteomics approach to understanding protein ubiquitination. Nat. Biotechnol. 21, 921–926 (2003).
Rahighi, S. et al. Specific recognition of linear ubiquitin chains by NEMO is important for NF-kappaB activation. Cell 136, 1098–1109 (2009).
Baker, R.T. et al. Using deubiquitylating enzymes as research tools. Methods Enzymol. 398, 540–554 (2005).
Povlsen, L.K. et al. Systems-wide analysis of ubiquitylation dynamics reveals a key role for PAF15 ubiquitylation in DNA-damage bypass. Nat. Cell Biol. 14, 1089–1098 (2012).
Haas, T.L. et al. Recruitment of the linear ubiquitin chain assembly complex stabilizes the TNF-R1 signaling complex and is required for TNF-mediated gene induction. Mol. Cell 36, 831–844 (2009).
Takiuchi, T. et al. Suppression of LUBAC-mediated linear ubiquitination by a specific interaction between LUBAC and the deubiquitinases CYLD and OTULIN. Genes Cells 19, 254–272 (2014).
Lewis, M.J. et al. UBE2L3 polymorphism amplifies NF-κB activation and promotes plasma cell development, linking linear ubiquitination to multiple autoimmune diseases. Am. J. Hum. Genet. 96, 221–234 (2015).
Müller-Rischart, A.K. et al. The E3 ligase Parkin maintains mitochondrial integrity by increasing linear ubiquitination of NEMO. Mol. Cell 49, 908–921 (2013).
Khan, M., Syed, G.H., Kim, S.J. & Siddiqui, A. Hepatitis B virus-induced Parkin-dependent recruitment of linear ubiquitin assembly complex (LUBAC) to mitochondria and attenuation of innate immunity. PLoS Pathog. 12, e1005693 (2016).
Lamothe, B. et al. Site-specific Lys-63-linked tumor necrosis factor receptor-associated factor 6 auto-ubiquitination is a critical determinant of I kappa B kinase activation. J. Biol. Chem. 282, 4102–4112 (2007).
Yang, W.L. et al. The E3 ligase TRAF6 regulates Akt ubiquitination and activation. Science 325, 1134–1138 (2009).
Zhang, M. et al. HDAC6 deacetylates and ubiquitinates MSH2 to maintain proper levels of MutSα. Mol. Cell 55, 31–46 (2014).
Deng, L. et al. Activation of the IkappaB kinase complex by TRAF6 requires a dimeric ubiquitin-conjugating enzyme complex and a unique polyubiquitin chain. Cell 103, 351–361 (2000).
Aguileta, M.A. et al. The E3 ubiquitin ligase Parkin is recruited to the 26 S proteasome via the proteasomal ubiquitin receptor Rpn13. J. Biol. Chem. 290, 7492–7505 (2015).
Bremm, A. & Komander, D. Synthesis and analysis of K11-linked ubiquitin chains. Methods Mol. Biol. 832, 219–228 (2012).
Lin, D.Y., Diao, J., Zhou, D. & Chen, J. Biochemical and structural studies of a HECT-like ubiquitin ligase from Escherichia coli O157:H7. J. Biol. Chem. 286, 441–449 (2011).
Umebayashi, K., Stenmark, H. & Yoshimori, T. Ubc4/5 and c-Cbl continue to ubiquitinate EGF receptor after internalization to facilitate polyubiquitination and degradation. Mol. Biol. Cell 19, 3454–3462 (2008).
Stieglitz, B., Morris-Davies, A.C., Koliopoulos, M.G., Christodoulou, E. & Rittinger, K. LUBAC synthesizes linear ubiquitin chains via a thioester intermediate. EMBO Rep. 13, 840–846 (2012).
Catanzariti, A.M., Soboleva, T.A., Jans, D.A., Board, P.G. & Baker, R.T. An efficient system for high-level expression and easy purification of authentic recombinant proteins. Prot. Sci. 13, 1331–1339 (2004).
Licchesi, J.D. et al. An ankyrin-repeat ubiquitin-binding domain determines TRABID's specificity for atypical ubiquitin chains. Nat. Struct. Mol. Biol. 19, 62–71 (2011).
Martin, S.R. & Schilstra, M.J. Circular dichroism and its application to the study of biomolecules. Methods Cell Biol. 84, 263–293 (2008).
Shevchenko, A., Tomas, H., Havlis, J., Olsen, J.V. & Mann, M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat. Protoc. 1, 2856–2860 (2006).
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2, 1896–1906 (2007).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Vizcaíno, J.A. et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 44, D447–D456 (2016).
Acknowledgements
We thank M. Asada (Nippon Medical School), R.T. Baker (Clinical Genomics), A. Bremm (Goethe University School of Medicine), C. Behrends (Goethe University School of Medicine), J. Chen (Rockefeller University), K.-H. Chun (Yonsei University College of Medicine), I. Dikic (Goethe University School of Medicine), H. Jiang (National Institute of Biological Sciences), J.U. Jung (University of Southern California), M. Kobayashi (Kanazawa University), C. White (Rosalind Franklin University of Medicine and Science) and A. Wittinghofer (Max Planck Institute of Molecular Physiology) for providing reagents and M. Akutsu, A. Carpy, J. Lopez-Mosqueda, M. Olma and S. Wahl for initial help with the project. We thank S. Schaubeck for excellent technical assistance. We are especially grateful to J.-I. Inoue (University of Tokyo) for providing us with TRAF6−/− MEFs and J.W. Bowman for providing us with the detailed LUBAC purification protocol. We thank A. Bremm, J. Lopez-Mosqueda and B. Srinivasan for discussions, comments and reading of the manuscript. We would also like to thank Reviewer 1 for their comments, which helped us to substantially improve the manuscript. K.K. was supported by the UPStream grant (EU, FP7, ITN project 290257).
Author information
Authors and Affiliations
Contributions
K.H. and K.K. developed the concept and designed the experiments. K.H. developed the INT-Ub tool, prepared the constructs and performed initial validation experiments. K.K. optimized the method, carried out all the cell biology and biochemical experiments, prepared all the inducible cell lines and samples for MS experiments and analyzed the results, including MS data. B.M., C.T. and M.F.-W. generated and analyzed MS data. B.S., I.P. and S.K. performed and analyzed CD and NMR experiments. K.H. and K.K. wrote the manuscript with contribution from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 The design and in vitro validation of INT-Ub.
(A) Ub chain building potential of various Ub mutants (top). Ub molecule with an insertion point of STREP-tag II (black) and the following color-coded residues: K (red), M (orange), G76 (blue) and I44 (yellow). This Figure was generated with PyMOL software by modifying a 1UBQ.pdb file (bottom). (B) Coomassie staining of recombinant Ub variants used for CD measurements in (C) Far-UV CD spectra of wild-type Ub (blue), Ub.7KR (red), INT-Ub (green) and INT-Ub.7KR (violet). (D) and (E) A direct comparison of in vitro ubiquitination reaction for untagged Ub, INT-Ub and INT-Ub.ΔGG. (F) Pull-down assay between INT-Ub and UBDs that can recognise single Ub moieties. (G) A direct comparison of diUb and INT-diUb ability to bind linear polyUb-specific UBD UBAN. (H) A direct comparison of USP2-cc DUB assay with diUb and INT-diUb. (I) A direct comparison of OTULIN DUB assay with diUb and INT-diUb. Blot and gel images for Supplementary Figure 1b,f-i were cropped to improve the conciseness of the presentation (full blots and gel images are available in Supplementary Figure 12).
Supplementary Figure 2 In vivo validation of INT-Ub.
(A) A comparison of INT-Ub (left panel), INT-Ub.7KR (middle panel), INT-Ub.ΔGG (right panel) and endogenous Ub levels in T-REx HEK293T cells upon doxycycline induction. To visualise levels of unconjugated Ub proteins, lysates were treated with USP2-cc. (B) A comparison of the effect of untagged Ub and Ub.ΔGG on NFκB transcriptional activity (control for Figure 2f). NFκB transcriptional activity was measured by a luciferase assay. Results are shown as means and s.e.m. (n=3). n.s=no statistically significant difference (p>0.05), determined by the two tailed Student’s t-test. (C) The effect of OTULIN C129A overexpression on the abundance of HMW linear polyubiquitinated proteins, was estimated by the linear Ub-specific Lub9 antibody upon denaturing INT-Ub.7KR Strep-Tactin PD. (D) The effect of TNFa stimulation (15 min) on the appearance of HMW ubiquitinated forms of NEMO upon denaturing INT-Ub.7KR Strep-Tactin PD. Blot images for Supplementary Figure 2a,c,d were cropped to improve the conciseness of the presentation (full blots are available in Supplementary Figure 12).
Supplementary Figure 3 Mass-spectrometry-based identification of novel linear polyUb-modified substrates.
(A) A schematic representation of the preliminary SILAC-based MS proteomic approach for the identification of linear polyUb-modified substrates upon 15 min of TNFα stimulation. Note that the SILAC labels were reversed in the subsequent triplicate MS analysis. (B) Scatter plots shown as a function of protein intensity and SILAC ratios (left: M/L, right: H/L) for preliminary MS data in (A) and Supplementary Table 1. SILAC ratios of log2>1 are marked in red. Several putative LUBAC substrates selected for the final validation are indicated in the plots. The number of identified protein pairs is marked in left upper part of each plot. (C) Experimental reproducibility of three independent experimental replicates of MS proteomic experiments from Figure 4a. The Pearson Correlation Coefficients are depicted in the left upper part of each plot. (D) Expression levels of putative linear polyUb-modified proteins in two different sets of T-REx HEK293T cell lines used in MS screens. Blot images for Supplementary Figure 3d were cropped to improve the conciseness of the presentation (full blots are available in Supplementary Figure 12).
Supplementary Figure 4 Validation of novel linear polyUb-modified substrates.
(A) Effect of active LUBAC overexpression on the appearance of HMW species above expected protein size for transiently transfected HA-tagged selected MS hits. Inactive LUBAC (HOIP C885A/HOIL-1L) was used as negative control. (B) Interaction studies between selected MS candidates and LUBAC components HOIP and HOIL-1. HOIP and/or HOIL-1L were transiently overexpressed in HEK293T cells and used for MBP or GST pull-down assays with recombinant putative substrates. (C) In vitro ubiquitination assays with recombinant LUBAC components and recombinant HIS-tagged putative substrates. (D) In vitro ubiquitination sample containing recombinant LUBAC complex and TRAF6 was treated with recombinant OTULIN to confirm TRAF6 modification by linear polyUb chains. (E) MS/MS spectra showing the presence of linear Ub signature peptide GGMQIFVK in IP sample, in which transiently overexpressed HA-TRAF6 and HOIP/HOIL-1L were immunoprecipitated with HA agarose under denaturing conditions. (F) Analysis of the presence of HMW species of endogenous SEPT2 (panel 1), HDAC6 (panel 2), VDAC1 (panel 3) and TRAF6 (panel 4) in the presence of ectopic LUBAC complex in HEK293T cells by denaturing linear Ub IP. Blot images for Supplementary Figure 4 (except e) were cropped to improve the conciseness of the presentation (full blots are available in Supplementary Figures 13, 14).
Supplementary Figure 5 TRAF6 is a novel LUBAC substrate in NFκB signalling.
(A) The effect of HOIP silencing on LUBAC-dependent linear polyubiquitination of TRAF6 upon IL-1β-stimulation. (B) MS/MS spectra showing identified ubiquitinated residues of TRAF6. HA-TRAF6 and LUBAC components were transiently overexpressed in HEK293T cells and used for GST pull-down with NEMO UBANx3 and further processed for MS analysis. (C) In vitro ubiquitination assays with recombinant LUBAC components and either HIS-tagged TRAF6 C70A or TRAF6 C70A K339/K497/K518R mutant. (D) Levels of recombinant HIS-tagged TRAF6 C70A and C70A K339/K497/ K518R mutants. The difference in protein size observed on gradient gels (Supplementary Figure 5c) is not visible on non-gradient 10 % SDS-PAGE. (E) IκBα phosphorylation and degradation kinetics in reconstituted TRAF6-/- MEFs. Cells were stimulated with IL-1β (10 ng/ml) for the indicated time periods. Quantification of the protein levels was performed by ImageJ software. Mean of three independent experimental replicas was calculated and is shown below figure panels (compared to time point 0 for each cell line). (F) Expression levels of various HA-TRAF6 variants stably reconstituted in TRAF6-/- MEFs. Blot images for Supplementary Figure 5 (except b) were cropped to improve the conciseness of the presentation (full blots are available in Supplementary Figure 15).
Supplementary Figure 6 Full blot images for Figures 1 and 2.
Full blot names indicate their position in the original Figures 1 and 2. Unless specifically indicated next to the blot, all the blots had the same protein marker (with 2 bands marked with +), as indicated by the schematic protein marker representation. Red frames indicate cropped parts used in the original figures.
Supplementary Figure 7 Full blot images for Figure 3.
Full blot names indicate their position in the original Figure 3. Unless specifically indicated next to the blot, all the blots had the same protein marker (with 2 bands marked with +), as indicated by the schematic protein marker representation. Red frames indicate cropped parts used in the original figures.
Supplementary Figure 8 Full blot images for Figure 5a.
Full blot names indicate their position in the original Figure 5a. Unless specifically indicated next to the blot, all the blots had the same protein marker (with 2 bands marked with +), as indicated by the schematic protein marker representation. Red frames indicate cropped parts used in the original figures.
Supplementary Figure 9 Full blot images for Figure 5b.
Full blot names indicate their position in the original Figure 5b. Unless specifically indicated next to the blot, all the blots had the same protein marker (with 2 bands marked with +), as indicated by the schematic protein marker representation. Red frames indicate cropped parts used in the original figures.
Supplementary Figure 10 Full blot images for Figure 5c.
Full blot names indicate their position in the original Figure 5c. Unless specifically indicated next to the blot, all the blots had the same protein marker (with 2 bands marked with +), as indicated by the schematic protein marker representation. Red frames indicate cropped parts used in the original figures.
Supplementary Figure 11 Full blot images for Figure 6.
Full blot names indicate their position in the original Figure 6. Unless specifically indicated next to the blot, all the blots had the same protein marker (with 2 bands marked with +), as indicated by the schematic protein marker representation. Red frames indicate cropped parts used in the original figures.
Supplementary Figure 12 Full blot images for Supplementary Figures 1, 2, 3.
Full blot names indicate their position in the original Supplementary Figures 1, 2, 3. Unless specifically indicated next to the blot, all the blots had the same protein marker (with 2 bands marked with +), as indicated by the schematic protein marker representation. Red frames indicate cropped parts used in the original figures.
Supplementary Figure 13 Full blot images for Supplementary Figures 4a–d.
Full blot names indicate their position in the original Supplementary Figures 4a-d. Unless specifically indicated next to the blot, all the blots had the same protein marker (with 2 bands marked with +), as indicated by the schematic protein marker representation. Red frames indicate cropped parts used in the original figures.
Supplementary Figure 14 Full blot images for Supplementary Figure 4f.
Full blot names indicate their position in the original Supplementary Figure 4f. Unless specifically indicated next to the blot, all the blots had the same protein marker (with 2 bands marked with +), as indicated by the schematic protein marker representation. Red frames indicate cropped parts used in the original figures.
Supplementary Figure 15 Full blot images for Supplementary Figure 5.
Full blot names indicate their position in the original Supplementary Figure 5. Unless specifically indicated next to the blot, all the blots had the same protein marker (with 2 bands marked with +), as indicated by the schematic protein marker representation. Red frames indicate cropped parts used in the original figures.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15 and Supplementary Tables 3–5 (PDF 2356 kb)
Supplementary Table 1
Results of the mass spectrometry experiments. (XLSX 1454 kb)
Supplementary Table 2
Results of the triplicate SILAC-based mass spectrometryexperiment with INT-Ub variants (10 min of TNFα stimulation). (XLSX 4985 kb)
Rights and permissions
About this article
Cite this article
Kliza, K., Taumer, C., Pinzuti, I. et al. Internally tagged ubiquitin: a tool to identify linear polyubiquitin-modified proteins by mass spectrometry. Nat Methods 14, 504–512 (2017). https://doi.org/10.1038/nmeth.4228
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4228
This article is cited by
-
Profiling the interactome of oligonucleotide drugs by proximity biotinylation
Nature Chemical Biology (2024)
-
N4BP1 functions as a dimerization-dependent linear ubiquitin reader which regulates TNF signalling
Cell Death Discovery (2024)
-
The ABL-MYC axis controls WIPI1-enhanced autophagy in lifespan extension
Communications Biology (2023)
-
Histone 4 lysine 5/12 acetylation enables developmental plasticity of Pristionchus mouth form
Nature Communications (2023)
-
Linear ubiquitination of PTEN impairs its function to promote prostate cancer progression
Oncogene (2022)