Abstract
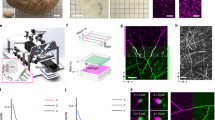
This protocol describes how in vivo–imaged dendrites and axons in adult mouse brains can subsequently be prepared and imaged with focused ion beam scanning electron microscopy (FIBSEM). The procedure starts after in vivo imaging with chemical fixation, followed by the identification of the fluorescent structures of interest. Their position is then highlighted in the fixed tissue by burning fiducial marks with the two-photon laser. Once the section has been stained and resin-embedded, a small block is trimmed close to these marks. Serially aligned EM images are acquired through this region, using FIBSEM, and the neurites of interest are then reconstructed semiautomatically by using the ilastik software (http://ilastik.org/). This reliable imaging and reconstruction technique avoids the use of specific labels to identify the structures of interest in the electron microscope, enabling optimal chemical fixation techniques to be applied and providing the best possible structural preservation for 3D analysis. The entire protocol takes ∼4 d.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Holtmaat, A. et al. Long-term, high-resolution imaging in the mouse neocortex through a chronic cranial window. Nat. Protoc. 4, 1128–1144 (2009).
Trachtenberg, J.T. et al. Long-term in vivo imaging of experience-dependent synaptic plasticity in adult cortex. Nature 420, 788–794 (2002).
Knott, G.W., Holtmaat, A., Wilbrecht, L., Welker, E. & Svoboda, K. Spine growth precedes synapse formation in the adult neocortex in vivo. Nat. Neurosci. 9, 1117–1124 (2006).
Shu, X. et al. A genetically encoded tag for correlated light and electron microscopy of intact cells, tissues, and organisms. PLoS Biol. 9, e1001041 (2011).
Knott, G.W., Holtmaat, A., Trachtenberg, J.T., Svoboda, K. & Welker, E. A protocol for preparing GFP-labeled neurons previously imaged in vivo and in slice preparations for light and electron microscopic analysis. Nat. Protoc. 4, 1145–1156 (2009).
Knott, G., Marchman, H., Wall, D. & Lich, B. Serial section scanning electron microscopy of adult brain tissue using focused ion beam milling. J. Neurosci. 28, 2959–2964 (2008).
Maco, B. et al. Correlative in vivo 2 photon and focused ion beam scanning electron microscopy of cortical neurons. PLoS ONE 8, e57405 (2013).
Bishop, D. et al. Near-infrared branding efficiently correlates light and electron microscopy. Nat. Methods 8, 568–570 (2011).
Kreshuk, A. et al. Automated detection and segmentation of synaptic contacts in nearly isotropic serial electron microscopy images. PLoS ONE 6, e24899 (2011).
Straehle, C.N., Köthe, U., Knott, G. & Hamprecht, F.A. Carving: scalable interactive segmentation of neural volume electron microscopy images. Med. Image Comput. Comput. Assist. Interv. 14, 653–660 (2011).
Denk, W. & Horstmann, H. Serial block-face scanning electron microscopy to reconstruct three-dimensional tissue nanostructure. PLoS Biol. 2, e329 (2004).
Helmstaedter, M. et al. Connectomic reconstruction of the inner plexiform layer in the mouse retina. Nature 500, 168–174 (2013).
Fiala, J.C. & Harris, K.M. Cylindrical diameters method for calibrating section thickness in serial electron microscopy. J. Microsc. 202, 468–472 (2001).
Cardona, A. et al. TrakEM2 software for neural circuit reconstruction. PLoS ONE 7, e38011 (2012).
Mostany, R. et al. Altered synaptic dynamics during normal brain aging. J. Neurosci. 33, 4094–4104 (2013).
Grillo, F.W. et al. Increased axonal bouton dynamics in the aging mouse cortex. Proc. Natl. Acad. Sci. USA 110, E1514–1523 (2013).
Sosinsky, G.E. et al. The combination of chemical fixation procedures with high pressure freezing and freeze substitution preserves highly labile tissue ultrastructure for electron tomography applications. J. Struct. Biol. 161, 359–371 (2008).
Acknowledgements
This work was supported by the Swiss National Foundation Synergia project grant CRF II313470/1 (G.W.K.) and project grant 31003A_135631 (A.H.).
Author information
Authors and Affiliations
Contributions
G.W.K. and A.H. conceived and designed experiments. A.H., G.W.K. and B.M. performed in vivo imaging, vibratome sectioning and laser mark branding. G.W.K., M.C. and B.M. performed electron microscopy and FIBSEM imaging. A.H., G.W.K., B.M., A.K. and F.A.H. analyzed the data and materials, and contributed reagents and analysis tools. G.W.K., B.M., A.H., M.C. and F.A.H. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Maco, B., Cantoni, M., Holtmaat, A. et al. Semiautomated correlative 3D electron microscopy of in vivo–imaged axons and dendrites. Nat Protoc 9, 1354–1366 (2014). https://doi.org/10.1038/nprot.2014.101
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.101
This article is cited by
-
A subpopulation of cortical VIP-expressing interneurons with highly dynamic spines
Communications Biology (2022)
-
Volume electron microscopy
Nature Reviews Methods Primers (2022)
-
Identifying long-range synaptic inputs using genetically encoded labels and volume electron microscopy
Scientific Reports (2022)
-
Along-axon diameter variation and axonal orientation dispersion revealed with 3D electron microscopy: implications for quantifying brain white matter microstructure with histology and diffusion MRI
Brain Structure and Function (2019)
-
Microglia remodel synapses by presynaptic trogocytosis and spine head filopodia induction
Nature Communications (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.