Abstract
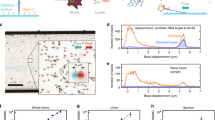
We describe a scheme for biomolecule enumeration by converting nanometer-scale specific molecular recognition events mediated by rolling-circle amplification to fluorescent micrometer-sized DNA molecules amenable to discrete optical detection. Our amplified single-molecule detection (SMD) approach preserves the discrete nature of the molecular population, allowing multiplex detection and highly precise quantification of molecules over a dynamic range of seven orders of magnitude. We apply the method for sensitive detection and quantification of the bacterial pathogen Vibrio cholerae.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Weiss, S. Fluorescence spectroscopy of single biomolecules. Science 283, 1676–1683 (1999).
Vogelstein, B. & Kinzler, K.W. Digital PCR. Proc. Natl. Acad. Sci. USA 96, 9236–9241 (1999).
Dressman, D., Yan, H., Traverso, G., Kinzler, K.W. & Vogelstein, B. Transforming single DNA molecules into fluorescent magnetic particles for detection and enumeration of genetic variations. Proc. Natl. Acad. Sci. USA 100, 8817–8822 (2003).
Mitra, R.D. et al. Digital genotyping and haplotyping with polymerase colonies. Proc. Natl. Acad. Sci. USA 100, 5926–5931 (2003).
Fire, A. & Xu, S.-Q. Rolling replication of short DNA circles. Proc. Natl. Acad. Sci. USA 92, 4641–4645 (1995).
Lizardi, P.M. et al. Mutation detection and single-molecule counting using isothermal rolling-circle amplification. Nat. Genet. 19, 225–232 (1998).
Nie, B., Shortreed, M.R. & Smith, L.M. Scoring single-nucleotide polymorphisms at the single-molecule level by counting individual DNA cleavage events on surfaces. Anal. Chem. 77, 6594–6600 (2005).
Larsson, C. et al. In situ genotyping individual DNA molecules by target-primed rolling-circle amplification of padlock probes. Nat. Methods 1, 227–232 (2004).
Nilsson, M. et al. Padlock probes: circularizing oligonucleotides for localized DNA detection. Science 265, 2085–2088 (1994).
Fredriksson, S. et al. Protein detection using proximity-dependent DNA ligation assays. Nat. Biotechnol. 20, 473–477 (2002).
Blab, G.A., Schmidt, T. & Nilsson, M. Sensitive and homogenous detection of single rolling-circle replication products. Anal. Chem. 76, 495–498 (2004).
Dahl, F. et al. Circle-to-circle amplification for precise and sensitive DNA analysis. Proc. Natl. Acad. Sci. USA 101, 4548–4553 (2004).
Reidl, J. & Klose, K.E. Vibrio cholerae and cholera: out of the water and into the host. FEMS Microbiol. Rev. 26, 125–139 (2002).
Rutledge, R.G. & Cote, C. Mathematics of quantitative kinetic PCR and the application of standard curves. Nucleic Acids Res. 31, e93 (2003).
Melin, J. et al. Thermoplastic microfluidic platform for single-molecule detection, cell culture, and actuation. Anal. Chem. 77, 7122–7130 (2005).
Acknowledgements
We thank O. Öhman at Åmic AB, L. Spångberg and S.E. Alm. This work was supported by the Wallenberg Foundation, the Swedish Defense Nanotechnology Program, Uppsala BioX, the EU Framework Programme 6 integrated project MolTools, the Swedish Research Council.
Author information
Authors and Affiliations
Contributions
J.J., S.F. and M.N. conceived of the described method. J.J., J.M., J.G. and M.N. contributed to the technical development of the method. J.J., J.M. and J.G. performed laboratory work. C.G.-R. and S.B. contributed with the bacterial application. J.S. designed microbial specific padlock probes. J.M., J.J., J.G., S.B. and M.N. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Extended dynamic range using circle to circle amplification (C2CA). (PDF 104 kb)
Supplementary Fig. 2
Ligation and RCA in complex matrices. (PDF 47 kb)
Supplementary Fig. 3
Multiplex detection. (PDF 827 kb)
Supplementary Fig. 4
Protein detection using proximity ligation. (PDF 41 kb)
Supplementary Table 1
Oligonucleotide sequences. (PDF 35 kb)
Rights and permissions
About this article
Cite this article
Jarvius, J., Melin, J., Göransson, J. et al. Digital quantification using amplified single-molecule detection. Nat Methods 3, 725–727 (2006). https://doi.org/10.1038/nmeth916
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth916
This article is cited by
-
A novel mutation tolerant padlock probe design for multiplexed detection of hypervariable RNA viruses
Scientific Reports (2019)
-
A microfluidic platform towards automated multiplexed in situ sequencing
Scientific Reports (2019)
-
Integration of microbead DNA handling with optomagnetic detection in rolling circle amplification assays
Microchimica Acta (2019)
-
Label-free detection of real-time DNA amplification using a nanofluidic diffraction grating
Scientific Reports (2016)
-
Compaction of rolling circle amplification products increases signal integrity and signal-to-noise ratio
Scientific Reports (2015)