Abstract
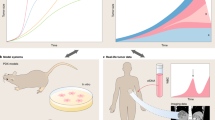
Cancer research attracts broad resources and scientists from many disciplines, and has produced some impressive advances in the treatment and understanding of this disease. However, a comprehensive mechanistic view of the cancer process remains elusive. To achieve this it seems clear that one must assemble a physically integrated team of interdisciplinary scientists that includes mathematicians, to develop mathematical models of tumorigenesis as a complex process. Examining these models and validating their findings by experimental and clinical observations seems to be one way to reconcile molecular reductionist with quantitative holistic approaches and produce an integrative mathematical oncology view of cancer progression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Anderson, A. R., Weaver, A. M., Cummings, P. T. & Quaranta, V. Tumor morphology and phenotypic evolution driven by selective pressure from the microenvironment. Cell 127, 905–915 (2006).
Gatenby, R. A., et al. Cellular adaptations to hypoxia and acidosis during somatic evolution of breast cancer. Br. J. Cancer 97, 646–653 (2007).
Araujo, R. P. & McElwain, D. L. A history of the study of solid tumour growth: the contribution of mathematical modelling. Bull. Math. Biol. 66, 1039–1091 (2004).
Kozusko, F. & Bourdeau, M. A unified model of sigmoid tumour growth based on cell proliferation and quiescence. Cell Prolif. 40, 824–834 (2007).
Anderson, A. R., Chaplain, M. A. J. & Rejniak, K. A. Single-Cell-Based Models in Biology and Medicine, (Birkhauser, Basel, 2007).
Weinberg, R. A. Using maths to tackle cancer. Nature 449, 978–981 (2007).
Janes, K. A., et al. A systems model of signaling identifies a molecular basis set for cytokine-induced apoptosis. Science 310, 1646–1653 (2005).
Kumar, N., Hendriks, B. S., Janes, K. A., de Graaf, D. & Lauffenburger, D. A. Applying computational modeling to drug discovery and development. Drug Discov. Today 11, 806–811 (2006).
Mogilner, A., Wollman, R. & Marshall, W. F. Quantitative modeling in cell biology: what is it good for? Dev. Cell 11, 279–287 (2006).
Anderson, A. R. A hybrid mathematical model of solid tumour invasion: the importance of cell adhesion. Math. Med. Biol. 22, 163–186 (2005).
Gerlee, P. & Anderson, A. R. An evolutionary hybrid cellular automaton model of solid tumour growth. J. Theor. Biol. 246, 583–603 (2007).
Mueller, M. M. & Fusenig, N. E. Friends or foes — bipolar effects of the tumour stroma in cancer. Nature Rev. Cancer 4, 839–849 (2004).
Wittekind, C., Compton, C. C., Greene, F. L. & Sobin, L. H. TNM residual tumor classification revisited. Cancer 94, 2511–2516 (2002).
Nowell, P. C. The clonal evolution of tumor cell populations. Science 194, 23–28 (1976).
Kerbel, R. S. Growth dominance of the metastatic cancer cell: cellular and molecular aspects. Adv. Cancer Res. 55, 87–132 (1990).
Vogelstein, B. & Kinzler, K. W. Cancer genes and the pathways they control. Nature Med. 10, 789–799 (2004).
Gould, S. J. & Lewontin, R. C. The spandrels of San Marco and the Panglossian paradigm: a critique of the adaptationist programme. Proc. R. Soc. Lond. B Biol. Sci. 205, 581–598 (1979).
Wade, M. & Wahl, G. M. c-Myc, genome instability, and tumorigenesis: the devil is in the details. Curr. Top. Microbiol Immunol. 302, 169–203 (2006).
Tomlinson, I. & Bodmer, W. Selection, the mutation rate and cancer: ensuring that the tail does not wag the dog. Nature Med. 5, 11–12 (1999).
Cairns, J. Mutation selection and the natural history of cancer. Nature 255, 197–200 (1975).
Kirschner, M. & Gerhart, J. Evolvability. Proc. Natl Acad. Sci. USA 95, 8420–8427 (1998).
Georgescu, W., et al. Model-controlled hydrodynamic focusing to generate multiple overlapping gradients of surface-immobilized proteins in microfluidic devices. Lab. Chip 21 Dec 2007 (doi: 10.b716203k).
Harpold, H. L., Alvord, E. C. Jr. & Swanson, K. R. The evolution of mathematical modeling of glioma proliferation and invasion. J. Neuropathol. Exp. Neurol. 66, 1–9 (2007).
Zaman, M. H., Kamm, R. D., Matsudaira, P. & Lauffenburger, D. A. Computational model for cell migration in three-dimensional matrices. Biophys. J. 89, 1389–1397 (2005).
Bild, A. H., et al. Oncogenic pathway signatures in human cancers as a guide to targeted therapies. Nature 439, 353–357 (2006).
Aldridge, B. B., Burke, J. M., Lauffenburger, D. A. & Sorger, P. K. Physicochemical modelling of cell signalling pathways. Nature Cell Biol. 8, 1195–1203 (2006).
Csikasz-Nagy, A., Battogtokh, D., Chen, K. C., Novak, B. & Tyson, J. J. Analysis of a generic model of eukaryotic cell-cycle regulation. Biophys. J. 90, 4361–4379 (2006).
Allen, G. E. in From Embryology to Evo–Devo: A History of Developmental Evolution (eds Laubichler, M. D. & Maienschein, J.) 123–167 (MIT, Cambridge, USA, 2007).
Bonner, J.T. The Evolution of Culture in Animals 5–9 (Princeton Univ., New Jersey, 1980).
Campbell, N. A. & Reece, J. B. Biology: Concepts and Connections 2–4 (Benjamin Cummings, Menlo Park, California, 2002).
Muller, G. B. in From Embryology to Evo–Devo: A History of Developmental Evolution (eds Laubichler, M. D. & Maienschein, J.) 499–524 (MIT, Cambridge, USA, 2007).
Anderson, A. R., Chaplain, M. A. J., Newman, E. L., Steele, R. J. & Thompson, A. M. Mathematical modelling of tumour invasion and metastasis. J. Theor. Biol. 2, 129–154 (2000).
Byrne, H. M. & Chaplain, M. A. J. Free boundary value problems associated with the growth and development of multicellular spheroids. Eur. J. App. Math. 8, 639–658 (1997).
Chaplain, M. A., Graziano, L. & Preziosi, L. Mathematical modelling of the loss of tissue compression responsiveness and its role in solid tumour development. Math. Med. Biol. 23, 197–229 (2006).
Enderling, H., Chaplain, M. A., Anderson, A. R. & Vaidya, J. S. A mathematical model of breast cancer development, local treatment and recurrence. J. Theor. Biol. 246, 245–259 (2007).
Ferreira, S. C., Jr., Martins, M. L. & Vilela, M. J. Reaction–diffusion model for the growth of avascular tumor. Phys. Rev. E 65, 021907 (2002).
Gatenby, R. A. & Gawlinski, E. T. A reaction–diffusion model of cancer invasion. Cancer Res. 56, 5745–5753 (1996).
Perumpanani, A. J., Sherratt, J. A., Norbury, J. & Byrne, H. M. Biological inferences from a mathematical model for malignant invasion. Invasion Metastasis 16, 209–221 (1996).
Sherratt, J. A. & Nowak, M. A. Oncogenes, anti-oncogenes and the immune response to cancer: a mathematical model. Proc. Biol. Sci. 248, 261–271 (1992).
Swanson, K. R., Bridge, C., Murray, J. D. & Alvord, E. C. Jr. Virtual and real brain tumors: using mathematical modeling to quantify glioma growth and invasion. J. Neurol. Sci. 216, 1–10 (2003).
Ward, J. P. & King, J. R. Mathematical modelling of avascular-tumour growth. II: Modelling growth saturation. IMA J. Math. Appl. Med. Biol. 16, 171–211 (1999).
Sachs, R. K., Chan, M., Hlatky, L. & Hahnfeldt, P. Modeling intercellular interactions during carcinogenesis. Radiat. Res. 164, 324–331 (2005).
Macklin, P. & Lowengrub, J. Evolving interfaces via gradients of geometry-dependent interior Poisson problems: application to tumor growth. J. Comp. Phys. 203, 191 (2005).
Zheng, X., Wise, S. M. & Cristini, V. Nonlinear simulation of tumor necrosis, neo-vascularization and tissue invasion via an adaptive finite-element/level-set method. Bull. Math. Biol. 67, 211–259 (2005).
Frieboes, H. B., et al. Computer simulation of glioma growth and morphology. Neuroimage 37, S59–S70 (2007).
Alarcon, T., Byrne, H. M. & Maini, P. K. A multiple scale model for tumor growth. Multiscale Modeling Simulation 3, 440–475 (2005).
Dormann, S. & Deutsch, A. Modeling of self-organized avascular tumor growth with a hybrid cellular automaton. In Silico Biol. 2, 393–406 (2002).
Duchting, W. Tumor growth simulation. Comput. Graph. 14, 505 (1990).
Kansal, A. R., Torquato, S., Harsh, G. I., Chiocca, E. A. & Deisboeck, T. S. Simulated brain tumor growth dynamics using a three-dimensional cellular automaton. J. Theor. Biol. 203, 367–382 (2000).
Patel, A. A., Gawlinski, E. T., Lemieux, S. K. & Gatenby, R. A. A cellular automaton model of early tumor growth and invasion. J. Theor. Biol. 213, 315–331 (2001).
Qi, A. S., Zheng, X., Du, C. Y. & An, B. S. A cellular automaton model of cancerous growth. J. Theor. Biol. 161, 1–12 (1993).
Smallbone, K., Gatenby, R. A., Gillies, R. J., Maini, P. K. & Gavaghan, D. J. Metabolic changes during carcinogenesis: potential impact on invasiveness. J. Theor. Biol. 244, 703–713 (2007).
Smolle, J. & Stettner, H. Computer simulation of tumour cell invasion by a stochastic growth model. J. Theor. Biol. 160, 63–72 (1993).
Jiang, Y., Pjesivac-Grbovic, J., Cantrell, C. & Freyer, J. P. A multiscale model for avascular tumor growth. Biophys. J. 89, 3884–3894 (2005).
Stott, E. L., Britton, N. F., Glazier, J. A. & Zajac, M. Stochastic simulation of benign avascular tumour growth using the Potts model. Math. Comput. Modelling 30, 183 (1999).
Turner, S. & Sherratt, J. A. Intercellular adhesion and cancer invasion: a discrete simulation using the extended Potts model. J. Theor. Biol. 216, 85–100 (2002).
Zhang, L., Athale, C. A. & Deisboeck, T. S. Development of a three-dimensional multiscale agent-based tumor model: simulating gene–protein interaction profiles, cell phenotypes and multicellular patterns in brain cancer. J. Theor. Biol. 244, 96–107 (2007).
Drasdo, D. & Hohme, S. Individual-based approaches to birth and death in avascular tumors. Math. Comput. Modelling 37, 1163 (2003).
Rejniak, K. A. An immersed boundary framework for modelling the growth of individual cells: an application to the early tumour development. J. Theor. Biol. 247, 186–204 (2007).H
Acknowledgements
The authors gratefully acknowledge the support of the Integrative Cancer Biology Program funded by the National Cancer Institute.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary information S1 (movie)
Tumour growth in a uniform ECM (cf. Figure 4a) All of the 2 dimensional simulations show the spatio–temporal evolution of all four variables in a 1cm2 domain: tumour cells (upper left), proteinase (upper right), nutrient (lower left) and ECM (lower right) over a period of approximately 3 months. Cell colouration relates to either the tumour cell-cell adhesion status (low=blue, medium=cyan, high=yellow, very high=orange) or if the cell is dead (dark brown). Colouration of the other variables is done using the hot colourmap (high=white, medium=red, low=black) to represent varying concentrations. All parameters used in the simulations are identical with the exception of the different microenvironments. (MOV 4942 kb)
Supplementary information S2 (movie)
Tumour growth in a grainy ECM (cf. Figure 4b) All of the 2 dimensional simulations show the spatio–temporal evolution of all four variables in a 1cm2 domain: tumour cells (upper left), proteinase (upper right), nutrient (lower left) and ECM (lower right) over a period of approximately 3 months. Cell colouration relates to either the tumour cell–cell adhesion status (low=blue, medium=cyan, high=yellow, very high=orange) or if the cell is dead (dark brown). Colouration of the other variables is done using the hot colourmap (high=white, medium=red, low=black) to represent varying concentrations. All parameters used in the simulations are identical with the exception of the different microenvironments. (MOV 5892 kb)
Supplementary information S3 (movie)
Tumour growth under low nutrient conditions (cf. Figure 4c) All of the 2 dimensional simulations show the spatio–temporal evolution of all four variables in a 1cm2 domain: tumour cells (upper left), proteinase (upper right), nutrient (lower left) and ECM (lower right) over a period of approximately 3 months. Cell colouration relates to either the tumour cell–cell adhesion status (low=blue, medium=cyan, high=yellow, very high=orange) or if the cell is dead (dark brown). Colouration of the other variables is done using the hot colourmap (high=white, medium=red, low=black) to represent varying concentrations. All parameters used in the simulations are identical with the exception of the different microenvironments. (MOV 827 kb)
Supplementary information S4 (movie)
A sequence of slices through the 3–dimensional tumour shown in Fig. 5. Colouration reflects different levels of cell density (green=low, red=high.)Each of the 3 dimensional simulations are simply different renderings of the same final tumour morphology after growth in a grainy ECM domain 0.5cm3 for approximately 1.5 months. (MOV 2070 kb)
Supplementary information S5 (movie)
A sequence of slices building up the 3–dimensional tumour shown in Fig. 5. Colouration is only used to try to distinguish different individual cells and has no other meaning.Each of the 3 dimensional simulations are simply different renderings of the same final tumour morphology after growth in a grainy ECM domain 0.5cm3 for approximately 1.5 months. (MOV 1699 kb)
Related links
Rights and permissions
About this article
Cite this article
Anderson, A., Quaranta, V. Integrative mathematical oncology. Nat Rev Cancer 8, 227–234 (2008). https://doi.org/10.1038/nrc2329
Issue Date:
DOI: https://doi.org/10.1038/nrc2329
This article is cited by
-
Comparing mechanism-based and machine learning models for predicting the effects of glucose accessibility on tumor cell proliferation
Scientific Reports (2023)
-
Fractional calculus in mathematical oncology
Scientific Reports (2023)
-
Investigation of phase-field models of tumor growth based on a reduced-order meshless Galerkin method
Engineering with Computers (2023)
-
Integrating a dynamic central metabolism model of cancer cells with a hybrid 3D multiscale model for vascular hepatocellular carcinoma growth
Scientific Reports (2022)
-
Can the Kuznetsov Model Replicate and Predict Cancer Growth in Humans?
Bulletin of Mathematical Biology (2022)