Abstract
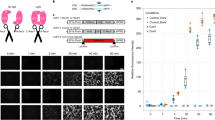
To facilitate a functional analysis of neuronal connectivity in a mammalian nervous system that is tightly packed with billions of cells, we developed a new technique that uses inducible genetic manipulations in fluorescently labeled single neurons in mice. Our technique, single-neuron labeling with inducible Cre-mediated knockout (SLICK), is achieved by coexpressing a drug-inducible form of Cre recombinase and a fluorescent protein in a small subsets of neurons, thus combining the powerful Cre recombinase system for conditional genetic manipulation with fluorescent labeling of single neurons for imaging. Here, we demonstrate efficient inducible genetic manipulation in several types of neurons using SLICK. Furthermore, we applied SLICK to eliminate synaptic transmission in a small subset of neuromuscular junctions. Our results provide evidence for the long-term stability of inactive neuromuscular synapses in adult animals and demonstrate a Cre-loxP compatible system for dissecting gene functions in single identifiable neurons.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
International Mouse Knockout Consortium. Collins, F.S., Rossant, J. & Wurst, W. A mouse for all reasons. Cell 128, 9–13 (2007).
Brecht, M. et al. Novel approaches to monitor and manipulate single neurons in vivo. J. Neurosci. 24, 9223–9227 (2004).
Callaway, E.M. A molecular and genetic arsenal for systems neuroscience. Trends Neurosci. 28, 196–201 (2005).
Deisseroth, K. et al. Next-generation optical technologies for illuminating genetically targeted brain circuits. J. Neurosci. 26, 10380–10386 (2006).
Marek, K.W. & Davis, G.W. Controlling the active properties of excitable cells. Curr. Opin. Neurobiol. 13, 607–611 (2003).
Miesenbock, G. Genetic methods for illuminating the function of neural circuits. Curr. Opin. Neurobiol. 14, 395–402 (2004).
Miyawaki, A. Fluorescence imaging of physiological activity in complex systems using GFP-based probes. Curr. Opin. Neurobiol. 13, 591–596 (2003).
Luo, L., Callaway, E.M. & Svoboda, K. Genetic dissection of neural circuits. Neuron 57, 634–660 (2008).
Young, P. & Feng, G. Labeling neurons in vivo for morphological and functional studies. Curr. Opin. Neurobiol. 14, 642–646 (2004).
Zong, H., Espinosa, J.S., Su, H.H., Muzumdar, M.D. & Luo, L. Mosaic analysis with double markers in mice. Cell 121, 479–492 (2005).
Feng, G. et al. Imaging neuronal subsets in transgenic mice expressing multiple spectral variants of GFP. Neuron 28, 41–51 (2000).
Grutzendler, J., Kasthuri, N. & Gan, W.B. Long-term dendritic spine stability in the adult cortex. Nature 420, 812–816 (2002).
Kasthuri, N. & Lichtman, J.W. Structural dynamics of synapses in living animals. Curr. Opin. Neurobiol. 14, 105–111 (2004).
Trachtenberg, J.T. et al. Long-term in vivo imaging of experience-dependent synaptic plasticity in adult cortex. Nature 420, 788–794 (2002).
Mizrahi, A. & Katz, L.C. Dendritic stability in the adult olfactory bulb. Nat. Neurosci. 6, 1201–1207 (2003).
Feil, R., Wagner, J., Metzger, D. & Chambon, P. Regulation of Cre recombinase activity by mutated estrogen receptor ligand-binding domains. Biochem. Biophys. Res. Commun. 237, 752–757 (1997).
Branda, C.S. & Dymecki, S.M. Talking about a revolution: the impact of site-specific recombinases on genetic analyses in mice. Dev. Cell 6, 7–28 (2004).
Metzger, D. & Chambon, P. Site- and time-specific gene targeting in the mouse. Methods 24, 71–80 (2001).
Soriano, P. Generalized lacZ expression with the ROSA26 Cre reporter strain. Nat. Genet. 21, 70–71 (1999).
Burrone, J. & Murthy, V.N. Synaptic gain control and homeostasis. Curr. Opin. Neurobiol. 13, 560–567 (2003).
Hua, J.Y. & Smith, S.J. Neural activity and the dynamics of central nervous system development. Nat. Neurosci. 7, 327–332 (2004).
Misgeld, T. et al. Roles of neurotransmitter in synapse formation: development of neuromuscular junctions lacking choline acetyltransferase. Neuron 36, 635–648 (2002).
Brandon, E.P. et al. Aberrant patterning of neuromuscular synapses in choline acetyltransferase–deficient mice. J. Neurosci. 23, 539–549 (2003).
Chen, C. & Tonegawa, S. Molecular genetic analysis of synaptic plasticity, activity-dependent neural development, learning, and memory in the mammalian brain. Annu. Rev. Neurosci. 20, 157–184 (1997).
Lee, T. & Luo, L. Mosaic analysis with a repressible cell marker for studies of gene function in neuronal morphogenesis. Neuron 22, 451–461 (1999).
Wang, X. et al. Prolongation of evoked and spontaneous synaptic currents at the neuromuscular junction after activity blockade is caused by the upregulation of fetal acetylcholine receptors. J. Neurosci. 26, 8983–8987 (2006).
Wang, X. et al. Activity-dependent presynaptic regulation of quantal size at the mammalian neuromuscular junction in vivo. J. Neurosci. 25, 343–351 (2005).
Son, Y.J. & Thompson, W.J. Nerve sprouting in muscle is induced and guided by processes extended by Schwann cells. Neuron 14, 133–141 (1995).
Johns, D.C., Marx, R., Mains, R.E., O'Rourke, B. & Marban, E. Inducible genetic suppression of neuronal excitability. J. Neurosci. 19, 1691–1697 (1999).
Sweeney, S.T., Broadie, K., Keane, J., Niemann, H. & O'Kane, C.J. Targeted expression of tetanus toxin light chain in Drosophila specifically eliminates synaptic transmission and causes behavioral defects. Neuron 14, 341–351 (1995).
Arenkiel, B.R. et al. In vivo light-induced activation of neural circuitry in transgenic mice expressing channelrhodopsin-2. Neuron 54, 205–218 (2007).
Wang, H. et al. High-speed mapping of synaptic connectivity using photostimulation in Channelrhodopsin-2 transgenic mice. Proc. Natl. Acad. Sci. USA 104, 8143–8148 (2007).
Shaner, N.C. et al. Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. Nat. Biotechnol. 22, 1567–1572 (2004).
Lewandoski, M. Conditional control of gene expression in the mouse. Nat. Rev. Genet. 2, 743–755 (2001).
Sauer, B. Inducible gene targeting in mice using the Cre/lox system. Methods 14, 381–392 (1998).
Caroni, P. Overexpression of growth-associated proteins in the neurons of adult transgenic mice. J. Neurosci. Methods 71, 3–9 (1997).
Hogan, B., Constantini, F. & Lacey, E. Production of transgenic mice. in Manipulating the Mouse Embryo: A Laboratory Manual (eds Hogan, B., Constantini, F. & Lacey, E.) 217–252 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 1994).
Acknowledgements
We thank J. Sanes for providing the Chat conditional-knockout mice and the antibody to β-galactosidase. We thank A. Mizrahi for performing two-photon imaging of SLICK-3 mice and R. Kotloski for advice regarding tamoxifen administration. We also thank the Duke Neurotransgenic Core Facility for generating the SLICK mice. We thank J. McNamara, M.D. Ehlers and M. McCaffrey for access to confocal microscopes. We are grateful to B. Arenkiel, M.D. Ehlers, A. West, K. Dean and members of the Feng laboratory for critically reading and discussing the manuscript. This work was supported by the Ruth K. Broad Biomedical Research Foundation, the Science Foundation Ireland and the US National Institutes of Health.
Author information
Authors and Affiliations
Contributions
P.Y. and G.F. conceived the SLICK method, designed the experiments and wrote the manuscript. P.Y. and J.G. generated the SLICK transgenic mice. P.Y., L.Q., S.Z. and D.W. characterized the SLICK mice. P.Y. and S.Z. carried out the quantification and data analysis.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Methods and Discussion (PDF 6860 kb)
Rights and permissions
About this article
Cite this article
Young, P., Qiu, L., Wang, D. et al. Single-neuron labeling with inducible Cre-mediated knockout in transgenic mice. Nat Neurosci 11, 721–728 (2008). https://doi.org/10.1038/nn.2118
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.2118
This article is cited by
-
Circadian clock regulator Bmal1 gates axon regeneration via Tet3 epigenetics in mouse sensory neurons
Nature Communications (2023)
-
FAM19A5/TAFA5, a novel neurokine, plays a crucial role in depressive-like and spatial memory-related behaviors in mice
Molecular Psychiatry (2021)
-
The effect of NAMPT deletion in projection neurons on the function and structure of neuromuscular junction (NMJ) in mice
Scientific Reports (2020)
-
Brainstem development requires galactosylceramidase and is critical for pathogenesis in a model of Krabbe disease
Nature Communications (2020)
-
Simultaneous fluorescence imaging of distinct nerve and blood vessel patterns in dual Thy1-YFP and Flt1-DsRed transgenic mice
Angiogenesis (2020)