Abstract
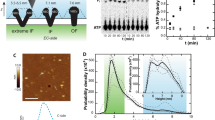
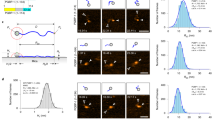
Membrane proteins comprise 30% of the proteome of higher organisms. They mediate energy conversion, signal transduction, solute transport and secretion. Their native environment is a bilayer in a physiological buffer solution, hence their structure and function are preferably assessed in this environment. The surface structure of single membrane proteins can be determined in buffer solutions by atomic force microscopy (AFM) at a lateral resolution of less than 1 nm and a vertical resolution of 0.1–0.2 nm. Moreover, single proteins can be directly addressed, stuck to the AFM stylus and subsequently unfolded, revealing the molecular interactions of the protein studied. The examples discussed here illustrate the power of AFM in the structural analysis of membrane proteins in a native environment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Müller, D.J., Fotiadis, D., Scheuring, S., Müller, S.A. & Engel, A. Electrostatically balanced subnanometer imaging of biological specimens by atomic force microscope. Biophys. J. 76, 1101–1111 (1999).
Hoogenboom, B.W. et al. Quantitative dynamic-mode scanning force microscopy in liquid. Appl. Phys. Lett. 88, 193109 (2006).
Putman, C.A., van der Werf, K.O., de Grooth, B.G., van Hulst, N.F. & Greve, J. Tapping mode atomic force microscopy in liquid. Appl. Phys. Lett. 64, 2454–2456 (1994).
Hansma, P.K. et al. Tapping mode atomic force microscopy in liquids. Appl. Phys. Lett. 64, 1738–1740 (1994).
Engel, A. & Müller, D.J. Observing single biomolecules at work with the atomic force microscope. Nat. Struct. Biol. 7, 715–718 (2000).
Yokokawa, M. et al. Fast-scanning atomic force microscopy reveals the ATP/ADP-dependent conformational changes of GroEL. EMBO J. 25, 4567–4576 (2006).
Oesterhelt, F. et al. Unfolding pathways of individual bacteriorhodopsins. Science 288, 143–146 (2000).
Müller, D.J., Büldt, G. & Engel, A. Force-induced conformational change of bacteriorhodopsin. J. Mol. Biol. 249, 239–243 (1995).
Viani, M.B. et al. Small cantilevers for force spectroscopy of single molecules. J. Appl. Phys. 86, 2258–2262 (1999).
Ando, T. et al. A high-speed atomic force microscope for studying biological macromolecules. Proc. Natl. Acad. Sci. USA 98, 12468–12472 (2001).
Yasumura, K.Y. et al. Quality factors in micron- and submicron-thick cantilevers. J. Microelectromech. Syst. 9, 117–125 (2000).
Chon, J.W.M., Mulvaney, P. & Sader, J.E. Experimental validation of theoretical models for the frequency response of atomic force microscope cantilever beams immersed in fluids. J. Appl. Phys. 87, 3978–3988 (2000).
Sader, J.E., Chon, J.W.M. & Mulvaney, P. Calibration of rectangular atomic force microscope cantilevers. Rev. Sci. Instrum. 70, 3967–3969 (1999).
Czajkowsky, D.M., Hotze, E.M., Shao, Z. & Tweten, R.K. Vertical collapse of a cytolysin prepore moves its transmembrane beta-hairpins to the membrane. EMBO J. 23, 3206–3215 (2004).
Fotiadis, D. et al. Structural analysis of the reaction center light-harvesting complex I photosynthetic core complex of Rhodospirillum rubrum using atomic force microscopy. J. Biol. Chem. 279, 2063–2068 (2004).
Scheuring, S. & Sturgis, J.N. Chromatic adaptation of photosynthetic membranes. Science 309, 484–487 (2005).
Müller, D.J. & Engel, A. Voltage and pH-induced channel closure of porin OmpF visualized by atomic force microscopy. J. Mol. Biol. 285, 1347–1351 (1999).
Müller, D.J., Amrein, M. & Engel, A. Adsorption of biological molecules to a solid support for scanning probe microscopy. J. Struct. Biol. 119, 172–188 (1997).
Butt, H.J. et al. Scan speed limit in atomic force microscopy. J. Microsc. 169, 75–84 (1993).
Engel, A., Schoenenberger, C.A. & Müller, D.J. High-resolution imaging of native biological sample surfaces using scanning probe microscopy. Curr. Opin. Struct. Biol. 7, 279–284 (1997).
Schwarz, U.D., Haefke, H., Reimann, P. & Guntherodt, H.J. Tip artefacts in scanning force microscopy. J. Microsc. 173, 183–197 (1994).
Möller, C., Allen, M., Elings, V., Engel, A. & Müller, D.J. Tapping-mode atomic force microscopy produces faithful high-resolution images of protein surfaces. Biophys. J. 77, 1150–1158 (1999).
Stark, M., Möller, C., Müller, D.J. & Guckenberger, R. From images to interactions: high-resolution phase imaging in tapping-mode atomic force microscopy. Biophys. J. 80, 3009–3018 (2001).
Garcia, R. & Perez, R. Dynamic atomic force microscopy methods. Surf. Sci. Rep. 47, 197–301 (2002).
Kedrov, A., Janovjak, H., Sapra, K.T. & Muller, D.J. Deciphering molecular interactions of native membrane proteins by single-molecule force spectroscopy. Annu. Rev. Biophys. Biomol. Struct. 36, 233–260 (2007).
Seelert, H. et al. Structural biology. Proton-powered turbine of a plant motor. Nature 405, 418–419 (2000).
Stahlberg, H. et al. Bacterial Na+-ATP synthase has an undecameric rotor. EMBO Rep. 2, 229–233 (2001).
Fotiadis, D. et al. Atomic-force microscopy: rhodopsin dimers in native disc membranes. Nature 421, 127–128 (2003).
Yu, J., Bippes, C.A., Hand, G.M., Muller, D.J. & Sosinsky, G.E. Aminosulfonate modulated pH-induced conformational changes in connexin26 hemichannels. J. Biol. Chem. 282, 8895–8904 (2007).
Acknowledgements
The authors acknowledge support by the Deutsche Forschungsgemeinschaft, Volkswagenstiftung, European Union, the Free State of Saxony, the Swiss National Research Foundation, the M.E. Müller Foundation, the Swiss National Center of Competence in Research (NCCR) 'Structural Biology', the NCCR 'Nanoscale Science' and the NANOMOT project of the EU (grant NEST2004 PathSYS29084).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Müller, D., Engel, A. Atomic force microscopy and spectroscopy of native membrane proteins. Nat Protoc 2, 2191–2197 (2007). https://doi.org/10.1038/nprot.2007.309
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2007.309
This article is cited by
-
Gasdermin-A3 pore formation propagates along variable pathways
Nature Communications (2022)
-
Imaging in Biologically-Relevant Environments with AFM Using Stiff qPlus Sensors
Scientific Reports (2018)
-
The Hessian Blob Algorithm: Precise Particle Detection in Atomic Force Microscopy Imagery
Scientific Reports (2018)
-
Protein-enriched outer membrane vesicles as a native platform for outer membrane protein studies
Communications Biology (2018)
-
Microfluidic deposition for resolving single-molecule protein architecture and heterogeneity
Nature Communications (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.