Abstract
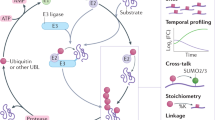
The small ubiquitin-like modifiers (SUMOs) are posttranslationally conjugated to eukaryotic cellular proteins with generally unpredictable consequences. SUMO substrates are found in many cellular systems, and functional analysis has revealed that substrate SUMOylation often has an important role in their regulation. Here we describe a cell-based protocol which can be used to detect the SUMOylation of a protein that relies on the enrichment of SUMO conjugates by purification of 6His-SUMO under denaturing conditions, followed by western blotting for the protein of interest. By purifying under denaturing conditions this method not only reduces the risk of false-positive identifications by non-covalent interactions, but also preserves SUMO-substrate conjugates by inhibiting SUMO-specific proteases—two caveats that may complicate other less stringent purification methods. In preliminary form, this protocol takes 4–5 d to perform, and it can be further elaborated to provide a multi-angled approach to investigate protein conjugation by SUMO.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Johnson, E.S. Protein modification by SUMO. Annu. Rev. Biochem. 73, 355–382 (2004).
Geiss-Friedlander, R. & Melchior, F. Concepts in sumoylation: a decade on. Nat. Rev. 8, 947–956 (2007).
Hayashi, T. et al. Ubc9 is essential for viability of higher eukaryotic cells. Exp. Cell Res. 280, 212–221 (2002).
Seufert, W., Futcher, B. & Jentsch, S. Role of a ubiquitin-conjugating enzyme in degradation of S- and M-phase cyclins. Nature 373, 78–81 (1995).
Nacerddine, K. et al. The SUMO pathway is essential for nuclear integrity and chromosome segregation in mice. Dev. Cell 9, 769–779 (2005).
Johnson, E.S., Schwienhorst, I., Dohmen, R.J. & Blobel, G. The ubiquitin-like protein Smt3p is activated for conjugation to other proteins by an Aos1p/Uba2p heterodimer. EMBO J. 16, 5509–5519 (1997).
Kerscher, O., Felberbaum, R. & Hochstrasser, M. Modification of proteins by ubiquitin and ubiquitin-like proteins. Annu. Rev. Cell Dev. Biol. 22, 159–180 (2006).
Rodriguez, M.S., Dargemont, C. & Hay, R.T. SUMO-1 conjugation in vivo requires both a consensus modification motif and nuclear targeting. J. Biol. Chem. 276, 12654–12659 (2001).
Sampson, D.A., Wang, M. & Matunis, M.J. The small ubiquitin-like modifier-1 (SUMO-1) consensus sequence mediates Ubc9 binding and is essential for SUMO-1 modification. J. Biol. Chem. 276, 21664–21669 (2001).
Vertegaal, A.C. et al. Distinct and overlapping sets of SUMO-1 and SUMO-2 target proteins revealed by quantitative proteomics. Mol. Cell Proteomics 5, 2298–2310 (2006).
Saitoh, H. & Hinchey, J. Functional heterogeneity of small ubiquitin-related protein modifiers SUMO-1 versus SUMO-2/3. J. Biol. Chem. 275, 6252–6258 (2000).
Mahajan, R., Delphin, C., Guan, T., Gerace, L. & Melchior, F. A small ubiquitin-related polypeptide involved in targeting RanGAP1 to nuclear pore complex protein RanBP2. Cell 88, 97–107 (1997).
Matunis, M.J., Wu, J. & Blobel, G. SUMO-1 modification and its role in targeting the Ran GTPase-activating protein, RanGAP1, to the nuclear pore complex. J. Cell Biol. 140, 499–509 (1998).
Lallemand-Breitenbach, V. et al. Role of promyelocytic leukemia (PML) sumolation in nuclear body formation, 11S proteasome recruitment, and As2O3-induced PML or PML/retinoic acid receptor alpha degradation. J. Exp. Med. 193, 1361–1371 (2001).
Girdwood, D. et al. P300 transcriptional repression is mediated by SUMO modification. Mol. Cell 11, 1043–1054 (2003).
Desterro, J.M., Rodriguez, M.S. & Hay, R.T. SUMO-1 modification of IkBa inhibits NF-kB activation. Mol. Cell 2, 233–239 (1998).
Hoege, C., Pfander, B., Moldovan, G.L., Pyrowolakis, G. & Jentsch, S. RAD6-dependent DNA repair is linked to modification of PCNA by ubiquitin and SUMO. Nature 419, 135–141 (2002).
Rosas-Acosta, G., Russell, W.K., Deyrieux, A., Russell, D.H. & Wilson, V.G. A universal strategy for proteomic studies of SUMO and other ubiquitin-like modifiers. Mol. Cell Proteomics 4, 56–72 (2005).
Blomster, H.A. et al. Novel proteomics strategy brings insight into the prevalence of SUMO-2 target sites. Mol. Cell Proteomics 8, 1382–1390 (2009).
Golebiowski, F. et al. System-wide changes to SUMO modifications in response to heat shock. Sci. Signal 2, ra24 (2009).
Schimmel, J. et al. The ubiquitin-proteasome system is a key component of the SUMO-2/3 cycle. Mol. Cell Proteomics 7, 277–289 (2008).
Gocke, C.B. & Yu, H. Identification of SUMO targets through in vitro expression cloning. Methods Mol. Biol. 497, 51–61 (2009).
Werner, A., Moutty, M.C., Moller, U. & Melchior, F. Performing in vitro sumoylation reactions using recombinant enzymes. Methods Mol. Biol. 497, 187–199 (2009).
Xirodimas, D.P., Saville, M.K., Bourdon, J.C., Hay, R.T. & Lane, D.P. Mdm2-mediated NEDD8 conjugation of p53 inhibits its transcriptional activity. Cell 118, 83–97 (2004).
Rodriguez, M.S. et al. SUMO-1 modification activates the transcriptional response of p53. EMBO J. 18, 6455–6461 (1999).
Vertegaal, A.C. et al. A proteomic study of SUMO-2 target proteins. J. Biol. Chem. 279, 33791–33798 (2004).
Vethantham, V., Rao, N. & Manley, J.L. Sumoylation regulates multiple aspects of mammalian poly(A) polymerase function. Genes Dev. 22, 499–511 (2008).
Jaffray, E.G. & Hay, R.T. Detection of modification by ubiquitin-like proteins. Methods 38, 35–38 (2006).
Bailey, D. & O'Hare, P. Comparison of the SUMO1 and ubiquitin conjugation pathways during the inhibition of proteasome activity with evidence of SUMO1 recycling. Biochem. J. 392, 271–281 (2005).
Windecker, H. & Ulrich, H.D. Architecture and assembly of poly-SUMO chains on PCNA in Saccharomyces cerevisiae. J. Mol. Biol. 376, 221–231 (2008).
Girdwood, D.W., Tatham, M.H. & Hay, R.T. SUMO and transcriptional regulation. Semin. Cell Dev. Biol. 15, 201–210 (2004).
Gill, G. Something about SUMO inhibits transcription. Curr. Opin. Genet. Dev. 15, 536–541 (2005).
Stehmeier, P. & Muller, S. Regulation of p53 family members by the ubiquitin-like SUMO system. DNA Repair 8, 491–498 (2009).
Acknowledgements
M.H.T. is supported by a Cancer research UK program grant, and M.S.R. is funded by the Ramón y Cajal Program, Ministerio de Educación y Ciencia grant BFU 2005-04091, Fondo de Investigaciones Sanitarias, CIBERhed. D.P.X. is a fellow of the Association for International Cancer Research.
Author information
Authors and Affiliations
Contributions
M.H.T. was involved in the development of the protocol and prepared the manuscript. M.S.R. created the 6His-SUMO cell lines, devised and refined the purification technique and edited the manuscript. D.P.X. was involved in developing and refining the technique and edited the manuscript. R.T.H. is the principal investigator that supervised the work and edited the manuscript.
Corresponding author
Rights and permissions
About this article
Cite this article
Tatham, M., Rodriguez, M., Xirodimas, D. et al. Detection of protein SUMOylation in vivo. Nat Protoc 4, 1363–1371 (2009). https://doi.org/10.1038/nprot.2009.128
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2009.128
This article is cited by
-
ISG15 conjugation to proteins on nascent DNA mitigates DNA replication stress
Nature Communications (2022)
-
SUMOylation controls the binding of hexokinase 2 to mitochondria and protects against prostate cancer tumorigenesis
Nature Communications (2021)
-
Neuronal surface P antigen (NSPA) modulates postsynaptic NMDAR stability through ubiquitination of tyrosine phosphatase PTPMEG
BMC Biology (2020)
-
DDX39B interacts with the pattern recognition receptor pathway to inhibit NF-κB and sensitize to alkylating chemotherapy
BMC Biology (2020)
-
The HDAC inhibitor SAHA regulates CBX2 stability via a SUMO-triggered ubiquitin-mediated pathway in leukemia
Oncogene (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.