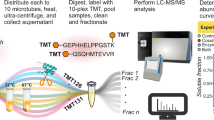
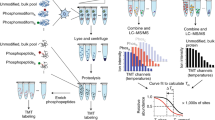
Abstract
Activity-based protein profiling (ABPP) utilizes active site-directed chemical probes to monitor the functional state of enzymes directly in native biological systems. Identification of the specific sites of probe labeling on enzymes remains a major challenge in ABPP experiments. In this protocol, we describe an advanced ABPP platform that utilizes a tandem orthogonal proteolysis (TOP) strategy coupled with mass spectrometric analysis to simultaneously identify probe-labeled proteins together with their exact sites of probe modification. Elucidation of probe modification sites reveals fundamental insights into the molecular basis of specific probe–protein interactions. The TOP-ABPP method can be applied to any type of proteomic sample, including those derived from in vitro or in vivo labeling experiments, and is compatible with a variety of chemical probe structures. Completion of the entire protocol, including chemical synthesis of key reagents, requires approximately 8–10 days.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Anderson, N.L. & Anderson, N.G. Proteome and proteomics: new technologies, new concepts, and new words. Electrophoresis 19, 1853–1861 (1998).
Mann, M., Hendrickson, R.C. & Pandey, A. Analysis of proteins and proteomes by mass spectrometry. Annu. Rev. Biochem. 70, 437–473 (2001).
Kobe, B. & Kemp, B.E. Active site-directed protein regulation. Nature 402, 373–376 (1999).
Evans, M.J. & Cravatt, B.F. Mechanism-based profiling of enzyme families. Chem. Rev. 106, 3279–3301 (2006).
Oda, Y., Nagasu, T. & Chait, B.T. Enrichment analysis of phosphorylated proteins as a tool for probing the phosphoproteome. Nat. Biotechnol. 19, 379–382 (2001).
Zhou, H., Watts, J.D. & Aebersold, R. A systematic approach to the analysis of protein phosphorylation. Nat. Biotechnol. 19, 375–378 (2001).
Tai, H.C., Khidekel, N., Ficarro, S.B., Peters, E.C. & Hsieh-Wilson, L.C. Parallel identification of O-GlcNAc-modified proteins from cell lysates. J. Am. Chem. Soc. 126, 10500–10501 (2004).
Vocadlo, D.J., Hang, H.C., Kim, E.J., Hanover, J.A. & Bertozzi, C.R. A chemical approach for identifying O-GlcNAc-modified proteins in cells. Proc. Natl. Acad. Sci. USA 100, 9116–9121 (2003).
Adam, G.C., Sorensen, E.J. & Cravatt, B.F. Chemical strategies for functional proteomics. Mol. Cell. Proteomics 1, 781–790 (2002).
Speers, A.E. & Cravatt, B.F. Chemical strategies for activity-based proteomics. Chembiochem 5, 41–47 (2004).
Liu, Y., Patricelli, M.P. & Cravatt, B.F. Activity-based protein profiling: the serine hydrolases. Proc. Natl. Acad. Sci. USA 96, 14694–14699 (1999).
Kato, D. et al. Activity-based probes that target diverse cysteine protease families. Nat. Chem. Biol. 1, 33–38 (2005).
Patricelli, M.P. et al. Functional interrogation of the kinome using nucleotide acyl phosphates. Biochemistry 46, 350–358 (2007).
Adam, G.C., Sorensen, E.J. & Cravatt, B.F. Proteomic profiling of mechanistically distinct enzyme classes using a common chemotype. Nat. Biotechnol. 20, 805–809 (2002).
Barglow, K.T. & Cravatt, B.F. Discovering disease-associated enzymes by proteome reactivity profiling. Chem. Biol. 11, 1523–31 (2004).
Evans, M.J., Saghatelian, A., Sorensen, E.J. & Cravatt, B.F. Target discovery in small-molecule cell-based screens by in situ proteome reactivity profiling. Nat. Biotechnol. 23, 1303–1307 (2005).
Speers, A.E., Adam, G.C. & Cravatt, B.F. Activity-based protein profiling in vivo using a copper(i)-catalyzed azide-alkyne [3 + 2] cycloaddition. J. Am. Chem. Soc. 125, 4686–4687 (2003).
Speers, A.E. & Cravatt, B.F. Profiling enzyme activities in vivo using click chemistry methods. Chem. Biol. 11, 535–546 (2004).
Kolb, H.C., Finn, M.G. & Sharpless, K.B. Click chemistry: diverse chemical function from a few good reactions. Angew. Chem. Int. Ed. Engl. 40, 2004–2021 (2001).
Speers, A.E. & Cravatt, B.F. A tandem orthogonal proteolysis strategy for high-content chemical proteomics. J. Am. Chem. Soc. 127, 10018–10019 (2005).
Dougherty, W.G., Cary, S.M. & Parks, T.D. Molecular genetic analysis of a plant virus polyprotein cleavage site: a model. Virology 171, 356–364 (1989).
Washburn, M.P., Wolters, D. & Yates, J.R. 3rd Large-scale analysis of the yeast proteome by multidimensional protein identification technology. Nat. Biotechnol. 19, 242–247 (2001).
Adam, G.C., Burbaum, J., Kozarich, J.W., Patricelli, M.P. & Cravatt, B.F. Mapping enzyme active sites in complex proteomes. J. Am. Chem. Soc. 126, 1363–1368 (2004).
Okerberg, E.S. et al. High-resolution functional proteomics by active-site peptide profiling. Proc. Natl. Acad. Sci. USA 102, 4996–5001 (2005).
Jessani, N. et al. A streamlined platform for high-content functional proteomics of primary human specimens. Nat. Methods 2, 691–697 (2005).
Liu, H., Sadygov, R.G. & Yates, J.R. 3rd A model for random sampling and estimation of relative protein abundance in shotgun proteomics. Anal. Chem. 76, 4193–4201 (2004).
Old, W.M. et al. Comparison of label-free methods for quantifying human proteins by shotgun proteomics. Mol. Cell. Proteomics 4, 1487–1502 (2005).
Cooper, H.J., Hakansson, K. & Marshall, A.G. The role of electron capture dissociation in biomolecular analysis. Mass Spectrom. Rev. 24, 201–222 (2005).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Weerapana, E., Speers, A. & Cravatt, B. Tandem orthogonal proteolysis-activity-based protein profiling (TOP-ABPP)—a general method for mapping sites of probe modification in proteomes. Nat Protoc 2, 1414–1425 (2007). https://doi.org/10.1038/nprot.2007.194
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2007.194
This article is cited by
-
Bioorthogonal photocatalytic proximity labeling in primary living samples
Nature Communications (2024)
-
Bactericidal bissulfone B7 targets bacterial pyruvate kinase to impair bacterial biology and pathogenicity in plants
Science China Life Sciences (2024)
-
Monitoring Fe–S cluster occupancy across the E. coli proteome using chemoproteomics
Nature Chemical Biology (2023)
-
Spatiotemporal-resolved protein networks profiling with photoactivation dependent proximity labeling
Nature Communications (2022)
-
Advances in covalent drug discovery
Nature Reviews Drug Discovery (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.