Abstract
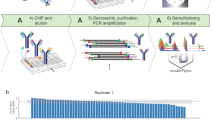
Chromatin and transcriptional processes are among the most intensively studied fields of biology today. The introduction of chromatin immunoprecipitations (ChIP) represents a major advancement in this area. This powerful method allows researchers to probe specific protein-DNA interactions in vivo and to estimate the density of proteins at specific sites genome-wide. We have introduced several improvements to the traditional ChIP assay, which simplify the procedure, greatly reducing the time and labor required to complete the assay. The simplicity of the method yields highly reproducible results. Our improvements facilitate the probing of multiple proteins in a single experiment, which allows for the simultaneous monitoring of many genomic events. This method is particularly useful in kinetic studies where multiple samples are processed at the same time. Starting with sheared chromatin, PCR-ready DNA can be isolated from 16–24 ChIP samples in 4–6 h using the fast method.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bernstein, E. & Allis, C.D. RNA meets chromatin. Genes Dev. 19, 1635–1655 (2005).
Schubeler, D. & Elgin, S.C. Defining epigenetic states through chromatin and RNA. Nat. Genet. 37, 917–918 (2005).
Felsenfeld, G. & Groudine, M. Controlling the double helix. Nature 421, 448–453 (2003).
Sims, R.J. III, Mandal, S.S. & Reinberg, D. Recent highlights of RNA-polymerase-II-mediated transcription. Curr. Opin. Cell Biol. 16, 263–271 (2004).
Thiriet, C. & Hayes, J.J. Chromatin in need of a fix: phosphorylation of H2AX connects chromatin to DNA repair. Mol. Cell 18, 617–622 (2005).
Kuo, M.H. & Allis, C.D. In vivo cross-linking and immunoprecipitation for studying dynamic Protein: DNA associations in a chromatin environment. Methods 19, 425–433 (1999).
Orlando, V., Strutt, H. & Paro, R. Analysis of chromatin structure by in vivo formaldehyde cross-linking. Methods 11, 205–214 (1997).
Impey, S. et al. Defining the CREB regulon: a genome-wide analysis of transcription factor regulatory regions. Cell 119, 1041–1054 (2004).
Nelson, J.D., Denisenko, O., Sova, P. & Bomsztyk, K. Fast chromatin immunoprecipitation assay. Nucleic Acids Res. 34, e2 (2006).
Ostrowski, J., Kawata, Y., Schullery, D.S., Denisenko, O.N. & Bomsztyk, K. Transient recruitment of the hnRNP K protein to inducibly transcribed gene loci. Nucleic Acids Res. 31, 3954–3962 (2003).
Dundr, M. et al. A kinetic framework for a mammalian RNA polymerase in vivo. Science 298, 1623–1626 (2002).
Cheutin, T. et al. Maintenance of stable heterochromatin domains by dynamic HP1 binding. Science 299, 721–725 (2003).
Cosma, M.P., Tanaka, T. & Nasmyth, K. Ordered recruitment of transcription and chromatin remodeling factors to a cell cycle- and developmentally regulated promoter. Cell 97, 299–311 (1999).
Liu, C.L. et al. Single-nucleosome mapping of histone modifications in S. cerevisiae. PLoS Biol. 3, e328 (2005).
Metivier, R. et al. Transcriptional complexes engaged by apo-estrogen receptor-alpha isoforms have divergent outcomes. EMBO J. 23, 3653–3666 (2004).
Koyanagi, M. et al. EZH2 and histone 3 trimethyl lysine 27 associated with Il4 and Il13 gene silencing in Th1 cells. J. Biol. Chem. 280, 31470–31477 (2005).
Solomon, M.J. & Varshavsky, A. Formaldehyde-mediated DNA-protein crosslinking: a probe for in vivo chromatin structures. Proc. Natl. Acad. Sci. USA 82, 6470–6474 (1985).
Thorne, A.W., Myers, F.A. & Hebbes, T.R. Native chromatin immunoprecipitation. Methods Mol. Biol. 287, 21–44 (2004).
Bernstein, B.E. et al. Genomic maps and comparative analysis of histone modifications in human and mouse. Cell 120, 169–181 (2005).
Pokholok, D.K. et al. Genome-wide map of nucleosome acetylation and methylation in yeast. Cell 122, 517–527 (2005).
Cawley, S. et al. Unbiased mapping of transcription factor binding sites along human chromosomes 21 and 22 points to widespread regulation of noncoding RNAs. Cell 116, 499–509 (2004).
Orlando, V. Mapping chromosomal proteins in vivo by formaldehyde-crosslinked- chromatin immunoprecipitation. Trends Biochem. Sci. 25, 99–104 (2000).
Chen, R. et al. Ultrasound-accelerated immunoassay, as exemplified by enzyme immunoassay of choriogonadotropin. Clin. Chem. 30, 1446–1451 (1984).
Chaya, D. & Zaret, K.S. Sequential chromatin immunoprecipitation from animal tissues. Methods Enzymol. 376, 361–372 (2004).
Schwabish, M.A. & Struhl, K. Evidence for eviction and rapid deposition of histones upon transcriptional elongation by RNA polymerase II. Mol. Cell. Biol. 24, 10111–10117 (2004).
Denisenko, O. & Bomsztyk, K. Yeast hnRNP K-like genes are involved in regulation of the telomeric position effect and telomere length. Mol. Cell. Biol. 22, 286–297 (2002).
Kurdistani, S.K., Tavazoie, S. & Grunstein, M. Mapping global histone acetylation patterns to gene expression. Cell 117, 721–733 (2004).
Sandoval, J. et al. RNAPol-ChIP: a novel application of chromatin immunoprecipitation to the analysis of real-time gene transcription. Nucleic Acids Res. 32, e88 (2004).
Waugh, D.S. Making the most of affinity tags. Trends Biotechnol. 23, 316–320 (2005).
Van Seuningen, I., Ostrowski, J. & Bomsztyk, K. Description of an IL-1-responsive kinase that phosphorylates the K protein. Enhancement of phosphorylation by sequence-selective DNA and RNA motifs. Biochemistry 34, 5644–5650 (1995).
Acknowledgements
We thank members of the K.B. lab for valuable discussions of the method. This work was supported by the US National Institutes of Health (DK45978 and GM45134) and the Juvenile Diabetes Research Foundation (K.B.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Nelson, J., Denisenko, O. & Bomsztyk, K. Protocol for the fast chromatin immunoprecipitation (ChIP) method. Nat Protoc 1, 179–185 (2006). https://doi.org/10.1038/nprot.2006.27
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.27
This article is cited by
-
PARylated PDHE1α generates acetyl-CoA for local chromatin acetylation and DNA damage repair
Nature Structural & Molecular Biology (2023)
-
Functional and structural diversification of incomplete phosphotransferase system in cellulose-degrading clostridia
The ISME Journal (2023)
-
Endogenous bioluminescent reporters reveal a sustained increase in utrophin gene expression upon EZH2 and ERK1/2 inhibition
Communications Biology (2023)
-
BAP18 facilitates CTCF-mediated chromatin accessible to regulate enhancer activity in breast cancer
Cell Death & Differentiation (2023)
-
Glucocorticoid activation of anti-inflammatory macrophages protects against insulin resistance
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.