Key Points
-
Gene therapy for blood-cell diseases can be performed with retroviral vectors that insert into the genome of haematopoietic stem cells.
-
A recent trial of gene therapy for infants with X-linked severe combined immune deficiency (XSCID) successfully restored the immune systems of most subjects.
-
Two subjects developed T-cell leukaemia more than 2 years after gene therapy commenced. This cancer seems to be caused by retroviral-vector activation of a cellular oncogene at the site of integration, a process known as 'insertional oncogenesis'.
-
The complication of leukaemia has not occurred in any other clinical trial, nor in any large animal model that used retroviral vectors to modify haematopoietic stem cells. Leukaemia has been linked to vector integration in only one mouse study using this approach.
-
Multiple factors could have contributed to the development of leukaemia in the patients involved in this trial. These include the high level of engraftment and expansion of the genetically modified cells, unique properties of the haematopoietic stem and progenitor cells in bone marrow of X-linked SCID patients, the immune deficiency of the X-linked SCID patients and/or the transferred gene itself.
-
Further use of current gene-transfer methods for the treatment of SCID poses an ethical dilemma in the consideration of the complex benefits and risks.
-
It might be possible to develop retroviral vectors or other gene-therapy methods that are less likely to lead to insertional oncogenesis and still retain the therapeutic benefits. The use of tissue-specific, regulated transcription units should, in principle, diminish the risk of proto-oncogene transactivation.
Abstract
Recombinant viral vectors have allowed gene transfer to be developed as a promising approach to the treatment of genetic diseases. Recently, gene therapy of children with X-linked severe combined immune deficiency resulted in impressive levels of immune reconstitution — a triumph that was later overshadowed by the development of leukaemia in two patients. What were the causes of this cancer, and how can the therapeutic benefits of gene therapy be achieved while minimizing risk to the patient?
Similar content being viewed by others
Main
Gene transfer to human cells is being considered as an approach for treating numerous congenital and acquired disorders. This concept is based on the knowledge that specific cellular functions can be restored, modified or enhanced through intervention at the genetic level. An ever-expanding spectrum of gene-therapy applications is under investigation — from the correction of inborn errors in metabolism to vaccination, oncolysis, tissue repair and pain control. The first step towards implementing these therapeutic strategies is to efficiently introduce the therapeutic gene into somatic cells. The target cell can either be in culture, thereby allowing ex vivo gene transfer, or reside in organs or tumours, which would require in vivo gene transfer. Of the many target cell types under investigation, stem cells are unique for their ability to regenerate tissues or organs and eventually allow long-term therapeutic benefits. Haematopoietic stem cells (HSCs) are the best-studied adult stem cells. Gene-therapy experiments performed in these cells could therefore establish important models for the entire field of stem-cell-based gene therapy.
The potential to perform gene therapy for monogenic disorders that affect blood-cell production or function has been under study for more than 15 years1. As human genes were cloned and gene-transfer methods developed, the idea emerged of inserting a normal copy of a relevant human gene into the patient's own HSCs as an alternative to allogeneic bone-marrow transplantation. HSCs from bone marrow, cytokine-mobilized peripheral-blood stem cells or umbilical-cord blood could therefore be transduced with a therapeutic gene, providing a renewable source of genetically corrected cells. Expression of the normal gene product in the relevant blood cells and/or their progenitors (for example, T and B lymphocytes for immune deficiencies, stem cells for Fanconi's anaemia and erythrocytes for severe haemoglobinopathies) could alleviate the genetic deficiency. This general concept is applicable, in principle, to a number of congenital or acquired disorders (Table 1).
Recombinant viruses are, at present, the most effective vectors for transferring genes into primary cells. Viruses replicate through many mechanisms, but all involve the transfer of viral genetic information to a host cell. Viral gene products then participate in highly evolved and often complex interactions with host-cell functions to assure virus propagation. Molecular geneticists have exploited the ability of viruses to enter cells and integrate their genomes, and viral vectors are now commonly used to deliver genes to cells. This harnessing of viruses as gene-transfer vectors has required a detailed knowledge of the viral life cycle. Researchers have, however, succeeded in engineering viruses with disrupted replication functions that still retain the ability to enter cells, express vector genes, and in some cases persist in EPISOMAL or integrated forms.
Selected mammalian viruses have been engineered as viral vectors to treat human diseases (Table 2). These fall into two classes — those that persist in the host and are therefore best suited for long-term gene-transfer and gene-replacement therapies, or viruses that are solely lytic in nature. Lytic viruses provide powerful tools for 'hit-and-run' applications, such as vaccination or tumour destruction.
So far, promising integrating vectors (those that integrate into the host cell genome) include the oncoretroviruses and lentiviruses. Each of these has advantages and disadvantages with regards to safety and transduction efficiency. Retroviral integration can occur in a large number of different locations on different chromosomes, although it seems that preferred sites are within or near genes that are transcriptionally active2,3,4. Adeno-associated virus (AAV) vectors have also been created that can integrate into the host chromosomes in a non-directed manner5, in contrast to the wild-type virus, which integrates primarily into the q arm of chromosome 19 (Ref. 6). Directed integration is dependent on the function of the viral rep gene product, which is absent from therapeutic vectors. Vector integration occurs primarily in dividing cells, and is a rare event in non-dividing cells such as liver and muscle — the most frequently targeted tissue for therapeutic gene delivery. SV40 vectors can integrate into the DNA of both dividing and non-dividing cells7.
Non-integrating vectors that replicate independently of the host cell genome include the human herpesviruses. The latter have also been proposed as long-term gene-therapy vectors, because their natural biology involves life-long latency in different cell types. Whereas all or most of the gene sequences that encode viral proteins can be deleted from the relatively small and simple genomes of oncoretroviral, lentiviral and AAV vectors, the larger human herpesviruses are complex by comparison, requiring the activity of viral gene products for replication, and more sophisticated methods for engineering and vector production. The lytic viruses include adenoviruses and pox viruses. Each of these have a considerable history of use in vaccines, and, more recently, adenoviruses have been used extensively for transient gene expression in patients. As with all vectors, the key issues to address are the trade-offs between safety and efficacy. Whereas no vector is perfect, gene-transfer tools are becoming more highly refined and more amenable to use in treating human diseases.
Integrating vectors are the tools of choice to ensure gene delivery to the progeny of cells that undergo extensive clonal expansion, such as HSCs. The primary vector system that is currently used for gene transfer into HSCs is derived from mouse oncoretroviruses. These vectors are replication-defective retroviruses that encode a therapeutic cDNA placed under the transcriptional control of the retroviral long terminal repeat (LTR), or of an internal enhancer promoter of viral or mammalian origin8,9. One of the main limitations to this approach is that oncoretroviral vectors cannot integrate unless their host cell divides, and most HSCs are quiescent. Using cytokine stimulation to provoke cell proliferation, transduction efficiency in human HSCs has gradually improved over the years10,11,12. These vectors have therefore been used in gene therapy for selected diseases (Box 1).
A recent gene-therapy trial by Cavazzana-Calvo et al. tested the ability of a retroviral vector to treat patients who had a form of severe combined immunodeficiency (SCID), and resulted in restored immune function in most patients13,14. After several years, however, two patients developed a T-cell leukaemia that seemed to be linked to the activation of a known oncogene adjacent to the vector insertion sites. This severe adverse event raises serious safety concerns about the use of retroviral vectors for gene transfer in stem cells. It is therefore important to analyse the mechanism of cancer pathogenesis in these two patients, based on our knowledge of retroviral biology and our accumulated experience with retroviral-mediated gene transfer into HSCs.
Therapeutic success in gene therapy of SCID
Severe combined immune deficiency (SCID) is a group of genetic diseases that are caused by mutations in certain genes involved in immune-system development and function15. SCID usually causes infant mortality, due to overwhelming infections. SCID was the first human disease that was cured by bone-marrow transplantation, which involves transfer of normal HSCs from an HLA-matched sibling donor. Unfortunately, most infants with SCID lack an HLA-matched sibling bone-marrow donor, and transplantation from a parent or unrelated donor has a significantly lower success rate (the survival rate is near 90% in patients that received sibling bone marrow, versus 50–70% in patients who have received bone marrow from other types of donor)16.
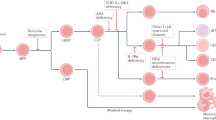
Another approach to treatment is to restore function through gene transfer to autologous HSCs (Fig. 1). Genetically corrected cells are likely to have a survival advantage, compared to the endogenous, genetically defective cells, and could therefore repopulate the lymphoid system. Nevertheless, initial clinical trials using retroviral vectors to treat SCID and other inherited immunodeficiency syndromes, including leukocyte adhesion defect (CD18 deficiency) and chronic granulomatous disease, were hampered by very low levels of gene transfer into HSCs. Recently, gene therapy for X-linked SCID (also referred to as XSCID or SCID-X1) and adenosine deaminase (ADA)-deficient SCID has been shown to result in effective immune restoration13,14,17.
a | In patients with severe combined immune deficiency (SCID), blocked lymphocyte production leads to immune deficiency. Blood cells are produced by the proliferation and differentiation of pluripotent haematopoietic stem cells (HSCs) through stages of lineage-restricted progenitors, the common lymphoid progenitor (CLP) and the common myeloid progenitor (CMP).These yield mature blood cells, including B cells (B), T cells (T), natural-killer cells (NK), granulocytes (Gran), monocytes (Mono), platelets (Plat) and erythrocytes (RBCs). Inherited mutations of genes that are needed for the production, survival or function of lymphocytes can cause severe combined immune deficiency (SCID), with absent or non-functional B, T and NK cells. b | The block to immune-cell production that occurs in patients with SCID leads to expansion of the progenitor-cell pool (CLPs). The number of CD34+ haematopoietic stem and progenitor cells obtained from the bone marrow of the youngest infant with X-linked SCID was relatively high for their size. This finding indicates that the pool of progenitor cells could be expanded in patients with X-linked SCID because of the absence of the γc-chain, which is required for cytokine signalling and differentiation of CLP. c | Gene correction of haematopoietic and progenitor cells leads to immune reconstitution. In gene therapy for X-linked SCID, a retroviral vector was used to transfer the normal human γc cDNA into HSCs that were isolated from the patient's bone marrow. Expression of γc restores the cytokine response and allows differentiation of CLP to B, T and NK cells, yielding immune reconstitution. The success in immune reconstitution in these patients might be due to the large pool of progenitor cells that were available for transduction, and to the high level of engraftment of the genetically altered cells, due to the absence of endogenous mature lymphocytes.
Cavazzana-Calvo et al.13 used a gene-therapy approach to treat ten infants that were born with X-linked SCID, which is caused by mutations in the γc gene (Box 2). In this study, they harvested bone marrow from the infants, isolated CD34+ STEM AND PROGENITOR CELLS, and transduced them with a retroviral vector that carries the normal human γc cDNA. This cDNA was under the transcriptional control of the retroviral LTR — a constitutively active, non-tissue-specific promoter. Significant levels of immune reconstitution occurred in all but one of the subjects, who was ill from a pre-existing infection and did not undergo HSC engraftment14. Protective T-cell immunity, characterized by proliferative responses to multiple antigens, was seen within 2–3 months after gene therapy. There was also evidence of at least partial restoration of B-cell and natural-killer (NK)-cell populations. NK-cell numbers have remained at subnormal levels in these patients, but have not been associated with any significant viral infections.
This timescale for immune reconstitution matches that observed in patients who received bone marrow from a normal sibling donor. It is also significantly shorter than the 6 months or more needed for adequate T-cell function to develop after patients receive bone marrow from a donor other than an HLA-matched sibling. The relatively rapid immune reconstitution that was achieved with this gene-therapy approach could therefore be beneficial to patients with SCID who have ongoing infections and lack an HLA-matched sibling donor. Bordignon et al. subsequently reported effective immune restoration in two subjects with SCID (due to ADA-deficiency) using similar methods of retroviral-mediated gene transfer to bone-marrow CD34+ cells17.
Leukaemia following gene therapy
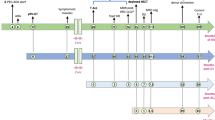
Unfortunately, almost 3 years after therapy was completed, T-cell leukaemia developed in two of the subjects that were involved in the trial of Cavazzana-Calvo et al.13. Significantly elevated white-blood-cell counts were detected during routine examinations, and further investigations showed the development of clonal T-cell proliferation, splenic enlargement and infiltration of the bone marrow — all classic clinical signs of leukaemia. In both cases, the leukaemia cells contained a single intact copy of the retroviral vector, which had integrated into chromosome 11, in or near the LMO2 gene. In the first case, the vector inserted into the first intron of the LMO2 gene, in a reverse orientation relative to that of LMO2 transcription. In the second case, the vector was located approximately 5 kb upstream from the LMO2 gene in the same orientation (Fig. 2a).
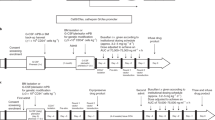
a | Integration sites of the therapeutic vector, which consists of the 5′ long terminal repeat (LTR), the γc cDNA and the 3′ LTR. Splice donor sites (SD) and splice acceptor sites (SA) are shown. In the first reported leukaemia patient (top), the vector integrated in the first intron, which lies between exon (Ex) 1 and 2, of the LMO2 gene. In the second patient (below), the integration site was upstream of the LMO2 promoter. [Alain Fischer, Marina Cavazzana-Calvo and Christoph von Kalle have shared publicly all of their scientific findings from their investigation of these cases and the information is adapted from presentations to the Biological Response Modifiers Advisory Committee of the US FDA, October 2002 and February 2003. See also RefS 14 and 18]. b | Inactivation of the 3′ LTR by U3 deletion (dLTR)54, which would eliminate LMO2 transcription from an upstream 3′ LTR. Vector elements include an internal promoter (P), the therapeutic transgene (TG). c | Abrogation of transcriptional read-through by transcriptional attenuation (TA) and polyadenylation (pA)55,56. d | Lineage-restricted, differentiation-stage-restricted transcription. In this example, an erythroid-specific vector59, carrying a β-globin gene promoter (p) and enhancer (e), as well as a locus control region (HS2, HS3, HS4), integrated into the first intron of LMO2 — this would not be expected to transactivate the LMO2 promoter in T cells. e | Transactivation of adjacent promoters might be prevented in vectors flanked by insulator elements (Ins) that have enhancer-blocking activity.
Although the leukaemias were detected 2.5 years after gene therapy, the specific T-cell clone from which the cancer arose could be detected, retrospectively, in samples that were collected as early as 1 year after therapy. The vector was detected using polymerase chain reaction (PCR) primers that were specific for the vector and cellular chromosomal DNA junctions. The relative prevalence of the vector in the T-cell population increased steadily over time. Presumably, the vector integrated into the LMO2 gene when therapy began, and the leukaemic clone eventually emerged through selective proliferation and/or survival. In the first subject, a t(6;13) translocation was detected in the leukaemia cells at the time of clinical presentation. In the second subject, IMMUNOSCOPE ANALYSIS of T-cell receptor patterns showed that at least three distinct transformed T-cell clones were present. This finding indicates that the vector integration and cellular transformation occurred in a pre-thymic stem or progenitor cell. These leukaemic cells also contained chromosomal anomalies, such as chromosome 11 trisomy. Both patients are now being treated with chemotherapy.
This serious adverse complication in a trial that initially produced such promising results has brought similar clinical trials to a halt, pending further investigation. The mechanism by which these two patients developed leukaemia is not clear. In more than a decade, leukaemias have not developed in any patients who are enrolled in other clinical trials that involved retroviral transduction of HSCs. Better insight into leukaemia pathogenesis is required to be able to assess and, ultimately, minimize the relative risk of this complication in patients who receive gene therapy for blood-cell diseases.
Possible mechanisms of leukaemogenesis
The presence of the integrated vector in the leukaemia cells of both patients raises the question of the role of the vector in the transformation or persistence of these clones. Its integration near a known oncogene is consistent with a direct role in leukaemogenesis. Furthermore, the integration into or near the same oncogene in two unrelated cases indicates that LMO2 transactivation is likely to be involved in transformation, and that LMO2 expression might be a powerful transforming event in the setting of gene therapy for X-linked SCID. However, the relatively long latency period before the onset of disease also indicates that LMO2 transactivation is not in itself sufficient to cause leukaemia — additional factors are required. This is consistent with current multistep models of leukaemogenesis, whereby initial mutations in progenitor cells increase cell proliferation and therefore the chance of accumulating secondary mutations19.
These two cases of leukaemia raise several important questions. Is retroviral integration causally linked to leukaemogenesis? Is integration near LMO2 either a frequent event or one that is uniquely prevalent/selected for in patients with X-linked SCID? Is the therapeutic gene itself, γc, a risk factor, when expressed from a recombinant retroviral vector under the control of the vector's LTR? Are there other genetic co-factors that cooperate with LMO2 to promote the development of leukaemia?
Retroviral insertion-induced oncogenesis. Retroviruses can cause tumour formation through various mechanisms20. Transactivation of an oncogene that is normally silenced is one mechanism that has been well-established in birds and mammals (Box 3). Retroviral insertion-induced mutagenesis is, in fact, an effective approach for uncovering novel oncogenes. The replication of endogenous retroviruses in selected strains of mice leads to frequent formation of tumours that are associated with multiple integration events in the same cell21,22,23. The analysis of retroviral integration sites (RISs) in these tumour cells is a powerful approach for oncogene discovery. Vector insertion-induced mutagenesis has also been used to activate oncogenes in cultured cell lines24.
However, there have been no other reports of gene-therapy vector insertion-induced oncogenesis in humans. Tumours have not been reported to develop in any of the more than 250 patients enrolled in more than 40 clinical trials that involved retroviral transfer of genes to HSCs25. In fact, in all the studies of mice that received retrovirally transduced HSCs, there is only a single report of tumour formation due to oncogene activation by an integrated retroviral vector26. In this study, Li et al. described the development of acute myeloid leukaemia (AML) in one of five mice that received retrovirally transduced bone-marrow cells26. The vector insertion site contributed to oncogenesis, because the vector integrated near the cellular proto-oncogene ecotropic viral integration site 1 ( Evi1 ). Evi1, which encodes a zinc-finger protein, was previously identified as a gene that is upregulated through insertional mutagenesis in neonatal mice that develop AML after infection with Moloney murine leukaemia virus27.
So, retroviral insertion-induced oncogenesis is a plausible mechanism by which cancer can develop in patients who have been treated with genetically modified HSCs. Extensive pre-clinical and clinical studies, however, gave Cavazzana-Calvo et al. no indication that cancer would develop in two out of the ten patients who were enrolled in their trial13,14.
Dysregulation of LMO2 expression. The most striking feature of these two leukaemia cases was the integration of the vector near LMO2 (Fig 2a). LMO2 is a widely expressed proto-oncogene that encodes a LIM DOMAIN PROTEIN that binds to transcription factors such as SCL/TAL1, GATA1 and GATA2 (Refs 28,29). It is expressed by haematopoietic progenitors and cells of the myeloid lineage, but not in post-thymic T cells. So, disruption of/insertion into the LMO2 locus could potentially affect haematopoiesis and lymphocyte development. LMO2 transcription is activated in childhood acute lymphoblastic leukaemia (ALL) cells as a consequence of a t(11;14)(p13;q11) translocation30. Transgenic mice that express LMO2/rhombotin-2 under the transcriptional control of the CD2 enhancer/promoter develop T-cell tumours as early as 5 months after birth31. In transgenic mice generated using the metallothionein promoter (active in most cell types) to direct expression of LMO2, the number of thymocytes at the most primitive stage of differentiation (CD4−CD8−) increased tenfold, with thymic tumours developing in 15% of the mice within 9–18 months after birth32.
LMO2 is therefore a leukaemogenic oncoprotein, but not one that causes the rapid onset of leukaemia in immunocompetent transgenic mice. Furthermore, LMO2 is not one of the most commonly activated oncogenes found in spontaneous human T-cell leukaemias33,34. Whether the LMO2 locus is a frequent RIS in human CD34+ cells — or at least in patients with X-linked SCID — or whether other powerful co-factors are required for leukaemogenesis in these patients, is unknown. Although these two possibilities are not mutually exclusive, it is essential to distinguish between them, because they have markedly different implications in terms of risk assessment.
Retovirally encoded γc-chain. The prolonged latency that preceded the development of overt leukaemia in patients indicated that the insertional activation of LMO2 expression either confers only a small increase in proliferation, which takes time to become apparent, or that secondary transforming events are required. It is therefore important to consider all possible co-factors that were common to both of the SCID patients who developed leukaemia and that could have contributed to the establishment or maintenance of a pre-leukaemic or leukaemic clone. One potential pathogenic factor to consider is the transgene product itself. There are many instances in which retroviral vectors have been shown to induce cellular transformation through the genes that they carry20. Genes that activate cell replication or inhibit apoptosis could potentially promote transformation.
At present, it is unknown whether the retrovirally encoded γc cDNA can act as a co-factor in transformation. Constitutive expression of γc under the transcriptional control of the LTR was expected to be safe, because γc is normally widely expressed in haematopoietic and lymphoid cells. Furthermore, other specific receptor proteins within the γc heteromeric complexes, along with receptor ligands, are required for signal transduction (Box 2). Nevertheless, the γc cDNA encodes a cytokine receptor chain that could provide a subtle proliferative or anti-apoptotic effect, and could promote transformation in combination with the insertional activation of LMO2.
There are indeed precedents whereby γc cytokine signalling has been shown to enhance leukaemogenesis. The IL-7 and IL-15 receptors have both been implicated in T-cell survival and homeostatic proliferation35,36,37. Transgenic mice that overexpress IL-7 are more susceptible to pro-B and pre-B cell tumours38, whereas those that overexpress IL-15 frequently develop T-NK lymphocytic leukaemias39. Interestingly, the incidence of T lymphoblastic lymphomas and large-B-cell lymphomas, induced by retroviral transfer of the NPM – ALK fusion gene product into bone-marrow cells, is increased in mice that transgenically express IL-9 (Refs 40,41). Furthermore, IL-15 has been shown to promote leukaemia-cell survival and/or proliferation42, whereas IL-7 can promote leukaemia-cell survival by downregulating p27 (Ref. 43). IL-7, which is required for thymocyte development and T-cell homeostasis35,36, is of particular interest because its levels are elevated in patients with SCID44. Hypothetically, increased signalling through the IL-7 receptor, which contains the γc-chain, could therefore have primed haematopoietic cells of SCID patients for transformation. The additional upregulation of LMO2 could compound this effect, leading the cells one step further along the road to transformation. Such a unique interaction between the specific therapeutic gene (γc) and a gene activated by insertional activation (LMO2) is one possible mechanism by which leukaemia could have developed in the two patients who were involved in the trial of Cavazzana-Calvo et al.13. This hypothesis could be tested experimentally in animal models.
Importantly, however, no serious adverse events were reported in any of the gene-complementation studies that were performed in γc-deficient mice25. Although mouse studies obviously differ from clinical studies, there has been no evidence so far that a γc transgene could have transforming potential.
In a study by Li et al., mice developed AML as a result of vector integration into the Evi1 locus. In this case, the retroviral vector encoded a truncated form of the human nerve growth-factor receptor p75 (ΔLNGFR)26. ΔLNGFR and other p75 mutants are cell-surface proteins that are used to identify transduced cells by fluorescence-activated cell sorting, or to sort transduced cells by immunomagnetic selection45. ΔLNGFR was originally claimed to be functionally disabled46, but Li et al. reported that the AML cells that were isolated from the transgenic mice proliferated in response to nerve growth factor26. They speculated that ΔLNGFR could therefore promote leukaemogenesis by cooperating with the activated Evi1 gene to transform myeloid cells. However, this point remains uncertain45, and it is noteworthy that the ΔLNGFR reporter construct has been used in several animal and clinical studies without any other report of leukaemia development47.
Immune deficits. It is also important to consider the context in which leukaemias emerged in the two patients. SCID is a state of immune deficiency that is characterized by deficits in T-, B- and NK-cell function. An overwhelming amount of experimental data indicates that tumours that would not thrive in normal mice can do so in immunodeficient mice. For example, the incidence of chemically-induced tumour formation is increased in Ifnγ−/−, Stat1−/− or Rag1−/− mice, compared with their immune-competent counterparts48. Furthermore, tumours that arise in immunodeficient strains of mice are easily rejected when transplanted into immunocompetent congenic mice49.
In humans, the association between Epstein–Barr-virus (EBV)-associated lymphoproliferative disease and immunosuppressed patients is well documented50,51. In the case of haematological tumours, immune surveillance by NK cells is especially important52. It is noteworthy, in this respect, that NK reconstitution of γc-deficient mice through gene therapy, using the same vector tested in the clinical trial, has been inconsistent53. In the patients involved in the trial of Cavazzana-Calvo et al., the NK-cell numbers were improved, but had not been returned to normal levels14. So, there is ample evidence to indicate that immunodeficient gene-therapy recipients could be, in general, at greater risk of developing tumours than those who are immune competent.
Other possible leukaemogenic co-factors. Although the absolute numbers of autologous CD34+ cells that the infants received during therapy were not exceptionally high, this number was more than tenfold higher, on a per kilogram basis (20–30 × 106/kg), than the dosages administered to adult patients. The ability of these patients to produce such large numbers of CD34+ cells indicates that these infants had expanded stem- or progenitor-cell pools. It is possible that infants with X-linked SCID accumulate more lymphoid progenitors in their bone marrow, due to the maturation block that is imparted by the inability to respond to cytokines that induce their differentiation. LMO2 is expressed in stem and progenitor cells, which might increase the frequency of insertion of vectors into this locus. So, these X-linked SCID infants could possess an expanded pool of progenitor cells that are susceptible to LMO2 disruption. Furthermore, it is noteworthy that the two subjects were the youngest enrolled in this clinical trial of SCID gene therapy. The hypothetical impact of age on susceptibility to insertional oncogenesis is unknown at present.
Reassessing the safety of integrating vectors
Understanding the risk of oncogenesis induction by vector integration requires further investigation into the mechanisms that underlie transformation, and the development of approaches to minimize the probability of leukaemia or tumour formation.
Analysis of vector integration sites. The first priority will be to analyse the patterns of integration-site selection for different integrating vectors (Table 1). In the case of oncoretroviruses, it has long been established that integration is biased towards DNAse I hypersensitive chromatin2,3. The same seems to be true for HIV-1 (Ref. 4). This bias results in preferential integration in gene-rich regions — particularly in or near actively transcribed genes. This preference is likely to be conserved in the replication-defective vectors that are derived from these viruses. However, there is at present no exhaustive compilation of all the potential RISs, nor is there an estimated frequency of their targeting.
Studies in mouse models of retroviral insertion-induced leukaemogenesis indicate that some genomic sites are more susceptible to vector integration than others. In a series of 1,299 RISs that were identified in mouse B-cell, T-cell and myeloid tumours, Suzuki et al.21 identified 152 loci at which gene insertion occurred more than once. Frequently disrupted genes encoded members of the RAS, NOTCH, JAK/STAT and NF-κB signalling pathways21. These data, however, represent the outcome of a highly selective assay that is likely to increase the representation of oncogenes in certain transformation complementation groups. Similarly, integration of the vector into LMO2 might also represent a highly selected event in cells from patients with X-linked SCID. Analyses of vector integration sites performed in tumour cells, therefore, do not alleviate the need to prospectively assess all possible integration sites in the absence of selective pressures.
This assessment requires methodologies that allow the rapid and accurate determination of RISs in small-tissue samples. Several PCR methods54,55 are now available to clone genomic regions that flank the provirus, and thanks to the availability of the human and mouse genome sequences, the integration site can be pinpointed based on the sequence of the region flanking the vector. A large database of human RISs is likely to develop over the next decade. The potential RISs and their targeting frequency will probably differ between vector types, as well as between target cell types.
Implications for preclinical toxicity studies. Onco-retroviruses are known to be leukaemogenic when allowed to replicate unchecked in neonatal mice, due to the high incidence of chromosomal integration. In fact, lymphomas developed in three of ten rhesus monkeys after transplantation of bone marrow that was exposed to replication-competent retroviral contaminants56. However, it was generally thought that vector insertion-induced oncogenesis would be extremely rare, if it even occurred at all, with replication-incompetent retroviral vectors, because these only integrate at the time of cell transduction. Therefore, one of the main elements in the certification of retroviral-vector preparations for use in clinical trials has been stringent assays to detect replication-competent retroviruses. Importantly, since 1989, no instance of unwanted retroviral-vector replication has been observed in any of the more than 100 clinical trials involving the use of oncoretroviral vectors1.
The recent awareness of the risk of vector insertion-induced oncogenesis will, however, affect the design of future toxicology studies. In the future, whenever a serious adverse event occurs in preclinical or clinical studies, the RIS will be one of the first factors investigated. This will require that valid assays for RIS analysis be established by gene-therapy investigators. Conversely, a prospective analysis of all possible RISs cannot be expected for every new gene-therapy proposal. As mentioned above, no single study can comprehensively address the issue of insertional mutagenesis. This will require the establishment of mouse and human registries, to which all investigators can contribute their data.
On the other hand, it could be required that each therapeutic gene to be delivered by an integrating vector be evaluated a priori, in transgenic mice and in animal disease models, to determine if it possesses the potential to promote insertional oncogenesis. Genes with an increased likelihood of causing oncogenesis would need to be considered in a risk/benefit ratio analysis for potential clinical applications.
Reducing the probability of oncogenesis
There are many approaches under development for reducing the risk of viral vector-induced oncogenesis. Within the confines of recombinant retroviruses, several modifications can be made to reduce the probability of oncogene activation. These include features to reduce transcriptional activation of neighbouring genes, to express a controlled suicide gene, and, in principle, to influence integration-site selection (Table 3).
Until now, all vectors tested in gene-therapy trials have been LTR-driven transcription units. The first modification that could increase vector safety would be to delete the U3 REGION OF THE VECTOR'S 3′ LTR57. This leads to self-inactivation (SIN) of the enhancer and promoter of both LTRs following integration. The therapeutic transgene could then be expressed from an internal enhancer/promoter. This SIN vector configuration reduces the number of active enhancer/promoter elements within the vector from two (the 5′ and 3′ LTR) to one (the internal enhancer/promoter), and abolishes downstream transcriptional read-through from the proviral 3′ LTR (Fig. 2b). Either of these effects could lower the risk of activation of a cellular gene by an integrated vector. The possibility still remains of transcription initiation from the internal enhancer/promoter. Transcriptional processivity through the inactivated 3′ LTR, which encodes a relatively weak polyadenylation signal58,59, could be reduced, in principle, by inclusion of a strong polyadenylation signal and eventually a transcriptional attenuator within the vector sequence (Fig. 2c). However, these measures to decrease the transcriptional activity of the LTR often lead to decreased viral titres.
Additional modifications to retroviral vectors that could improve safety are based on cell-specific transgene expression, in contrast to using non-specific enhancer/promoters that function in all cells. A model for such vectors is provided by the globin vectors, which are under investigation for the treatment of severe haemoglobinopathies60,61. These vectors, which are designed to express the β-chain of haemoglobin specifically in maturing proerythroblasts, require that many transcriptional control elements and chromatin-structure determinants be optimally combined61,62. The results obtained with such complex vectors show the feasibility of achieving tissue-specific transgene expression in the haematopoietic tissue62. If, for example, an erythroid-specific vector integrated into the LMO2 gene, these lineage-restricted promoter elements would not be expected to transactivate the LMO2 promoter in lymphocytes (Fig. 2d).
Other approaches to controlling transgene expression involve administration of specific small molecules that activate synthetic promoters. Artificial transcriptional activators that respond to drugs such as tetracycline or ecdysone have also shown promise in controlling transgene activation63. These could also be used to reduce the duration of transcriptional activity in the vector.
INSULATORS are DNA elements with enhancer-blocking activity that can also be engineered into therapeutic vectors to inhibit transactivation of adjacent genes in the genomic DNA (Fig. 2e). Such elements have been described in Drosophila, chicken and mammals, and are typically found in gene-boundary regions64,65. Unlike silencers, they are directional elements that only block enhancer-promoter interactions when they are physically interposed between the two. In oncoretroviral vectors, insulators have been shown to attenuate position effects66,67. Initially used to shelter the vector from the flanking chromatin, these elements could be even more useful in protecting flanking genes from the effects of the vector. It will be important to test the efficacy of these elements in appropriate animal models.
A distinctly different approach to tackle the risk of insertional oncogenesis is to incorporate into the vector a safety function that could be activated in the case of tumour formation. This strategy, which involves the use of a suicide gene, has been proposed for the elimination of alloreactive T lymphocytes68. In this approach, vectors that encode the herpes simplex virus-1 thymidine kinase ( HSVtk ) are constitutively expressed in the transduced T cells. Treatment with the prodrug ganciclovir, which is phosphorylated by HSVtk, prevents DNA elongation and selectively eliminates the proliferating T cells. The functional reliability, transcriptional regulation and immunogenicity of existing suicide systems, such as HSVtk, cytosine deaminase and FAS fusion proteins, remains to be established45.
Certainly, it would be ideal to develop transduction methods that allow transgene insertion into specific and safe chromosome sites, or that can correct mutations within endogenous loci. Targeted gene insertion can be implemented in certain experimental settings, but it remains a significant challenge to establish procedures that are sufficiently efficient and reliable for clinical application. Until targeted gene delivery becomes feasible in human cells, we will continue to depend on randomly integrating recombinant viruses to transfer therapeutic genes into cells that yield abundant progeny, such as stem cells. At present, the screening of integration sites before HSC transplantation poses a major challenge.
Interestingly, the random integration of transposable elements is not always biased towards actively transcribed genes. In yeast, certain retrotransposons show a preference for heterochromatin69. Some pre-integration complexes can therefore be targeted to chromosomal regions other than DNAse-I-sensitive sites. In a similar manner, manipulation of oncoretroviral and lentiviral pre-integration complexes could be used to influence integration site selection, although this approach has not been successful so far70.
Future directions
Should we postpone gene therapy for SCID patients until safer vectors are developed? To put this issue into perspective, tissue transplantation, chemotherapy and surgery have each, for many years, raised similar issues about risk to benefit assessment, and now this form of gene therapy brings us to a similar decision point. In the United States, this question has been grappled with by federal regulatory agencies, including the Food and Drug Administration (FDA) and the Recombinant DNA Advisory Committee (RAC) of the Office of Biotechnology Assessment, National Institutes of Health. The debate has hinged on the central question of the relative benefits of a life-saving treatment compared with the risks associated with the treatment itself.
The precise relative benefits and risks from gene therapy for patients with SCID, compared to non-autologous HSC transplantation, remain to be fully defined, as the total number of gene-therapy recipients remains low. The high rate of rapid immune reconstitution in patients who have received gene therapy could ultimately result in better outcomes than non-autologous HSC transplantation — even with the inherent risk of insertional oncogenesis. At the present time, proposals for clinical trials of gene therapy for SCID will be evaluated by the FDA and the RAC on a case-by-case basis, with the proven benefits and risks of gene therapy compared to those of the alternative treatments. Patients involved in gene-therapy trials will be carefully monitored, with long-term follow-up, as required by the FDA.
At present, there are no data to indicate that the risk of insertional oncogenesis is the same for all blood disorders. The available information, which is arguably incomplete, indicates that a concurrence of a constellation of cooperating factors is required to increase the risk of developing leukaemia to a frequency of two out of ten, as seen in the clinical trial of patients with X-linked SCID.
In the past two decades, the concept of gene therapy has ignited the imagination of medical scientists, engendering a great deal of research and speculation about new ways to treat human diseases. Indeed, the largely successful treatment of children with X-linked SCID strongly demonstrates the benefits of the gene-therapy approach, and represents a milestone in medicine. However, the inherent risks of insertional oncogenesis that are associated with the current methodologies have also been revealed. This dichotomy mandates that more effective and safer vector technology be developed, and this need rests squarely on the shoulders of vector biologists.
References
Kohn, D. B. Gene therapy for genetic and haematological disorders and immunodeficiency. J. Int. Med. 249, 379–390 (2001).
Vijaya, S. Acceptor sites for retroviral integrations map near DNase I-hypersensitive sites in chromatin. J. Virol. 60, 683–692 (1986).
Rohdewohld, H. et al. Retrovirus integration and chromatin structure: Moloney murine leukemia proviral integration sites map near DNase I-hypersensitive sites. J. Virol. 61, 336–343 (1987).
Schroder, A. R. et al. HIV-1 integration in the human genome favors active genes and local hotspots. Cell 110, 521–529 (2002). A paper that catalogued more than 500 independent genomic integration sites of HIV in human cells. A modest preference was seen for integration into gene-rich regions that were actively transcribed.
Nakai, H. et al. AAV serotype 2 vectors preferentially integrate into active genes in mice. Nature Genet. 1 Jun 2003 (doi:10.1038/ng1179).
Kotin, R. M. et al. Site specific integration by adeno-associated virus. Proc. Natl Acad. Sci. USA. 87, 2211–2215 (1990).
Strayer, D. S. et al. Durability of transgene expression and vector integration: recombinant SV40-derived gene therapy vectors. Mol. Ther. (in the press).
Miller, A. D. in Retroviruses (eds Coffin, F. M., Hughes, S. H. & Varmus, H. E.) 437–474 (Cold Spring Harbor Laboratory Press, New York, 1997).
Baum, C., Ostertag, W., Stocking, C. & von Laber, D. in Gene Therapy of Cancer (eds Lattime, E. C. & Gerson, S. L.) 3–30 (Academic Press, Missouri, USA, 2002).
Heim, D. A. & Dunbar, C. E. Hematopoietic stem cell gene therapy: towards clinically significant gene transfer efficiency. Immunol. Rev. 178, 29–38 (2000).
Halene, S. & Kohn, D. B. Gene therapy using hematopoietic stem cells: Sisyphus approaches the crest. Hum. Gene Ther. 11, 1259–1267 (2000).
Sadelain, M. et al. Issues in the manufacture and transplantation of genetically modified hematopoietic stem cells. Curr. Opin. Hematol. 7, 364–377 (2000).
Cavazzana-Calvo, M. et al. Gene therapy of human severe combined immunodeficiency (SCID)-X1 disease. Science 288, 669–672 (2000). The first report on the effects of gene therapy in two patients with X-linked SCID subjects. This paper marks the milestone of the first 'cure' of a disease by gene therapy.
Hacein-Bey-Abina, S. et al. Sustained correction of X-linked severe combined immunodeficiency by ex vivo gene therapy. N. Engl. J. Med. 346, 1185–1193 (2002). A second report with longer follow-up of the first two subjects reported in reference 7, with three additional subjects reported. Restoration of immunity was seen in four of five subjects. Two of these subjects subsequently developed T-cell leukaemia.
Weinberg, K. I. & Kohn, D. B. Gene therapy for congenital lymphoid immunodeficiency diseases. Semin. Hematol. 35, 354–366 (1998).
Antoine, C. S. et al. Long-term survival and transplantation of haemopoietic stem cells for immunodeficiencies: report of the European experience 1968–99. Lancet 361, 553–560 (2003).
Aiuti, A. et al. Correction of ADA-SCID by stem cell gene therapy combined with nonmyeloablative conditioning. Science 296, 2410–2413 (2002). The first report of immune restoration in two patients with ADA-deficient SCID. Unlike all previous gene-therapy studies for this form of SCID, these subjects were not treated with PEG-ADA enzyme-replacement therapy, which is believed to reduce the selective advantage of the gene-corrected cells. Additionally, these subjects received bone-marrow cytoreduction with a moderate dosage of the chemotherapeutic agent busulphan, which could promote engraftment of the gentically modified cells.
Marshall, E. Gene therapy. Second child in French trial is found to have leukemia. Science 299, 320 (2003).
Wu, X. & Pandolfi, P. P. Mouse models for multistep tumorigenesis. Trends Cell Biol. 11, S2–S9 (2001).
Rosenberg, N. et al. in Retroviruses (eds Coffin, A. M., Hughes, S. H. & Varmus, H. E.) 475–585 (Cold Spring Harbor Laboratory Press, New York, 1997).
Suzuki, T. et al. New genes involved in cancer identified by retroviral tagging. Nature Genet. 32, 166–174 (2002). In this study (and also references 22 and 23), researchers sequenced retroviral insertional sites in the mouse genome to examine cellular oncogenes that are involved in transformation. Multiple genes in specific signalling pathways were shown to promote transformation.
Mikkers, H., J. et al. High-throughput retroviral tagging to identify components of specific signaling pathways in cancer. Nature Genet. 32, 153–159 (2002).
Joosten, M. et al. Large-scale identification of novel potential disease loci in mouse leukemia applying an improved strategy for cloning common virus integration sites. Oncogene 21, 7247–7255 (2002).
Morishita, K. et al. Retroviral activation of a novel gene encoding a zinc finger protein in IL-3-dependent myeloid leukemia cell line. Cell 54, 831–840 (1988).
Kohn, D. B. et al. Report of the ad hoc sub-committee of the American Society of Gene Therapy on clinical trials conducted using retroviral-mediated gene transfer to hematopoietic stem cells. Mol. Ther. (in the press).
Li, Z. et al. Murine leukemia induced by retroviral gene marking. Science 296, 497 (2002). The first reported observation of insertional oncogenesis by a replication-incompetent retroviral vector. The retroviral vector was inserted near the Evi1 locus — a gene that is associated with acute myeloid leukaemia.
Nucifora, G. in Transcription Factors: Normal and Malignant Development of Blood Cells (eds Ravid, K. & Licht, J. D.) 393–404 (Wiley-Liss, New Jersey, USA 2001).
Osada, H. et al. Association of erythroid transcription factors: complexes involving the LIM protein RBTN2 and the zinc-finger protein GATA1. Proc. Natl Acad. Sci. USA 92, 9585–9589 (1995).
Rabbitts, T. H. et al. The effect of chromosomal translocations in acute leukemias: the LMO2 paradigm in transcription and development. Cancer Res. 59 (7 Suppl), 1794s–1798s (1999).
Royer-Pokora, B., Loos, U. & Ludwig, W. D. TTG-2, a new gene encoding a cysteine-rich protein with the LIM motif, is overexpressed in acute T-cell leukaemia with the t(11;14)(p13;q11). Oncogene 6, 1887–1893 (1991).
Larson, R. C. et al. The oncogenic LIM protein Rbtn2 causes thymic developmental aberrations that precede malignancy in transgenic mice. Oncogene 11, 853–862 (1995).
Neale, G. A, Rehg, J. E. & Goorha, R. M. Ectopic expression of rhombotin-2 causes selective expansion of CD4−CD8− lymphocytes in the thymus and T-cell tumors in transgeneic mice. Blood 86, 3060–3071 (1995).
Rabbits, T. H. Chromosomal translocation master genes, mouse models and experimental therapeutics. Oncogene 20, 5763–5777 (2001).
Ferrando, A. A. et al. Gene expression signatures define novel oncogenic pathways in T cell acute lymphoblastic leukemia. Cancer Cell 1, 75–87 (2002).
Puel, A., Ziegler, S. F., Buckley, R. H. & Leonard, W. J. Defective IL7R expression in T(−)B(+)NK(+) severe combined immunodeficiency. Nature Genet. 20, 394–397 (1998).
Schluns, K. S., Kieper, W. C., Jameson, S. C. & Lefrancois, L. Interleukin-7 mediates the homeostasis of naive and memory CD8 T cells in vivo. Nature Immunol. 1, 426–432 (2000).
Schluns, K. S. & Lefrancois, L. Cytokine control of memory T-cell development and survival. Nature Rev. Immunol. 3, 269–279 (2003).
Valenzona, H. O. Prelymphomatous B cell hyperplasia in the bone marrow of interleukin-7 transgenic mice: precursor B cell dynamics, microenvironmental organization and osteolysis. Exp. Hematol. 24, 1521–1529 (1996).
Fehniger, T. A. et al. Fatal leukemia in interleukin-15 transgenic mice. Blood Cells Mol. Dis. 27, 223–230 (2001).
Renauld, J. C. et al. Thymic lymphomas in interleukin 9 transgenic mice. Oncogene 9, 1327–1332 (1994).
Lange, K. et al. Overexpression of NPM-ALK induces different types of malignant lymphomas in IL-9 transgenic mice. Oncogene 22, 517–527 (2003).
Fehniger, T. A. & Caligiuri, M. A. Interleukin-15: biology and relevance to human disease. Blood 97, 14–32 (2001).
Barata, J. T., Cardoso, A. A., Nadler, L. M. & Boussiotos, V. A. Interleukin-7 promotes survival and cell cycle progression of T cell acute lymphoblastic leukemia cells by down-regulating the cyclin-dependent kinase inhibitor p27(kip1). Blood 98, 1524–1531 (2001).
Bolotin, E. et al. Serum levels of IL-7 in bone marrow transplant recipients: relationship to clinical characteristics and lymphocyte count. Bone Marrow Transplant. 23, 783–788 (1999).
Sadelain, M. & Riviere, I. Sturm und drang over suicidal lymphocytes. Mol. Ther. 5, 655–657 (2002).
Mavilio, F. et al. Peripheral blood lymphocytes as target cells of retroviral vector-mediated gene transfer. Blood 83, 1988–1997 (1994).
Bonini, C. et al. Safety of retroviral gene marking with a truncated NGF receptor. Nature Med. 9, 367–368 (2003).
Dunn, G. P. et al. Cancer immunoediting: from immunosurveillance to tumor escape. Nature Immunol. 3, 991–998 (2002).
Shankaran, V. et al. IFNgamma and lymphocytes prevent primary tumour development and shape tumour immunogenicity. Nature 410, 1107–1111 (2001).
O'Reilly, R. J. et al. Biology and adoptive cell therapy of EBV-associated lymphoproliferative disorders in recipients of marrow allografts. Immunol. Rev. 157, 195–216 (1997).
Okano, M. & Gross, T. G. A review of Epstein–Barr virus infection in patients with immunodeficiency disorders. Am. J. Med. Sci. 319, 392–396 (2000).
Smyth, M. J., Hayakawa, Y., Takeda, K. & Yagita, H. New aspects of natural-killer-cell surveillance and therapy of cancer. Nature Rev. Cancer 2, 850–861 (2002).
Soudais, C. et al. Stable and functional lymphoid of common cytokine reception chain deficient mice by retroviral-mediated gene transfer. Blood 95, 3071–3077 (2000).
Valk, P. J. A rapid RT-PCR based method to isolate complementary DNA fragments flanking retrovirus integration sites. Nucleic Acids Res. 25, 4419–4421 (1997).
Schmidt, M. et al. Detection and direct genomic sequencing of multiple rate unknown flanking DNA in highly complex samples. Hum. Gene Ther. 12, 743–749 (2001). Description of the LAM-PCR technique that was used by Christoph von Kalle and colleagues to uncover the LMO2 integration sites in the two leukaemia patients.
Donahue, R. E. et al. Helper virus induced t cell lymphoma in nonhuman primates after retroviral mediated gene transfer. J. Exp. Med. 176, 1125–1135 (1992).
Yu, S. F. et al. Self-inactivating retroviral vectors designed for transfer of whole genes into mammalian cells. Proc. Natl Acad. Sci. USA 83, 3194–3198 (1986).
Ismail, S. I., Rohll, J. B., Kingsman, S. M., Kingsman, A. J. & Uden, M. Use of intron-disrupted polyadenylation sites to enhance expression and safety of retroviral vectors. J. Virol. 75, 199–204 (2001).
Zaiss, A. K. et al. RNA 3′ readthrough of oncoretrovirus and lentivirus: implications for vector safety and efficacy. J. Virol. 76, 7209–7219 (2002).
Sadelain, M. Globin gene transfer for the treatment of severe hemoglobinopathies: a paradigm for stem cell-based gene therapy. J. Gene Med. 4, 113–121 (2002).
Rivella, S. & Sadelain, M. Therapeutic globin gene delivery using lentiviral vectors. Curr. Opin. Mol. Ther. 4, 505–514 (2002).
May, C. et al. Therapeutic haemoglobin synthesis in beta-thalassaemic mice expressing lentivirus-encoded human beta-globin. Nature 406, 82–86 (2000). Shows that lentiviral vector-mediated erythroid-specific expression of a globin gene can be used to treat haemoglobinopathies. This was an elusive goal for investigators who were trying the same approach using retroviral vectors, due to instability conferred by the necessary globin genomic regulatory sequences.
Baron, U. Tetracycline-controlled transcription in eukaryotes: novel transactivators with graded transactivation potential. Nucleic Acids Res. 25, 2723–2729 (1997).
Gerasimova, T. I. & Corces, V. G. Chromatin insulators and boundaries: effects on transcription and nuclear organization. Annu. Rev. Genet. 35, 193–208 (2001).
West, A. G., Gazner, M. & Felsenfold, G. Insulators: many functions, many mechanisms. Genes Dev. 16, 271–288 (2002).
Rivella, S. et al. The cHS4 insulator increases the probability of retroviral expression at random chromosomal integration sites. J. Virol. 74, 4679–4687 (2000).
Emery, D. W. et al. A chromatin insulator protects retrovirus vectors from chromosomal position effects. Proc. Natl Acad. Sci. USA 97, 9150–9155 (2000).
Cohen, J. L. et al. Suicide gene-mediated modulation of graft-versus-host disease. Leuk. Lymph. 34, 473–480 (1999).
Boeke, J. D. et al. Yeast retrotransposons: finding a nice quiet neighborhood. Cell 93, 1087–1089 (1998).
Bushman, F. Targeting retroviral integration? Mol. Ther. 6, 570–571 (2002).
Buckley, R. H. et al. Human severe combined immunodeficiency: genetic, phenotypic, and functional diversity in one hundred eight infants. J. Pediatr. 130, 378–387 (1997).
Fischer, A. et al. Naturally occurring primary deficiencies of the immune system. Annu. Rev. Immunol. 15, 93–124 (1997).
Leonard, W. J. The molecular basis of X-linked severe combined immunodeficiency: defective cytokine receptor signaling. Annu. Rev. Med. 47, 229–239 (1996).
Noguchi, M. et al. Interleukin-2 receptor gamma chain mutation results in X-linked severe combined immunodeficiency in humans. Cell 73, 147–157 (1993).
de Saint Basile, G. et al. Close linkage of the locus for X chromosome-linked severe combined immunodeficiency to polymorphic DNA markers in Xq11–q13. Proc. Natl Acad. Sci. USA 84, 7576–7579 (1987).
Puck, J. M., Nussbaum, R. L., Smead, D. L. & Conley, M. E. X-linked severe combined immunodeficiency: localization within the region Xq13.1-q21.1 by linkage and deletion analysis. Am. J. Hum. Genet. 44, 724–730 (1989).
Giri, J. et al. Elevated levels of shed type II IL-1 receptor in sepsis. Potential role for type II receptor in regulation of IL-1 responses. J. Immunol. 153, 5802–5809 (1994).
Kondo, M. et al. Sharing of the interleukin-2 (IL-2) receptor gamma chain between receptors for IL-2 and IL-4. Science 262, 1874–1877 (1993).
Russell, S. M. et al. Interleukin-2 receptor gamma chain: a functional component of the interleukin-4 receptor. Science 262, 1880–1883 (1993).
Noguchi, M. et al. Interleukin-2 receptor gamma chain: a functional component of the interleukin-7 receptor. Science 262, 1877–1880 (1993).
Kondo, M. et al. Functional participation of the IL-2 receptor gamma chain in IL-7 receptor complexes. Science 263, 1453–1454 (1994).
Russell, S. M. et al. Interaction of IL-2R beta and gamma c chains with Jak1 and Jak3: implications for XSCID and XCID. Science 266, 1042–1045 (1994).
Kimura, Y. et al. Sharing of the IL-2 receptor gamma chain with the functional IL-9 receptor complex. Int. Immunol. 7, 115–120 (1995).
Asao, H. et al. Cutting edge: the common gamma-chain is an indispensable subunit of the IL-21 receptor complex. J. Immunol. 167, 1–5 (2001).
Leonard, W. J. Cytokines and immunodeficiency diseases. Nature Rev. Immunol. 1, 200–208 (2001).
Kennedy, M. K. et al. Reversible defects in natural killer and memory CD8 T cell lineages in interleukin 15-deficient mice. J. Exp. Med. 191, 771–780 (2000).
Lodolce, J. P. et al. IL-15 receptor maintains lymphoid homeostasis by supporting lymphocyte homing and proliferation. Immunity 9, 669–676 (1998).
Macchi, P. et al. Mutations of Jak-3 gene in patients with autosomal severe combined immune deficiency (SCID). Nature 377, 65–68 (1995).
Russell, S. M. et al. Mutation of Jak3 in a patient with SCID: essential role of Jak3 in lymphoid development. Science 270, 797–800 (1995).
Coffin, J. M. in Fields Virology (eds Fields, B., Knipe, N., Howley, D. M.) 1767–1848 (Lippincott-Raven, USA, 1996).
Bishop, J. M. Molecular themes in oncogenesis. Cell 64, 235–248 (1991).
Haran–Ghera, N. Potential leukemic cells among bone marrow cells of young AKR/J mice. Proc. Natl Acad. Sci. USA, 77, 2923–2926 (1980).
Asjo, B. et al. Influence of genotype and the organ of origin on the subtype of T-cell in Moloney lymphomas induced by transfer of preleukemic cells from athymic and thymus-bearing mice. Cancer Res. 45, 1040–1045 (1985).
Fan, H., Brightman, B. K., Davis, B. R. & Li, Q. X. in Viruses that Affect the Immune System (ed. Fan, H.Y. et al.) 155–174 (American Society for Microbiology, Washington DC, 1991).
Lazo, P. A., Lee, J. S. & Tsichlis, P. N. Long-distance activation of the Myc protooncogene by provirus insertion in Mlvi-1 or Mlvi-4 in rat T-cell lymphomas. Proc. Natl Acad. Sci. USA 87, 170–173 (1990).
Bartholomew, C. & Ihle, J. N. Retroviral insertions 90 kilobases proximal to the Evi-1 myeloid transforming gene activate transcription from the normal promoter. Mol. Cell. Biol. 11, 1820–1828 (1991).
Corcoran, L. M., Adams, J. M., Dunn, A. R. & Cory, S. Murine T lymphomas in which the cellular myc oncogene has been activated by retroviral insertion. Cell 37, 113–122 (1984).
Selten, G., Cuypers, H. T. & Berns, A. Proviral activation of the putative oncogene Pim-1 in MuLV induced T-cell lymphomas. EMBO J. 4, 1793–1798 (1985).
Author information
Authors and Affiliations
Related links
Related links
DATABASES
Cancer.gov
GenBank
LocusLink
OMIM
FURTHER INFORMATION
The American Society of Gene Therapy
The Center for Biologics Evaluation and Research (CBER), Food and Drug Administration (FDA)
Childrens Hospital Los Angeles
Glossary
- EPISOME
-
A DNA element that persists in the nucleus of a cell, independently of the chromosomal DNA. Episomes might be the genomes of some viruses, including herpesvirus and Epstein–Barr virus, or artifical chromosomes.
- CD34+ STEM AND PROGENITOR CELLS
-
CD34 is a glycoprotein that is present on the surface of approximately 1% of human bone-marrow cells. CD34+ cells have properties of haematopoietic stem-cell and progenitor cells, in that they can proliferate and produce blood cells of various lineages. CD34+ cells can be isolated from the bone marrow or peripheral blood using commercially available immunoaffinity devices, be modified by gene transfer with retroviral vectors and be transplanted back into their donor.
- IMMUNOSCOPE ANALYSIS
-
A method to analyse T-cell diversity and specificity in a sample such as peripheral blood. Immunoscope analysis uses a polymerase-chain-reaction-based assay to examine the rearrangement patterns of T-cell-receptor gene families.
- LIM DOMAIN PROTEIN
-
A cysteine- and histidine-rich, zinc-coordinating domain that is composed of two tandemly repeated zinc fingers. LIM domains do not seem to bind DNA but instead seem to mediate protein–protein interactions.
- U3 REGION OF THE RETROVIRAL LONG TERMINAL REPEAT (LTR)
-
LTRs are the DNA sequences of approximately 600–800 base pairs in length that are present at both ends (5′ and 3′) of the retroviral-vector genome (provirus) — even after integration into the host-cell DNA. The U3 region of the LTR contains strong transcriptional enhancers and a promoter that drives expression of genes that are carried by most retroviral vectors. The U3 of the retroviral vector can activate expression of cellular genes that are adjacent to the site of retroviral integration.
- INSULATORS
-
Insulators are DNA sequences that insert between enhancer and promoter regions of DNA, blocking the ability of the enhancer to activate the promoter. Chains of genes along chromosomes can then act as isolated transcriptional units, and adjacent genes can be regulated independently. Insulators can be used to prevent a vector gene from being influenced by flanking genomic DNA sequences, or, conversely, to protect the flanking cellular genes from being influenced by the vector.
Rights and permissions
About this article
Cite this article
Kohn, D., Sadelain, M. & Glorioso, J. Occurrence of leukaemia following gene therapy of X-linked SCID. Nat Rev Cancer 3, 477–488 (2003). https://doi.org/10.1038/nrc1122
Issue Date:
DOI: https://doi.org/10.1038/nrc1122
This article is cited by
-
Long-term outcomes following CAR T cell therapy: what we know so far
Nature Reviews Clinical Oncology (2023)
-
Chimeric antigen receptor T cells therapy in solid tumors
Clinical and Translational Oncology (2023)
-
CRISPR–Cas9 can cause chromothripsis
Nature Genetics (2021)
-
Are chimeric antigen receptor T cells (CAR-T cells) the future in immunotherapy for autoimmune diseases?
Inflammation Research (2021)
-
Episomal minicircles persist in periods of transcriptional inactivity and can be transmitted through somatic cell nuclear transfer into bovine embryos
Molecular Biology Reports (2019)