Abstract
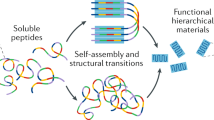
Sequence-specific polymers, such as oligonucleotides and peptides, can be used as building blocks for functional supramolecular nanomaterials. The design and selection of suitable self-assembling sequences is, however, challenging because of the vast combinatorial space available. Here we report a methodology that allows the peptide sequence space to be searched for self-assembling structures. In this approach, unprotected homo- and heterodipeptides (including aromatic, aliphatic, polar and charged amino acids) are subjected to continuous enzymatic condensation, hydrolysis and sequence exchange to create a dynamic combinatorial peptide library. The free-energy change associated with the assembly process itself gives rise to selective amplification of self-assembling candidates. By changing the environmental conditions during the selection process, different sequences and consequent nanoscale morphologies are selected.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Jones, M. R., Seeman, N. C. & Mirkin, C. A. Programmable materials and the nature of the DNA bond. Science 347, 1260901 (2015).
Webber, M. J., Appel, E. A., Meijer, E. & Langer, R. Supramolecular biomaterials. Nat. Mater. 15, 13–26 (2016).
Aida, T., Meijer, E. & Stupp, S. Functional supramolecular polymers. Science 335, 813–817 (2012).
Du, X., Zhou, J., Shi, J. & Xu, B. Supramolecular hydrogelators and hydrogels: from soft matter to molecular biomaterials. Chem. Rev. 15, 13165–13307 (2015).
Whitesides, G. M. & Grzybowski, B. Self-assembly at all scales. Science 295, 2418–2421 (2002).
Hartgerink, J. D., Beniash, E. & Stupp, S. L. Self-assembly and mineralization of peptide-amphiphile nanofibers. Science 294, 1684–1688 (2001).
Thomson, A. R. et al. Computational design of water-soluble α-helical barrels. Science 346, 485–488 (2014).
Omenetto, F. G. & Kaplan, D. L. New opportunities for an ancient material. Science 329, 528–531 (2010).
Inostroza-Brito, K. E. et al. Co-assembly, spatiotemporal control and morphogenesis of a hybrid protein–peptide system. Nat. Chem. 7, 897–904 (2015).
Zhang, S. Fabrication of novel biomaterials through molecular self-assembly. Nat. Biotechnol. 21, 1171–1178 (2003).
Reches, M. & Gazit, E. Casting metal nanowires within discrete self-assembled peptide nanotubes. Science 300, 625–627 (2003).
Görbitz, C. H. Nanotube formation by hydrophobic dipeptides. Chem. Eur. J. 7, 5153–5159 (2001).
Frederix, P. W. J. M. et al. Exploring the sequence space for (tri-)peptide self-assembly to design and discover new hydrogels. Nat. Chem. 7, 30–37 (2015).
Erdogan, H. et al. Morphological versatility in the self-assembly of Val-Ala and Ala-Val dipeptides. Langmuir 31, 7337–7345 (2015).
Adler-Abramovich, L. & Gazit, E. The physical properties of supramolecular peptide assemblies: from building block association to technological applications. Chem. Soc. Rev. 43, 6881–6893 (2014).
Fan, Z., Sun, L., Huang, Y., Wang, Y. & Zhang, M. Bioinspired fluorescent dipeptide nanoparticles for targeted cancer cell imaging and real-time monitoring of drug release. Nat. Nanotech. 11, 388–394 (2016).
Berger, O. et al. Light-emitting self-assembled peptide nucleic acids exhibit both stacking interactions and Watson–Crick base pairing. Nat. Nanotech. 10, 353–360 (2015).
Sanchez-de Groot, N., Parella, T., Aviles, F., Vendrell, J. & Ventura, S. Ile-Phe dipeptide self-assembly: clues to amyloid formation. Biophys. J. 92, 1732–1741 (2007).
Forrer, P., Jung, S. & Plückthun, A. Beyond binding: using phage display to select for structure, folding and enzymatic activity in proteins. Curr. Opin. Struct. Biol. 9, 514–520 (1999).
Whaley, S. R., English, D., Hu, E. L., Barbara, P. F. & Belcher, A. M. Selection of peptides with semiconductor binding specificity for directed nanocrystal assembly. Nature 405, 665–668 (2000).
Naik, R. R., Stringer, S. J., Agarwal, G., Jones, S. E. & Stone, M. O. Biomimetic synthesis and patterning of silver nanoparticles. Nat. Mater. 1, 169–172 (2002).
Ellington, A. D. & Szostak, J. W. In vitro selection of RNA molecules that bind specific ligands. Nature 346, 818–822 (1990).
Tuerk, C. & Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 249, 505–510 (1990).
Bastian, A. A., Marcozzi, A. & Herrmann, A. Selective transformations of complex molecules are enabled by aptameric protective groups. Nat. Chem. 4, 789–793 (2012).
Zhang, S., Holmes, T., Lockshin, C. & Rich, A. Spontaneous assembly of a self-complementary oligopeptide to form a stable macroscopic membrane. Proc. Natl Acad. Sci. USA 90, 3334–3338 (1993).
Ghadiri, M. R., Granja, J. R., Milligan, R. A., McRee, D. E. & Khazanovich, N. Self-assembling organic nanotubes based on a cyclic peptide architecture. Nature 366, 324–327 (1993).
Lehn, J. M. Dynamic combinatorial chemistry and virtual combinatorial libraries. Chem. Eur. J. 5, 2455–2463 (1999).
Corbett, P. T. et al. Dynamic combinatorial chemistry. Chem. Rev. 106, 3652–3711 (2006).
Moulin, E., Cormos, G. & Giuseppone, N. Dynamic combinatorial chemistry as a tool for the design of functional materials and devices. Chem. Soc. Rev. 41, 1031–1049 (2012).
Carnall, J. M. et al. Mechanosensitive self-replication driven by self-organization. Science 327, 1502–1506 (2010).
Krishnan Ghosh, Y. & Balasubramanian, S. Dynamic covalent chemistry on self templating peptides: formation of a disulfide linked β hairpin mimic. Angew. Chem. 115, 2221–2223 (2003).
Campbell, V. E. et al. Cascading transformations within a dynamic self-assembled system. Nat. Chem. 2, 684–687 (2010).
Swann, P. G. et al. Nonspecific protease-catalyzed hydrolysis/synthesis of a mixture of peptides: product diversity and ligand amplification by a molecular trap. Peptide Sci. 40, 617–625 (1996).
Williams, R. J. et al. Enzyme-assisted self-assembly under thermodynamic control. Nat. Nanotech. 4, 19–24 (2009).
Guilbaud, J. B. et al. Enzymatic catalyzed synthesis and triggered gelation of ionic peptides. Langmuir 26, 11297–11303 (2010).
Qin, X. et al. Enzyme-triggered hydrogelation via self-assembly of alternating peptides. Chem. Commun. 49, 4839–4841 (2013).
Ruff, Y., Garavini, V. & Giuseppone, N. Reversible native chemical ligation: a facile access to dynamic covalent peptides. J. Am. Chem. Soc. 136, 6333–6339 (2014).
Rodriguez-Garcia, M. et al. Formation of oligopeptides in high yield under simple programmable conditions. Nat. Commun. 6, 8385–8391 (2015).
Hofmeister, F. On the understanding of the effects of salts. Arch. Exp. Pathol. Pharmakol. (Leipzig) 24, 247–260 (1888).
Roy, S. et al. Dramatic specific-ion effect in supramolecular hydrogels. Chem. Eur. J. 18, 11723–11731 (2012).
Ulijn, R. V. et al. Solvent selection for solid-to-solid synthesis. Biotechnol. Bioeng. 80, 509–515 (2002).
Leonetti, G. & Otto, S. Solvent composition dictates emergence in dynamic molecular networks containing competing replicators. J. Am. Chem. Soc. 137, 2067–2072 (2015).
Gutierrez, J. M. P., Hinkley, T., Taylor, J. W., Yanev, K. & Cronin, L. Evolution of oil droplets in a chemorobotic platform. Nat. Commun. 5, 5571–5578 (2014).
Acknowledgements
The authors acknowledge P. Keating, S. Chakravartula and B. Yoo for the liquid chromatography–mass spectroscopy (LC–MS) experiments. We thank M. Mullin for the help with TEM imaging, N. T. Hunt for access to FTIR spectroscopy and S. Kelly for use of the CD equipment. The authors acknowledge the ‘NanoFabrication Facility’ from the CUNY ASRC. The authors also acknowledge R. Mart for the help with the design of the free-energy diagram (Fig. 1). The research leading to these results has received funding from the European Research Council under the European Union's Seventh Framework Programme (FP7/2007–2013) EMERgE/ERC (Grant Agreement Number 258775) and US Air Force (AFOSR, grant FA9550-15-1-0192). C.G.P. acknowledges Linn Products Ltd for funding and I.R.S. acknowledges the EC 7th Framework Programme Marie Curie Actions via the European ITN SMARTNET No. 316656.
Author information
Authors and Affiliations
Contributions
C.G.P., R.S., I.R.S and R.V.U. conceived and designed the experiments. C.G.P., R.S., I.R.S. and H.S. performed the experiments. C.G.P., R.S., I.R.S. and R.V.U. analysed the data. T.W. performed the TEM and cryo-TEM experiments. N.W. performed the rheological experiments. C.G.P. and V.N. performed the AFM experiments. R.A. performed the LC–MS experiments. C.G.P. and R.V.U. co-wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 3034 kb)
Rights and permissions
About this article
Cite this article
Pappas, C., Shafi, R., Sasselli, I. et al. Dynamic peptide libraries for the discovery of supramolecular nanomaterials. Nature Nanotech 11, 960–967 (2016). https://doi.org/10.1038/nnano.2016.169
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2016.169
This article is cited by
-
Self-assembled and Zn(II)-coordinated dipeptide nanoparticles with membrane-rupturing action on bacteria
Applied Microbiology and Biotechnology (2023)
-
Rapid discovery of self-assembling peptides with one-bead one-compound peptide library
Nature Communications (2021)
-
Emerging functional materials based on chemically designed molecular recognition
BMC Materials (2020)
-
Homopolymer self-assembly of poly(propylene sulfone) hydrogels via dynamic noncovalent sulfone–sulfone bonding
Nature Communications (2020)
-
A chemically fueled non-enzymatic bistable network
Nature Communications (2019)