Abstract
Circadian rhythm and sleep disruptions occur frequently in individuals with alcohol use disorders (AUD) and present significant barriers to treatment. Recently, a variant of adenosine transporter, equilibrative nucleoside transporter 1 (ENT1), was associated with the co-occurrence of sleep problems and AUD. We have previously shown that mice lacking ENT1 (ENT1 KO) have reduced adenosine levels in the striatum and drink more alcohol compared with wild types (WT). However, it is unknown whether ENT1 deletion disrupts circadian rhythms, which may contribute to alcohol preference in ENT1 KO mice. Here we used these mice to determine whether endogenous adenosine regulates circadian genetic and behavioral rhythms and influences alcohol intake during chronodisruption. We examined circadian locomotor activity in ENT1 KO vs WT littermates and found that ENT1 KO mice were both active earlier and hyperactive compared with WT mice at night. We used real-time PCR and immunohistochemistry to estimate striatal clock gene levels and found that PER2 expression in the striatum was blunted by ENT1 deletion or A2A receptor (A2AR) antagonism. Next, we exposed ENT1 KO and WT mice to constant light (LL) and found further elevation in ethanol intake in ENT1 KO, but not in WT mice, supporting the notion that circadian dysfunction may contribute to increased alcohol intake in ENT1 KO mice. Finally, we showed that A2AR agonist administration normalized PER1 and PER2 expression and circadian locomotor activity in ENT1 KO mice. Together, our results demonstrate that adenosine signaling regulates cellular and behavioral circadian timing and influences alcohol intake during chronodisruption.
Similar content being viewed by others
INTRODUCTION
Circadian rhythm and sleep problems are common, treatment-resistant components of alcohol use disorders (AUD) that greatly contribute to relapse (Brower, 2003; Brower and Perron, 2010). Alcohol is highly disruptive to circadian hormonal and activity rhythms, whereas conversely, circadian desynchrony arising from sleep disorders (Roehrs and Roth, 2001), shiftwork, or repeated jetlag (Gordon et al, 1986; Smart, 1979) increases risk of developing AUD. Thus, misalignment of circadian time may be central to the pathophysiology of AUD (Rosenwasser, 2010).
At the cellular level, circadian timing is determined by oscillatory loops of transcription and translation of clock genes (Lowrey and Takahashi, 2004). A central role for the circadian system in AUD is underscored by evidence linking most clock genes with AUD (Albrecht et al, 2001; Franken et al, 2000; Gamsby et al, 2013; Kovanen et al, 2010; Le-Niculescu et al, 2008; Lopez-Molina et al, 1997; Melendez et al, 2011; Perreau-Lenz et al, 2012; Spanagel et al, 2005; Steinlechner et al, 2002; Wang et al, 2012; Yang et al, 2012). In addition to ethanol (Melendez et al, 2011), morphine (Hood et al, 2011) and amphetamine (Wongchitrat et al, 2013) alter circadian variation in clock gene expression in the mesolimbic reward system during chronic use and/or withdrawal, which may contribute to addictive behavior and/or circadian misalignment. However, little is known about clock gene regulation outside of the suprachiasmatic nucleus (SCN) circadian clock.
Adenosine signaling has an essential role in acute ethanol intoxication, including ataxia, anxiolysis, and sedation, while long-term ethanol exposure leads to neuroadaptation in adenosine signaling that contributes largely to alcohol dependence (Asatryan et al, 2011; Dunwiddie and Masino, 2001; Ruby et al, 2010). Adenosine transporter equilibrative nucleoside transporter 1 (ENT1) is blocked by acute ethanol and downregulated by chronic ethanol exposure (Nagy et al, 1990; Nagy et al, 1989), and studies associate ENT1 with AUD in humans (Gass et al, 2010; Kim et al, 2011). Mice lacking ENT1 (ENT1 KO) display high tolerance to ethanol intoxication and drink alcohol excessively (Choi et al, 2004) because of low adenosine tone in the striatum (Nam et al, 2011; Wu et al, 2010). Recent evidence links a variant of ENT1 with circadian sleep disruption and AUD (Gass et al, 2010).
As a regulator of circadian timing (Antle et al, 2001; Hallworth et al, 2002; Sigworth and Rea, 2003), sleep (Halassa et al, 2009), and alcohol intake (Asatryan et al, 2011; Dunwiddie and Masino, 2001; Ruby et al, 2010), adenosine signaling may contribute to chronodisruption in AUD. Adenosine modulates both photic (light) and nonphotic (behavioral) circadian clock resetting (Antle et al, 2001; Hallworth et al, 2002; Sigworth and Rea, 2003). Caffeine, an adenosine A1 receptor (A1R)/A2A receptor (A2AR) antagonist, lengthens (Oike et al, 2011), whereas ethanol shortens (Seggio et al, 2009), free-running (endogenous) circadian period. Circadian variation in astrocytic ATP release, extracellular adenosine concentration, and ENT1 and A1R expression has been reported in sleep/wake-promoting areas of the brain (Alanko et al, 2003; Marpegan et al, 2011; Murillo-Rodriguez et al, 2004; Virus et al, 1984). Together, this evidence suggests that adenosine may regulate cellular circadian timekeeping.
To explore a role for endogenous adenosine in regulating circadian genetic and behavioral rhythms, we examined circadian entrainment (locomotor activity under a 12:12 light:dark, or LD, photocycle) and striatal clock gene expression in adenosine-challenged ENT1 KO mice (Nam et al, 2011) and wild-type (WT) littermates. We then measured ethanol intake in ENT1 KO and WT mice during constant light (LL), a circadian challenge. Finally, we investigated whether selective inhibition of A1R or A2AR in WT mice mimicked some of the clock gene expression changes we observed in ENT1 KO mice, and whether activation of A2AR improved circadian rhythms in ENT1 KO mice.
MATERIALS AND METHODS
Animals
ENT1 KO mice were generated (Choi et al, 2004). We used F2 hybrid ENT1 KO and WT littermates when they reached 10 weeks old, with a C57BL/6J × 129 × 1/SvJ background (Crusio et al, 2009) to avoid strain-specific influences on behavior or gene expression. Mice were housed in standard Plexiglas cages with food and water available ad libitum. The chamber was maintained on 12 h light (500 lux) and 12 h dark (<0.5 lux) cycle (LD) except during constant light (LL; see below). Animal care and handling procedures were approved by Mayo Clinic IACUC according to NIH guidelines.
Circadian Activity and Drinking
Mice under LD were individually caged and their circadian locomotor rhythms were monitored using infrared sensors interfaced with Clocklab (Coulbourn Instruments, Whitehall, PA). For 2 weeks, mice of each genotype were acclimated to lickometers (Coulbourn Instruments) to visualize circadian drinking patterns. After the acclimation period, mice received ethanol in two-bottle choice (see below); mice using lickometers received ethanol solutions in the lickometer for the duration of the experiment. After 2 weeks at 10% ethanol in LD, mice were exposed to LL for 2 weeks.
Entrainment to the LD photocycle was determined by calculating the daily activity onset relative to Zeitgeber Time 12 (ZT12), the beginning of the dark phase (active phase for nocturnal rodents). Onset was defined as the first 10 min activity that (1) exceeded 10% maximum daily rate; (2) was preceded by ⩾4 h inactivity; and (3) was followed by ⩾30 min sustained activity. Offset was defined as the final 10 min that was preceded by ⩾60 min sustained activity and followed by ⩾4 h inactivity. Alpha (the active phase) was calculated as the period between activity onset and activity offset (h). Activity distribution was assessed using activity bout analysis (Ruby et al, 2009). A bout was defined as a period of continuous activity (regardless of duration) separated by at least 10 min of inactivity. Activity duration (min of activity) and intensity (activity counts) were summed over the active phase (night) and over the inactive phase (day), and averaged over 8 days at each condition (basal, ethanol, and constant light) for each animal.
Quantitative, Real-Time PCR
Mice were anesthetized with CO2 and rapidly decapitated at ZT6, ZT10, ZT14, or ZT18. nucleus accumbens (NAc) was isolated under a surgical microscope. RNA was isolated using the RNAeasy-Plus Mini kit (Qiagen, Valencia, CA). Real-time, quantitative RT-PCR was performed with the iCycler IQ real-time PCR detection system (Bio-Rad, Hercules, CA) using QuantiTect SYBR Green RT-PCR Kit (Qiagen; (Wu et al, 2010). Gene-specific primers were purchased (Qiagen). Protocol: reverse transcription (30 min, 50 °C), denaturation (15 min, 95 °C), and 45 amplification and quantification cycles (15 s, 94 °C; 10 s, 55 °C; and 30 s, 72 °C), each with 1 fluorescence measurement. mRNA expression was normalized by GAPDH. Percentage changes were calculated by subtracting GAPDH Ct values from Ct values for the gene of interest (Livak and Schmittgen, 2001).
Immunohistochemistry
Mice were anesthetized with pentobarbital (80 mg/kg) at ZT22 (peak for PER2 protein in the striatum) and transcardially perfused with 4% paraformaldehyde (Sigma-Aldrich) in PBS. Brains were removed and post-fixed for 24 h in the same fixative at 4 °C. Brains were immersed in 30% sucrose for 24 h, frozen, and cut in 40 μm sections using a cryostat (Leica). Free-floating sections were incubated in 50% ethanol, then 10% normal donkey serum in PBS for 30 min, then antibodies against PER2 (1:250, Santa Cruz), and NeuN (1:100, Millipore) or GFAP (1:300, Cell Signaling) overnight. Sections were then incubated in 2% normal donkey serum in PBS for 10 min followed by Alexa 488-conjugated secondary goat anti-rabbit (for PER2, 1:1000, Cell Signaling) and Alexa 555-conjugated goat anti-mouse (for GFAP or NeuN, 1:1000, Cell Signaling) for 2 h. Images from each brain region of interest (NAc core, NAc shell, DMS, DLS, and SCN) were obtained using a LSM 510 confocal laser scanning microscope (Carl Zeiss). Areas of PER2 expression within regions of interest (450 × 450 μm) were quantified using NIH Image J software. Quantitative analysis was performed using a selected region-of-interest background correction method and calculated as a percentage of WT.
Pharmacological Treatment
For gene expression experiments, C57BL/6J mice were treated with either DPCPX (6.0 mg/kg, i.p.; Tocris), ZM-241385 (20.0 mg/kg, i.p.; Tocris), or vehicle (15% DMSO, 15% Cremaphor in 0.9% NaCl) for 3 days at ZT22 (peak for PER proteins in the NAc). ENT1 KO and WT mice were treated with CGS-21680 (2.0 mg/kg; Tocris) or vehicle for 3 days at ZT22. On day 4 at ZT14 (peak for Per mRNA in the NAc), mice were anesthetized with CO2, rapidly decapitated, and the NAc was isolated and frozen until real-time PCR was performed. For behavioral experiments, a separate group of ENT1 KO and WT mice was treated with CGS-21680 (1.0 mg/kg) or vehicle for 5 days at ZT22, and their behavior was analyzed using Clocklab. Drug doses and treatment length (3–5 days) were selected based on antidipsotropic efficacy (Nam et al, 2013); timing of doses and killing is shown in Figure 4a.
Ethanol Self-Administration
Oral alcohol self-administration and preference were examined using two-bottle choice (Lee et al, 2013). Mice had 24 h access to two bottles, one containing tap water and the other containing ethanol. The ethanol concentration was raised every fourth day from 3 to 6 to 10% (v/v) ethanol. After 2 weeks at 10% ethanol, mice were exposed to constant light (LL) for 2 weeks. A separate group was maintained on 10% ethanol for 4 weeks in LD to control for length of ethanol exposure. Fluid intake and body weight were measured every 48 h to calculate average ethanol consumption (g/kg/day) and preference (% ethanol solution/total fluid consumption).
Statistical Analyses
Student’s t-test or one-way ANOVA was used to compare circadian activity and gene expression within a single time point between genotypes or treatments. Two-way ANOVA was used to examine circadian gene expression between genotypes or behavioral response to drugs. Two-way RM ANOVA was used to compare ethanol intake between genotypes during LD vs LL. ANOVA were followed by Tukey post hoc tests where interactions were found. Results were significant where P<0.05.
RESULTS
Early Onset, Long Active-Phase and Nighttime Hyperactivity in ENT1 KO Mice
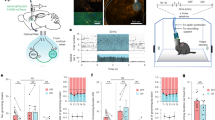
To explore the effect of endogenous adenosine concentration on circadian entrainment and activity level, we analyzed daily activity onset relative to ZT12 (beginning of the dark phase), alpha (active-phase length), and the duration and intensity of activity bouts during the night vs day in adenosine-challenged ENT1 KO mice vs WT littermates. Representative actograms from WT and ENT1 KO mice are shown in Figure 1a. In contrast to WT, ENT1 KO mice had a positive phase angle of entrainment in LD, becoming active ≈20 min before ZT12 (t=3.376, P=0.003; Figure 1b). Alpha was longer in ENT1 KO mice (t=3.381, P=0.003; Figure 1c). ENT1 KO mice showed greater 24 h activity duration (t=4.059, P<0.001) and intensity (t=2.419; P=0.025) owing to increased nighttime duration (t=4.643, P<0.001; Figure 1d) and intensity (t=2.519; P=0.020; Figure 1e), as daytime activity duration and intensity were similar between genotypes. These data indicate that ENT1 KO mice are hyperactive at night.
Adenosine transporter equilibrative nucleoside transporter 1 (ENT1) regulates circadian locomotor activity. (a) Representative actograms from wild-type (WT; left side) and ENT1 KO mice (right side). (b) ENT1 KO mice had a positive phase angle of entrainment relative to WT, becoming active before lights were turned off (Zeitgeber Time 12, ZT12). (c) ENT1 KO mice had longer alpha (active phase) relative to WT. (d) Increased nighttime, but not daytime, activity duration in ENT1 KO mice compared with WT. (e) Higher nighttime, but not daytime, activity intensity in ENT1 KO mice compared with WT. *P<0.05 by unpaired, two-tailed t-test; n=12 per genotype. Data are expressed as mean±SEM.
Altered Clock Gene Expression in the ENT1 KO Striatum
Clock gene expression in the striatum participates in the timing and intensity of daily locomotor activity (Iijima et al, 2002), yet little is known about striatal clock gene regulation. We found that circadian expression of Clock and Dbp was altered in ENT1 KO mice (Supplementary Figure S1), but the most striking difference was in Per2. Two-way ANOVA indicated a main effect of genotype (F1,38=13.617, P<0.001). A time-of-day effect indicated that Per2 expression was circadian (F3,38=14.230, P<0.001). A genotype × time-of-day interaction (F3,38=4.117, P=0.014) and subsequent post hoc testing revealed 60% reduction in peak Per2 mRNA levels in the ENT1 KO vs WT NAc at ZT14 (P<0.001; Figure 2a).
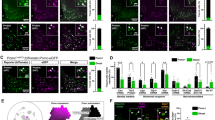
Equilibrative nucleoside transporter 1 (ENT1) regulates Per2 expression in the striatum. (a) ENT1 KO mice showed a reduction in Per2 mRNA in the nucleus accumbens (NAc) during the normal, wild-type (WT) peak time (Zeitgeber Time 14, ZT14) compared with WT littermates #P<0.05 for main effect of genotype by two-way ANOVA; *P<0.05 by post hoc Tukey test; n=5 per genotype. (b) The normal WT peak in PER2 protein expression (ZT22) was also reduced in the NAc and caudate–putamen (CPu) of ENT1 KO mice; *P<0.05 by unpaired, two-tailed t-test; n=8 per genotype. (c) Representative confocal images showing reduced PER2 immunofluorescence at ZT22 in the NAc and CPu of WT (left) vs ENT1 KO mice (right). Colocalization of PER2 with NeuN indicated that PER2 was expressed in neurons. Scale bar, 100 μm. Data are expressed as mean±SEM.
Immunofluorescence confirmed that peak PER2 protein levels (which occur at ZT22, ≈8 h after peak Per2 gene levels) were also reduced, both in the NAc (t=3.153, P=0.004) and caudate–putamen (CPu; t=3.741, P<0.001; Figure 2b). Background-corrected PER2 immunofluorescence-integrated density averaged 9.0 × 106±3.8 × 105 in WT and 6.9 × 106±4.6 × 105 in ENT1 KO mice in the NAc, and 8.4 × 106±3.4 × 105 vs 6.6 × 106±3.2 × 105 in the CPu, respectively. Confocal images showing reduced PER2 immunofluorescence in the striatum are shown in Figure 2c. Colocalization of PER2 with NeuN but not with GFAP (Supplementary Figure S2a) indicated that PER2 was neuronal. No genotype difference in PER2 immunofluorescence was seen in the SCN (Supplementary Figure S2b).
Chronodisruption-Induced Escalation in Ethanol Intake by ENT1 KO Mice
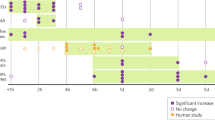
If circadian misalignment influences the baseline heavy-drinking phenotype of ENT1 KO mice (Choi et al, 2004), then chronodisruption should further increase their ethanol consumption. Higher baseline ethanol intake in ENT1 KO mice during training was confirmed (Supplementary Figure S3). Following 2 weeks at 10% ethanol, mice were exposed to 2 weeks LL. ENT1 KO mice consumed more ethanol (F1,103=4.731, P<0.034), while a genotype × photocycle interaction (F1,103=10.736, P=0.002) and post hoc testing revealed increased drinking (P=0.002) by ENT1 KO mice and a genotype difference (P=0.006) during LL (Figure 3a). A genotype × photocycle interaction on ethanol preference (F1,103=8.355, P=0.006) and post hoc testing showed that ethanol preference increased during LL only in ENT1 KO mice (P=0.003; Figure 3b). Representative, double-plotted drinkograms (circadian ethanol drinking patterns) are shown in Figure 3c. No changes in ethanol intake occurred in ENT1 KO or WT mice held in LD for the same duration (Lee et al, 2013). These data indicate that LL-induced chronodisruption further increases ethanol drinking in ENT1 KO mice.
Escalation in ethanol intake by equilibrative nucleoside transporter 1 (ENT1) KO mice during a constant light (LL) photocycle. (a) ENT1 KO mice showed increased ethanol consumption during LL compared with LD, whereas wild-type (WT) mice consumed the same amount of ethanol in LL and LD. (b) ENT1 KO mice also showed increased ethanol preference in LL compared with LD, whereas no such change occurred in WT. (c) Representative, double-plotted drinkograms showing circadian drinking patterns from WT and ENT1 KO mice during LD and LL (stars denote beginning of LL). #P<0.05 for main effect of genotype by two-way, repeated measures ANOVA; *P<0.05 by post hoc Tukey test; n=27 per genotype. Data are expressed as mean±SEM.
A2AR Regulates Per Expression and Circadian Behavior in ENT1 KO Mice
If an adenosine deficit decreases striatal Per expression in ENT1 KO mice, then inhibiting adenosine receptors in WT mice may mimic this change. Pharmacological agents were given for 3 days at ZT22 (PER protein peak); killing took place at ZT14 (per mRNA peak; Figure 4a). Treatment with A1R antagonist DPCPX reduced Per1 expression in the NAc (F1,23=7.906, P=0.003) vs vehicle. A1R antagonist DPCPX (P=0.005) and A2AR antagonist ZM-241385 (P<0.001) decreased Per2 expression (F1,21=12.816, P<0.001) vs vehicle (F1,21=12.816, P<0.001; Figure 4b).
A2A receptor (A2AR)-mediated regulation of Per gene expression and circadian behavioral rhythms in equilibrative nucleoside transporter 1 (ENT1) KO mice. (a) Diagram depicting times of treatment with adenosine receptor ligands (Zeitgeber Time (ZT22); wild-type (WT) peak for PER protein expression in striatum) and killing (ZT14; WT peak for Per gene expression in striatum). (b) Three-day treatment of A1R antagonist DPCPX (6.0 mg/kg, i.p.) or A2AR antagonist ZM-241385 (20.0 mg/kg, i.p.) at ZT22 (late night) differentially affected clock gene expression in WT mice, with the former downregulating peak expression of Per1 and Per2, while the latter only reduced expression of Per2; *P<0.05 by one-way ANOVA; n=6 per treatment. (c) Three-day treatment of A2AR agonist CGS-21680 (2.0 mg/kg) upregulated Per1 only in ENT1 KO mice and Per2 in both WT and ENT1 KO mice compared with vehicle-treated WT controls; P<0.05 by unpaired, two-tailed t-test; n=6 per treatment. (d) Representative actograms from WT (left side) and ENT1 KO (right side) mice treated with vehicle (top) or A2AR agonist CGS-21680 (bottom) for 5 days. Stars denote daily treatment time at ZT22; for simplicity, only the first treatment day is starred. (e) Five-day treatment with CGS-21680 (1.0 mg/kg, i.p.) at ZT22 (late night) delayed activity onset in ENT1 KO, but not in WT mice. (f) Alpha was reduced by 5-day CGS-21680 treatment in both ENT1 KO and WT mice, but to a greater degree in the former; #P<0.05 for main effect of genotype by two-way ANOVA; *P<0.05 by post hoc Tukey test; n=5 per genotype × treatment. Data are expressed as mean±SEM.
A2AR is enriched in the striatum and its inhibition by caffeine in the NAc produces locomotor activation (Lazarus et al, 2011). Increased alcohol consumption (Spanagel et al, 2005) and advanced entrainment (Brager et al, 2011) in Per2 mutant mice suggest that low Per2 expression may account for early entrainment in ENT1 KO mice. As inhibition of A2AR selectively downregulated Per2 expression, we used A2AR agonist CGS-21680 to determine whether A2AR activation would increase Per2. Three-day CGS-21680 treatment increased Per2 levels similarly (≈ 45%) in ENT1 KO and WT mice, but increased Per1 only in ENT1 KO mice relative to vehicle-treated WT controls (t=−3.229, P=0.012; Figure 4c).
If compromised A2AR-mediated striatal Per expression underlies circadian changes in ENT1 KO mice, then increasing Per expression with CGS-21680 should normalize behavior. As expected, we found main effects of genotype (F1,21=8.671, P=0.009), 5-day CGS-21680 treatment (F1,21=13.210, P=0.002), and a genotype × treatment interaction (F1,21=8.783, P=0.008). Post hoc analysis revealed that CGS-21680 delayed onset in ENT1 KO (P<0.001) but not in WT mice. Both genotypes responded similarly to vehicle, but differentially to CGS-21680 (P=0.001; Figure 4e). Main effects of genotype (F1,20=6.634, P=0.020), treatment (F1,20=42.613, P<0.001), and a genotype × treatment interaction (F1,20=5.454, P=0.032) were also shown for alpha. Post hoc analysis showed that CGS-21680 reduced alpha in ENT1 KO (P<0.001) and WT mice (P=0.005), albeit to a greater degree in the former (P=0.004; Figure 3f). Representative actograms of mice from each genotype × treatment are shown in Figure 4d. These data support a role for striatal A2AR-mediated Per expression in circadian activity.
DISCUSSION
We provide strong evidence that adenosine signaling regulates circadian clock gene expression in the striatum. We also demonstrate that endogenous adenosine levels regulate circadian entrainment, nightly activity level, and alcohol intake during chronodisruption. Our study supports a role for adenosine in maintaining circadian timing in reward circuitry and for striatal clock genes in regulating alcohol intake and activity.
The hyperactive phenotype of ENT1 KO mice is consistent with their low adenosine tone and hyperlocomotion produced by caffeine, the latter mediated by inhibition of A2AR in the NAc (El Yacoubi et al, 2000; Lazarus et al, 2011). In line with increased ethanol intake by A2AR KO mice (Naassila et al, 2002), ENT1 deletion also induces striatal A2AR signaling deficits that increase operant reward responding (Nam et al, 2013). In contrast, A2AR KO mice show lower exploratory activity, which may reflect the lack of congruence between constitutive and temporary reduction in A2AR signaling or differences between locomotor measures. In this regard, the present findings illustrate the pitfalls of relying on open-field locomotor testing, which is typically brief (⩽1 h) and performed during the daytime, when rodents sleep. The present data provide a relevant example: genotype differences in activity only occurred at night, whereas daytime circadian and open-field activity is comparable in ENT1 KO and WT mice (Chen et al, 2007; Choi et al, 2004; Ruby et al, 2011). As cellular timing controls the expression of many genes and proteins (Lowrey and Takahashi, 2004; Miller et al, 2007), the time of day at which experiments are performed may be critical in determining the contribution of genes and proteins to behavior.
Per2 mRNA and protein deficits in ENT1 KO mice are intriguing in light of the resemblance ENT1 KO mice bear to Per2 mutants. Both are well-established animal models of glutamatergic dysregulation and compromised astrocyte function in AUD (Choi et al, 2004; Lee et al, 2013; Nam et al, 2011; Spanagel et al, 2005). Early awakening in ENT1 KO mice is also consistent with the role of Per2 in circadian phase regulation (Albrecht et al, 2001). The circadian phenotype in Per2 mutants is severe; they become active 2 h before lights off (Bode et al, 2011; Brager et al, 2011), whereas ENT1 KO were active ≈20 min before ZT12. Per2 mRNA expression in ENT1 KO mice was also advanced relative to WT (ZT10 vs ZT14, respectively), which could indicate that peak PER2 protein expression occurred early. However, Per2 mRNA levels in ENT1 KO mice at ZT10 were still substantially lower than the ZT14 peak for WT mice. Given that PER proteins inhibit their transcripts, examining 24 h striatal PER2 protein expression in ENT1 KO mice may be informative.
ENT1 expression is very low in the hypothalamus (Guillen-Gomez et al, 2004), and therefore may not be a significant source of extracellular adenosine in the SCN. Thus, ENT1 deletion would not be expected to greatly alter SCN adenosine tonus. Although outside the scope of this study, proper assessment would require examining SCN adenosine levels, 24 h SCN PER2 expression, and phase-resetting responses to A1R-gated photic input (Sigworth and Rea, 2003) in ENT1 KO mice. However, the present data support a role for striatal adenosine and Per2 in the observed genotype difference in behavior, as SCN PER2 expression was comparable between genotypes. Moreover, activation of A2AR, which is enriched in the striatum but scant in the hypothalamus, reversed the deficits of ENT1 KO mice. In this regard, other evidence exists for clock genes in reward circuitry contributing to circadian amplitude (Mukherjee et al, 2010).
Our study indicates that cellular circadian timing in the striatum may influence alcohol consumption and preference, particularly during chronodisruption. LL usually lengthens circadian period (Figure 3c) and dampens Per expression (Ohta et al, 2005). Long-term LL (50+ days) constitutively elevates PER2 protein in the murine SCN, which negatively regulates Per2 transcripts (Munoz et al, 2005). Thus, the fact that LL did not change ethanol consumption or preference in WT mice indicates that disruption of SCN timing alone is insufficient to affect drinking. However, the additive effect of LL-induced chronodisruption on ethanol intake in ENT1 KO mice validates the idea that pre-existing circadian misalignment may increase ethanol consumption during environmental challenges to the circadian system.
Our findings implicate ENT1 and adenosine receptor-mediated signaling in alcohol-induced flattening of Per gene rhythms (Chen et al, 2004; Huang et al, 2010). Specifically, A1R inhibition downregulated Per1 and Per2 expression, whereas A2AR inhibition downregulated only Per2. However, A2AR agonist CGS-21680 treatment in Per1/Per2-deficient ENT1 KO mice increased expression of both genes, indicating that A2AR affects Per1 levels as well. Adenosinergic regulation of NAc clock genes has important implications regarding the influence of A1R/A2AR antagonist caffeine on alcohol consumption. Reduced sensitivity to intoxication may underlie the frequently toxic blood alcohol concentration achieved by consuming caffeinated alcoholic beverages (Attwood, 2012; Attwood et al, 2012). Here, A2AR antagonism only once a day for 3 days was sufficient to reduce striatal Per2, which itself increases alcohol intake (Spanagel et al, 2005). Furthermore, the dose of A2AR antagonist ZM-241385 we used herein increases alcohol intake (Nam et al, 2013).
As A2AR is enriched in the striatum, the observed A2AR-mediated improvements in Per expression and circadian behavior in ENT1 KO mice argue that the striatal cellular clock participates in the timing and intensity of daily activity. The dose range of CGS-21680 used also reduces alcohol drinking in mice (Nam et al, 2013), implying a relationship between circadian rhythm improvement and decreased drinking. Similar to several studies (Houchi et al, 2013; Jones et al, 2012; Knapp et al, 2001; Rimondini et al 1997), we observed that CGS-21680 reduced locomotor activity. Alpha (active-phase duration) was reduced in CGS-21680-treated mice of both genotypes vs vehicle-treated WT controls (Figure 4d). This is likely related to late-night treatment timing, which coincided with peak PER2 protein expression, but may have reduced late-night locomotion (Figure 4d and e). However, alpha decreased more markedly in ENT1 KO vs WT mice, as CGS-21680 delayed onset only in ENT1 KO mice. As CGS-21680 was administered during the day in Nam et al (2013), it is also unlikely that hypolocomotion underlies reduced ethanol drinking in that study.
Our results suggest a role for striatal adenosine signaling in psychiatric disorders with a prominent hyperactivity component and animal models thereof. Of note, adult ADHD is linked with blunted Per2 expression (Baird et al, 2012). It is also interesting that the other clock genes with altered circadian profiles in ENT1 KO mice, Clock and Dbp, are associated with ADHD (Kissling et al, 2008; Roybal et al, 2007; Xu et al, 2010), bipolar disorder (Le-Niculescu et al, 2008; Lee et al, 2010; Shi et al, 2008; Soria et al, 2010), anxiety (Le-Niculescu et al, 2011a), and AUD (Le-Niculescu et al, 2011b; Le-Niculescu et al, 2008). Interestingly, ENT1 KO mice display mania-like behavior (Ruby et al, 2011) similar to Dbp KO mice (Le-Niculescu et al, 2008). Stress-reactive Dbp KO mice show reduced prefrontal cortical EAAT2 (Le-Niculescu et al, 2008) consistent with the ENT1 KO striatum (Lee et al, 2013). Thus, future studies examining a role for Dbp in ENT1 KO mice are warranted.
Our findings reveal that striatal A2AR-mediated signaling may influence cellular and behavioral circadian timing and activity level. Although clock timing in the SCN itself does not appear to change alcohol intake, compromised reward network timing may confer susceptibility to increased drinking during chronodisruption. This has important implications for individuals who experience chronic environmental challenges to the circadian system, as in shiftwork, jetlag, and sleep disruption. Our study also implies that striatal dysfunction, which adversely affects mood and drives drug intake, may partly underlie the high prevalence of circadian misalignment in psychiatric and substance use disorders.
FUNDING AND DISCLOSURE
The authors declare no conflict of interest.
References
Alanko L, Stenberg D, Porkka-Heiskanen T (2003). Nitrobenzylthioinosine (NBMPR) binding and nucleoside transporter ENT1 mRNA expression after prolonged wakefulness and recovery sleep in the cortex and basal forebrain of rat. J Sleep Res 12: 299–304.
Albrecht U, Zheng B, Larkin D, Sun ZS, Lee CC (2001). MPer1 and mper2 are essential for normal resetting of the circadian clock. J Biol Rhythms 16: 100–104.
Antle MC, Steen NM, Mistlberger RE (2001). Adenosine and caffeine modulate circadian rhythms in the Syrian hamster. Neuroreport 12: 2901–2905.
Asatryan L, Nam HW, Lee MR, Thakkar MM, Saeed Dar M, Davies DL et al (2011). Implication of the purinergic system in alcohol use disorders. Alcohol Clin Exp Res 35: 584–594.
Attwood AS (2012). Caffeinated alcohol beverages: a public health concern. Alcohol Alcohol 47: 370–371.
Attwood AS, Rogers PJ, Ataya AF, Adams S, Munafo MR (2012). Effects of caffeine on alcohol-related changes in behavioural control and perceived intoxication in light caffeine consumers. Psychopharmacology (Berl) 221: 551–560.
Baird AL, Coogan AN, Siddiqui A, Donev RM, Thome J (2012). Adult attention-deficit hyperactivity disorder is associated with alterations in circadian rhythms at the behavioural, endocrine and molecular levels. Mol Psychiatry 17: 988–995.
Bode B, Taneja R, Rossner MJ, Oster H (2011). Advanced light-entrained activity onsets and restored free-running suprachiasmatic nucleus circadian rhythms in per2/dec mutant mice. Chronobiol Int 28: 737–750.
Brager AJ, Prosser RA, Glass JD (2011). Circadian and acamprosate modulation of elevated ethanol drinking in mPer2 clock gene mutant mice. Chronobiol Int 28: 664–672.
Brower KJ (2003). Insomnia, alcoholism and relapse. Sleep Med Rev 7: 523–539.
Brower KJ, Perron BE (2010). Sleep disturbance as a universal risk factor for relapse in addictions to psychoactive substances. Med Hypoth 74: 928–933.
Chen CP, Kuhn P, Advis JP, Sarkar DK (2004). Chronic ethanol consumption impairs the circadian rhythm of pro-opiomelanocortin and period genes mRNA expression in the hypothalamus of the male rat. J Neurochem 88: 1547–1554.
Chen J, Rinaldo L, Lim SJ, Young H, Messing RO, Choi DS (2007). The type 1 equilibrative nucleoside transporter regulates anxiety-like behavior in mice. Genes Brain Behav 6: 776–783.
Choi DS, Cascini MG, Mailliard W, Young H, Paredes P, McMahon T et al (2004). The type 1 equilibrative nucleoside transporter regulates ethanol intoxication and preference. Nat Neurosci 7: 855–861.
Crusio WE, Goldowitz D, Holmes A, Wolfer D (2009). Standards for the publication of mouse mutant studies. Genes Brain Behav 8: 1–4.
Dunwiddie TV, Masino SA (2001). The role and regulation of adenosine in the central nervous system. Annu Rev Neurosci 24: 31–55.
El Yacoubi M, Ledent C, Menard JF, Parmentier M, Costentin J, Vaugeois JM (2000). The stimulant effects of caffeine on locomotor behaviour in mice are mediated through its blockade of adenosine A(2A) receptors. Br J Pharmacol 129: 1465–1473.
Franken P, Lopez-Molina L, Marcacci L, Schibler U, Tafti M (2000). The transcription factor DBP affects circadian sleep consolidation and rhythmic EEG activity. J Neurosci 20: 617–625.
Gamsby JJ, Templeton EL, Bonvini LA, Wang W, Loros JJ, Dunlap JC et al (2013). The circadian Per1 and Per2 genes influence alcohol intake, reinforcement, and blood alcohol levels. Behav Brain Res 249: 15–21.
Gass N, Ollila HM, Utge S, Partonen T, Kronholm E, Pirkola S et al (2010). Contribution of adenosine related genes to the risk of depression with disturbed sleep. J Affect Disord 126: 134–139.
Gordon NP, Cleary PD, Parker CE, Czeisler CA (1986). The prevalence and health impact of shiftwork. Am J Public Health 76: 1225–1228.
Guillen-Gomez E, Calbet M, Casado J, de Lecea L, Soriano E, Pastor-Anglada M et al (2004). Distribution of CNT2 and ENT1 transcripts in rat brain: selective decrease of CNT2 mRNA in the cerebral cortex of sleep-deprived rats. J Neurochem 90: 883–893.
Halassa MM, Florian C, Fellin T, Munoz JR, Lee SY, Abel T et al (2009). Astrocytic modulation of sleep homeostasis and cognitive consequences of sleep loss. Neuron 61: 213–219.
Hallworth R, Cato M, Colbert C, Rea MA (2002). Presynaptic adenosine A1 receptors regulate retinohypothalamic neurotransmission in the hamster suprachiasmatic nucleus. J Neurobiol 52: 230–240.
Hood S, Cassidy P, Mathewson S, Stewart J, Amir S (2011). Daily morphine injection and withdrawal disrupt 24-h wheel running and PERIOD2 expression patterns in the rat limbic forebrain. Neuroscience 186: 65–75.
Houchi H, Persyn W, Legastelois R, Naassila M (2013). The adenosine A2A receptor agonist CGS 21680 decreases ethanol self-administration in both non-dependent and dependent animals. Addict Biol 18: 812–825.
Huang MC, Ho CW, Chen CH, Liu SC, Chen CC, Leu SJ (2010). Reduced expression of circadian clock genes in male alcoholic patients. Alcohol Clin Exp Res 34: 1899–1904.
Iijima M, Nikaido T, Akiyama M, Moriya T, Shibata S (2002). Methamphetamine-induced, suprachiasmatic nucleus-independent circadian rhythms of activity and mPer gene expression in the striatum of the mouse. Eur J Neurosci 16: 921–929.
Jones CK, Bubser M, Thompson AD, Dickerson JW, Turle-Lorenzo N, Amalric M et al (2012). The metabotropic glutamate receptor 4-positive allosteric modulator VU0364770 produces efficacy alone and in combination with L-DOPA or an adenosine 2A antagonist in preclinical rodent models of Parkinson's disease. J Pharmacol Exp Ther 340: 404–421.
Kim JH, Karpyak VM, Biernacka JM, Nam HW, Lee MR, Preuss UW et al (2011). Functional role of the polymorphic 647 T/C variant of ENT1 (SLC29A1) and its association with alcohol withdrawal seizures. PloS ONE 6: e16331.
Kissling C, Retz W, Wiemann S, Coogan AN, Clement RM, Hunnerkopf R et al (2008). A polymorphism at the 3'-untranslated region of the CLOCK gene is associated with adult attention-deficit hyperactivity disorder. Am J Med Genet B Neuropsychiatr Genet 147: 333–338.
Knapp CM, Foye MM, Cottam N, Ciraulo DA, Kornetsky C (2001). Adenosine agonists CGS 21680 and NECA inhibit the initiation of cocaine self-administration. Pharmacol Biochem Behav 68: 797–803.
Kovanen L, Saarikoski ST, Haukka J, Pirkola S, Aromaa A, Lonnqvist J et al (2010). Circadian clock gene polymorphisms in alcohol use disorders and alcohol consumption. Alcohol Alcohol 45: 303–311.
Lazarus M, Shen HY, Cherasse Y, Qu WM, Huang ZL, Bass CE et al (2011). Arousal effect of caffeine depends on adenosine A2A receptors in the shell of the nucleus accumbens. J Neurosci 31: 10067–10075.
Le-Niculescu H, Balaraman Y, Patel SD, Ayalew M, Gupta J, Kuczenski R et al (2011a). Convergent functional genomics of anxiety disorders: translational identification of genes, biomarkers, pathways and mechanisms. Transl Psychiatry 1: e9.
Le-Niculescu H, Case NJ, Hulvershorn L, Patel SD, Bowker D, Gupta J et al (2011b). Convergent functional genomic studies of omega-3 fatty acids in stress reactivity, bipolar disorder and alcoholism. Transl Psychiatry 1: e4.
Le-Niculescu H, McFarland MJ, Ogden CA, Balaraman Y, Patel S, Tan J et al (2008). Phenomic, convergent functional genomic, and biomarker studies in a stress-reactive genetic animal model of bipolar disorder and co-morbid alcoholism. Am J Med Genet B Neuropsychiatr Genet 147B: 134–166.
Lee KY, Song JY, Kim SH, Kim SC, Joo EJ, Ahn YM et al (2010). Association between CLOCK 3111T/C and preferred circadian phase in Korean patients with bipolar disorder. Prog Neuropsychopharmacol Biol Psychiatry 34: 1196–1201.
Lee MR, Ruby CL, Hinton DJ, Choi S, Adams CA, Young Kang N et al (2013). Striatal Adenosine Signaling Regulates EAAT2 and Astrocytic AQP4 Expression and Alcohol Drinking in Mice. Neuropsychopharmacology 38: 437–445.
Livak KJ, Schmittgen TD (2001). Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25: 402–408.
Lopez-Molina L, Conquet F, Dubois-Dauphin M, Schibler U (1997). The DBP gene is expressed according to a circadian rhythm in the suprachiasmatic nucleus and influences circadian behavior. EMBO J 16: 6762–6771.
Lowrey PL, Takahashi JS (2004). Mammalian circadian biology: elucidating genome-wide levels of temporal organization. Annu Rev Genomics Hum Genet 5: 407–441.
Marpegan L, Swanstrom AE, Chung K, Simon T, Haydon PG, Khan SK et al (2011). Circadian regulation of ATP release in astrocytes. J Neurosci 31: 8342–8350.
Melendez RI, McGinty JF, Kalivas PW, Becker HC (2011). Brain region-specific gene expression changes after chronic intermittent ethanol exposure and early withdrawal in C57BL/6J mice. Addict Biol 17: 351–364.
Miller BH, McDearmon EL, Panda S, Hayes KR, Zhang J, Andrews JL et al (2007). Circadian and CLOCK-controlled regulation of the mouse transcriptome and cell proliferation. Proc Natl Acad Sci USA 104: 3342–3347.
Mukherjee S, Coque L, Cao JL, Kumar J, Chakravarty S, Asaithamby A et al (2010). Knockdown of Clock in the ventral tegmental area through RNA interference results in a mixed state of mania and depression-like behavior. Biol Psychiatry 68: 503–511.
Munoz M, Peirson SN, Hankins MW, Foster RG (2005). Long-term constant light induces constitutive elevated expression of mPER2 protein in the murine SCN: a molecular basis for Aschoff's rule? J Biol Rhythms 20: 3–14.
Murillo-Rodriguez E, Blanco-Centurion C, Gerashchenko D, Salin-Pascual RJ, Shiromani PJ (2004). The diurnal rhythm of adenosine levels in the basal forebrain of young and old rats. Neuroscience 123: 361–370.
Naassila M, Ledent C, Daoust M (2002). Low ethanol sensitivity and increased ethanol consumption in mice lacking adenosine A2A receptors. J Neurosci 22: 10487–10493.
Nagy LE, Diamond I, Casso DJ, Franklin C, Gordon AS (1990). Ethanol increases extracellular adenosine by inhibiting adenosine uptake via the nucleoside transporter. J Biol Chem 265: 1946–1951.
Nagy LE, Diamond I, Collier K, Lopez L, Ullman B, Gordon AS (1989). Adenosine is required for ethanol-induced heterologous desensitization. Mol Pharmacol 36: 744–748.
Nam HW, Hinton DJ, Kang NY, Kim TH, Lee MR, Oliveros A et al (2013). Adenosine transporter ENT1 regulates the acquisition of goal-directed behavior and ethanol drinking through A2A receptors in the dorsomedial striatum. J Neurosci 33: 4329–4338.
Nam HW, Lee MR, Zhu Y, Wu J, Hinton DJ, Choi S et al (2011). Type 1 equilibrative nucleoside transporter regulates ethanol drinking through accumbal N-methyl-D-aspartate receptor signaling. Biol Psychiatry 69: 1043–1051.
Ohta H, Yamazaki S, McMahon DG (2005). Constant light desynchronizes mammalian clock neurons. Nat Neurosci 8: 267–269.
Oike H, Kobori M, Suzuki T, Ishida N (2011). Caffeine lengthens circadian rhythms in mice. Biochem Biophys Res Comm 410: 654–658.
Perreau-Lenz S, Vengeliene V, Noori HR, Merlo-Pich EV, Corsi MA, Corti C et al (2012). Inhibition of the casein-kinase-1-epsilon/delta prevents relapse-like alcohol drinking. Neuropsychopharmacology 37: 2121–2131.
Rimondini R, Ferre S, Ogren SO, Fuxe K (1997). Adenosine A2A agonists: a potential new type of atypical antipsychotic. Neuropsychopharmacology 17: 82–91.
Roehrs T, Roth T (2001). Sleep, sleepiness, and alcohol use. Alcohol Res Health 25: 101–109.
Rosenwasser AM (2010). Circadian clock genes: non-circadian roles in sleep, addiction, and psychiatric disorders? Neurosci Biobehav Rev 34: 1249–1255.
Roybal K, Theobold D, Graham A, DiNieri JA, Russo SJ, Krishnan V et al (2007). Mania-like behavior induced by disruption of CLOCK. Proc Natl Acad Sci USA 104: 6406–6411.
Ruby CL, Adams CA, Knight EJ, Nam HW, Choi DS (2010). An essential role for adenosine signaling in alcohol abuse. Curr Drug Abuse Rev 3: 163–174.
Ruby CL, Brager AJ, DePaul MA, Prosser RA, Glass JD (2009). Chronic ethanol attenuates circadian photic phase resetting and alters nocturnal activity patterns in the hamster. Am J Physiol Regul Integr Comp Physiol 297: R729–R737.
Ruby CL, Walker DL, An J, Kim J, Choi DS (2011). Sex-specific regulation of depression, anxiety-like behaviors and alcohol drinking in mice lacking ENT1. J Addict Res Ther S4: 1–6.
Seggio JA, Fixaris MC, Reed JD, Logan RW, Rosenwasser AM (2009). Chronic ethanol intake alters circadian phase shifting and free-running period in mice. J Biol Rhythms 24: 304–312.
Shi J, Wittke-Thompson JK, Badner JA, Hattori E, Potash JB, Willour VL et al (2008). Clock genes may influence bipolar disorder susceptibility and dysfunctional circadian rhythm. Am J Med Genet B Neuropsychiatr Genet 147B: 1047–1055.
Sigworth LA, Rea MA (2003). Adenosine A1 receptors regulate the response of the mouse circadian clock to light. Brain Res 960: 246–251.
Smart RG (1979). Drinking problems among employed, unemployed and shift workers. J Occup Med 21: 731–736.
Soria V, Martinez-Amoros E, Escaramis G, Valero J, Perez-Egea R, Garcia C et al (2010). Differential association of circadian genes with mood disorders: CRY1 and NPAS2 are associated with unipolar major depression and CLOCK and VIP with bipolar disorder. Neuropsychopharmacology 35: 1279–1289.
Spanagel R, Pendyala G, Abarca C, Zghoul T, Sanchis-Segura C, Magnone MC et al (2005). The clock gene Per2 influences the glutamatergic system and modulates alcohol consumption. Nat Med 11: 35–42.
Steinlechner S, Jacobmeier B, Scherbarth F, Dernbach H, Kruse F, Albrecht U (2002). Robust circadian rhythmicity of Per1 and Per2 mutant mice in constant light, and dynamics of Per1 and Per2 gene expression under long and short photoperiods. J Biol Rhythms 17: 202–209.
Virus RM, Baglajewski T, Radulovacki M (1984). Circadian variation of [3H]N6-(L-phenylisopropyl)adenosine binding in rat brain. Neurosci Lett 46: 219–222.
Wang X, Mozhui K, Li Z, Mulligan MK, Ingels JF, Zhou X et al (2012). A promoter polymorphism in the Per3 gene is associated with alcohol and stress response. Transl Psychiatry 2: e73.
Wongchitrat P, Mukda S, Phansuwan-Pujito P, Govitrapong P (2013). Effect of amphetamine on the clock gene expression in rat striatum. Neurosci Lett 542: 126–130.
Wu J, Lee MR, Choi S, Kim T, Choi DS (2010). ENT1 regulates ethanol-sensitive EAAT2 expression and function in astrocytes. Alcohol Clin Exp Res 34: 1110–1117.
Xu X, Breen G, Chen CK, Huang YS, Wu YY, Asherson P (2010). Association study between a polymorphism at the 3'-untranslated region of CLOCK gene and attention deficit hyperactivity disorder. Behav Brain Funct 6: 48.
Yang BZ, Han S, Kranzler HR, Farrer LA, Elston RC, Gelernter J (2012). Autosomal linkage scan for loci predisposing to comorbid dependence on multiple substances. Am J Med Genet B Neuropsychiatr Genet 159B: 361–369.
Acknowledgements
This project was funded by the Samuel C. Johnson Genomics of Addiction Program, the Ulm Foundation, the Godby Foundation at Mayo Clinic and in parts by grants from the National Institutes of Health (AA018779). We thank Mr Brandon Boudreaux for his assistance in editing the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Neuropsychopharmacology website
Rights and permissions
About this article
Cite this article
Ruby, C., Vadnie, C., Hinton, D. et al. Adenosinergic Regulation of Striatal Clock Gene Expression and Ethanol Intake During Constant Light. Neuropsychopharmacol 39, 2432–2440 (2014). https://doi.org/10.1038/npp.2014.94
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/npp.2014.94
This article is cited by
-
Neuroimaging of inflammation in alcohol use disorder: a review
Science China Information Sciences (2020)
-
Adenosine A2A receptor mediates hypnotic effects of ethanol in mice
Scientific Reports (2017)