Abstract
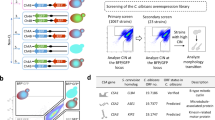
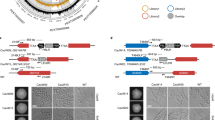
The recent discovery of haploids in Candida albicans and the construction of tool strains carrying multiple auxotrophic markers have enabled, for the first time, performing one-step gene deletions in this fungal human pathogen. This breakthrough promises to greatly facilitate the molecular and genetic study of C. albicans biology and pathogenicity. However, the construction of gene-deletion mutants in C. albicans haploids involves many technical difficulties, particularly low transformation efficiency and autodiploidization. Here we describe a highly effective protocol for designing and performing one-step gene deletion in C. albicans haploids, which takes ∼11 d to complete (not including plasmid construction, which may take ∼2 weeks). A gene deletion cassette is constructed on a plasmid and subsequently released for transformation by lithium acetate incubation or electroporation. Desired gene-deletion mutants are identified and their ploidy is assessed simultaneously by colony PCR before final confirmation by flow cytometry.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Brown, G.D. et al. Hidden killers: human fungal infections. Sci. Transl. Med. 4, 165rv113 (2012).
Homann, O.R., Dea, J., Noble, S.M. & Johnson, A.D. A phenotypic profile of the Candida albicans regulatory network. PLoS Genet. 5, e1000783 (2009).
Noble, S.M., French, S., Kohn, L.A., Chen, V. & Johnson, A.D. Systematic screens of a Candida albicans homozygous deletion library decouple morphogenetic switching and pathogenicity. Nat. Genet. 42, 590–598 (2010).
Blankenship, J.R., Fanning, S., Hamaker, J.J. & Mitchell, A.P. An extensive circuitry for cell wall regulation in Candida albicans. PLoS Pathog. 6, e1000752 (2010).
Ryan, O. et al. Global gene deletion analysis exploring yeast filamentous growth. Science 337, 1353–1356 (2012).
Noble, S.M. & Johnson, A.D. Strains and strategies for large-scale gene deletion studies of the diploid human fungal pathogen Candida albicans. Eukaryot. Cell 4, 298–309 (2005).
Fonzi, W.A. & Irwin, M.Y. Isogenic strain construction and gene mapping in Candida albicans. Genetics 134, 717–728 (1993).
Berman, J. & Sudbery, P.E. Candida albicans: a molecular revolution built on lessons from budding yeast. Nat. Rev. Genet. 3, 918–930 (2002).
De Backer, M.D., Magee, P.T. & Pla, J. Recent developments in molecular genetics of Candida albicans. Annu. Rev. Microbiol. 54, 463–498 (2000).
Wilson, R.B., Davis, D. & Mitchell, A.P. Rapid hypothesis testing with Candida albicans through gene disruption with short homology regions. J. Bacteriol. 181, 1868–1874 (1999).
Morschhauser, J., Michel, S. & Staib, P. Sequential gene disruption in Candida albicans by FLP-mediated site-specific recombination. Mol. Microbiol. 32, 547–556 (1999).
Alani, E., Cao, L. & Kleckner, N. A method for gene disruption that allows repeated use of URA3 selection in the construction of multiply disrupted yeast strains. Genetics 116, 541–545 (1987).
Wilson, R.B., Davis, D., Enloe, B.M. & Mitchell, A.P. A recyclable Candida albicans URA3 cassette for PCR product-directed gene disruptions. Yeast 16, 65–70 (2000).
Gola, S., Martin, R., Walther, A., Dunkler, A. & Wendland, J. New modules for PCR-based gene targeting in Candida albicans: rapid and efficient gene targeting using 100 bp of flanking homology region. Yeast 20, 1339–1347 (2003).
Walther, A. & Wendland, J. PCR-based gene targeting in Candida albicans. Nat. Protoc. 3, 1414–1421 (2008).
Gietz, R.D., Schiestl, R.H., Willems, A.R. & Woods, R.A. Studies on the transformation of intact yeast cells by the LiAc/SS-DNA/PEG procedure. Yeast 11, 355–360 (1995).
Walther, A. & Wendland, J. An improved transformation protocol for the human fungal pathogen Candida albicans. Curr. Genet. 42, 339–343 (2003).
Gietz, R.D. & Schiestl, R.H. High-efficiency yeast transformation using the LiAc/SS carrier DNA/PEG method. Nat. Protoc. 2, 31–34 (2007).
De Backer, M.D. et al. Transformation of Candida albicans by electroporation. Yeast 15, 1609–1618 (1999).
Thompson, J.R., Register, E., Curotto, J., Kurtz, M. & Kelly, R. An improved protocol for the preparation of yeast cells for transformation by electroporation. Yeast 14, 565–571 (1998).
Hickman, M.A. et al. The 'obligate diploid' Candida albicans forms mating-competent haploids. Nature 494, 55–59 (2013).
Gomez-Raja, J., Andaluz, E., Magee, B., Calderone, R. & Larriba, G. A single SNP, G929T (Gly310Val), determines the presence of a functional and a non-functional allele of HIS4 in Candida albicans SC5314: detection of the non-functional allele in laboratory strains. Fungal Genet. Biol. 45, 527–541 (2008).
Chibana, H., Uno, J., Cho, T. & Mikami, Y. Mutation in IRO1 tightly linked with URA3 gene reduces virulence of Candida albicans. Microbiol. Immunol. 49, 937–939 (2005).
Brand, A., MacCallum, D.M., Brown, A.J., Gow, N.A. & Odds, F.C. Ectopic expression of URA3 can influence the virulence phenotypes and proteome of Candida albicans but can be overcome by targeted reintegration of URA3 at the RPS10 locus. Eukaryot. Cell 3, 900–909 (2004).
Reuss, O., Vik, A., Kolter, R. & Morschhauser, J. The SAT1 flipper, an optimized tool for gene disruption in Candida albicans. Gene 341, 119–127 (2004).
Huxley, C., Green, E.D. & Dunham, I. Rapid assessment of S. cerevisiae mating type by PCR. Trends Genet. 6, 236 (1990).
Acknowledgements
We thank J. Berman and members of the Wang lab for critical reading of the manuscript. This work was funded by the Agency for Sciences, Technology and Research of Singapore.
Author information
Authors and Affiliations
Contributions
G.Z. designed and performed the experiments and wrote the first draft of the manuscript. F.Y.C. contributed to plasmid construction, and Y.-M.W. helped with flow cytometry analysis. Y.W. discussed and commented on the results at all stages and revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Zeng, G., Wang, YM., Chan, F. et al. One-step targeted gene deletion in Candida albicans haploids. Nat Protoc 9, 464–473 (2014). https://doi.org/10.1038/nprot.2014.029
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.029
This article is cited by
-
Rapid evolution of an adaptive multicellular morphology of Candida auris during systemic infection
Nature Communications (2024)
-
Candida albicans gains azole resistance by altering sphingolipid composition
Nature Communications (2018)
-
Sac7 and Rho1 regulate the white-to-opaque switching in Candida albicans
Scientific Reports (2018)
-
New “haploid biofilm model” unravels IRA2 as a novel regulator of Candida albicans biofilm formation
Scientific Reports (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.