Abstract
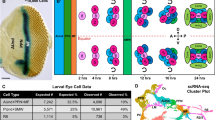
Cultured Drosophila melanogaster S2 and S2R+ cell lines have become important tools for uncovering fundamental aspects of cell biology as well as for gene discovery. Despite their utility, these cell lines are nonmotile and cannot build polarized structures or cell-cell contacts. Here we outline a previously isolated, but uncharacterized, Drosophila cell line named Dm-D17-c3 (or D17). These cells spread and migrate in culture, form cell-cell junctions and are susceptible to RNA interference (RNAi). Using this protocol, we describe how investigators, upon receiving cells from the Bloomington stock center, can culture cells and prepare the necessary reagents to plate and image migrating D17 cells; they can then be used to examine intracellular dynamics or observe loss-of-function RNAi phenotypes using an in vitro scratch or wound healing assay. From first thawing frozen ampules of D17 cells, investigators can expect to begin assaying RNAi phenotypes in D17 cells within roughly 2–3 weeks.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Schneider, I. Cell lines derived from late embryonic stages of Drosophila melanogaster. J. Embryol. Exp. Morphol. 27, 353–365 (1972).
Yanagawa, S., Lee, J.S. & Ishimoto, A. Identification and characterization of a novel line of Drosophila Schneider S2 cells that respond to wingless signaling. J. Biol. Chem. 273, 32353–32359 (1998).
Clemens, J.C. et al. Use of double-stranded RNA interference in Drosophila cell lines to dissect signal transduction pathways. Proc. Natl. Acad. Sci. USA 97, 6499–6503 (2000).
Rogers, S.L. Drosophila EB1 is important for proper assembly, dynamics, and positioning of the mitotic spindle. J. Cell. Biol. 158, 873–884 (2002).
Ramadan, N., Flockhart, I., Booker, M., Perrimon, N. & Mathey-Prevot, B. Design and implementation of high-throughput RNAi screens in cultured Drosophila cells. Nat. Protoc. 2, 2245–2264 (2007).
Adams, M.D. et al. The genome sequence of Drosophila melanogaster. Science 287, 2185–2195 (2000).
Rogers, G.C., Rusan, N.M., Peifer, M. & Rogers, S.L. A multicomponent assembly pathway contributes to the formation of acentrosomal microtubule arrays in interphase Drosophila cells. Mol. Biol. Cell. 19, 3163–3178 (2008).
Wheeler, S.R. et al. Neurexin IV and Wrapper interactions mediate Drosophila midline glial migration and axonal ensheathment. Development 136, 1147–1157 (2009).
Johnston, C.A., Hirono, K., Prehoda, K.E. & Doe, C.Q. Identification of an Aurora-A/PinsLINKER/Dlg spindle orientation pathway using induced cell polarity in S2 cells. Cell 138, 1150–1163 (2009).
Jiang, L., Rogers, S.L. & Crews, S.T. The Drosophila Dead end Arf-like3 GTPase controls vesicle trafficking during tracheal fusion cell morphogenesis. Dev. Biol. 311, 487–499 (2007).
Dorer, M.S., Kirton, D., Bader, J.S. & Isberg, R.R. RNA interference analysis of Legionella in Drosophila cells: exploitation of early secretory apparatus dynamics. PLoS. Pathog. 2, e34 (2006).
Wagner, E.J. et al. A genome-wide RNA interference screen reveals that variant histones are necessary for replication-dependent histone pre-mRNA processing. Mol. Cell. 28, 692–699 (2007).
D'Ambrosio, M.V. & Vale, R.D. A whole genome RNAi screen of Drosophila S2 cell spreading performed using automated computational image analysis. J. Cell. Biol. 191, 471–478 (2010).
Ui, K., Ueda, R. & Miyake, T. Cell lines from imaginal discs of Drosophila melanogaster. In Vitro Cell. Dev. Biol. 23, 707–711 (1987).
Wood, W., Faria, C. & Jacinto, A. Distinct mechanisms regulate hemocyte chemotaxis during development and wound healing in Drosophila melanogaster. J. Cell. Biol. 173, 405–416 (2006).
Wood, W. & Jacinto, A. Drosophila melanogaster embryonic haemocytes: masters of multitasking. Nat. Rev. Mol. Cell. Biol. 8, 542–551 (2007).
Celniker, S.E. et al. Unlocking the secrets of the genome. Nature 459, 927–930 (2009).
Rehorn, K.P., Thelen, H., Michelson, A.M. & Reuter, R. A molecular aspect of hematopoiesis and endoderm development common to vertebrates and Drosophila. Development 122, 4023–4031 (1996).
Cherbas, L. et al. The transcriptional diversity of 25 Drosophila cell lines. Genome Res. 21, 301–314 (2011).
Fasano, L. et al. The gene teashirt is required for the development of Drosophila embryonic trunk segments and encodes a protein with widely spaced zinc finger motifs. Cell 64, 63–79 (1991).
Rogers, S.L. & Rogers, G.C. Culture of Drosophila S2 cells and their use for RNAi-mediated loss-of-function studies and immunofluorescence microscopy. Nat. Protoc. 3, 606–611 (2008).
Zhang, D. et al. Drosophila katanin is a microtubule depolymerase that regulates cortical-microtubule plus-end interactions and cell migration. Nat. Cell. Biol. 13, 361–370 (2011).
Drechsel, D.N., Hyman, A.A., Hall, A. & Glotzer, M. A requirement for Rho and Cdc42 during cytokinesis in Xenopus embryos. Curr. Biol. 7, 12–23 (1997).
Paladi, M. & Tepass, U. Function of Rho GTPases in embryonic blood cell migration in Drosophila. J. Cell. Sci. 117, 6313–6326 (2004).
Stramer, B. Live imaging of wound inflammation in Drosophila embryos reveals key roles for small GTPases during in vivo cell migration. J. Cell. Biol. 168, 567–573 (2005).
Fessler, J.H., Nelson, R.E. & Fessler, L.I. Preparation of extracellular matrix. Methods Cell Biol. 44, 303–328 (1994).
Liang, C., Park, A.Y. & Guan, J. In vitro scratch assay: a convenient and inexpensive method for analysis of cell migration in vitro. Nat. Protoc. 2, 329–333 (2007).
Yarrow, J.C., Perlman, Z.E., Westwood, N.J. & Mitchison, T.J. A high-throughput cell migration assay using scratch wound healing, a comparison of image-based readout methods. BMC Biotechnol. 4, 21 (2004).
Vitorino, P. & Meyer, T. Modular control of endothelial sheet migration. Genes Dev. 22, 3268–3281 (2008).
Acknowledgements
We thank members of the Rogers lab for helpful discussions regarding the application of this protocol. We would also like to acknowledge M. Peifer for the generous gift of Canoe antibody and the DGRC for cell lines and helpful protocols.
Author information
Authors and Affiliations
Contributions
J.D.C. performed the protocol under the supervision of S.L.R. J.D.C. and S.L.R. wrote the protocol manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Video 1. D17 wound migration.
D17 cells treated with control dsRNA and imaged in intervals of 10 minutes over 16 hours. (MOV 8333 kb)
Supplementary Video 2. D17 single cell migration.
D17 cell transiently transfected with pMT GFP-Orbit and imaged every 3 seconds using spinning disc confocal microscopy. GFP-Orbit is a Drosophila microtubule plus end protein that labels the growing ends of microtubules in cells. (MOV 1660 kb)
Rights and permissions
About this article
Cite this article
Currie, J., Rogers, S. Using the Drosophila melanogaster D17-c3 cell culture system to study cell motility. Nat Protoc 6, 1632–1641 (2011). https://doi.org/10.1038/nprot.2011.397
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2011.397
This article is cited by
-
Drosophila ML-DmD17-c3 cells respond robustly to Dpp and exhibit complex transcriptional feedback on BMP signaling components
BMC Developmental Biology (2019)
-
Network-based prediction of polygenic disease genes involved in cell motility
BMC Bioinformatics (2019)
-
Comparative transcriptomic analysis of human and Drosophila extracellular vesicles
Scientific Reports (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.