Abstract
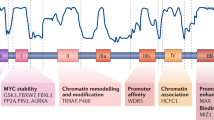
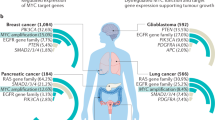
MYC regulates the transcription of thousands of genes required to coordinate a range of cellular processes, including those essential for proliferation, growth, differentiation, apoptosis and self-renewal. Recently, MYC has also been shown to serve as a direct regulator of ribosome biogenesis. MYC coordinates protein synthesis through the transcriptional control of RNA and protein components of ribosomes, and of gene products required for the processing of ribosomal RNA, the nuclear export of ribosomal subunits and the initiation of mRNA translation. We discuss how the modulation of ribosome biogenesis by MYC may be essential to its physiological functions as well as its pathological role in tumorigenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Pelengaris, S. & Khan, M. The many faces of c-MYC. Arch. Biochem. Biophys. 416, 129–136 (2003).
Eilers, M. & Eisenman, R. N. Myc's broad reach. Genes Dev. 22, 2755–2766 (2008).
Dang, C. V. et al. The c-Myc target gene network. Semin. Cancer Biol. 16, 253–264 (2006).
Grandori, C., Cowley, S. M., James, L. P. & Eisenman, R. N. The Myc/Max/Mad network and the transcriptional control of cell behavior. Annu. Rev. Cell Dev. Biol. 16, 653–699 (2000).
Adhikary, S. & Eilers, M. Transcriptional regulation and transformation by Myc proteins. Nature Rev. Mol. Cell Biol. 6, 635–645 (2005).
Dang, C. V. c-Myc target genes involved in cell growth, apoptosis, and metabolism. Mol. Cell. Biol. 19, 1–11 (1999).
Frank, S. R., Schroeder, M., Fernandez, P., Taubert, S. & Amati, B. Binding of c-Myc to chromatin mediates mitogen-induced acetylation of histone H4 and gene activation. Genes Dev. 15, 2069–2082 (2001).
Bouchard, C. et al. Regulation of cyclin D2 gene expression by the Myc/Max/Mad network: Myc-dependent TRRAP recruitment and histone acetylation at the cyclin D2 promoter. Genes Dev. 15, 2042–2047 (2001).
Knoepfler, P. S. et al. Myc influences global chromatin structure. EMBO J. 25, 2723–2734 (2006).
Cole, M. D. & Cowling, V. H. Transcription-independent functions of MYC: regulation of translation and DNA replication. Nature Rev. Mol. Cell Biol. 9, 810–815 (2008).
Dominguez-Sola, D. et al. Non-transcriptional control of DNA replication by c-Myc. Nature 448, 445–451 (2007).
Grandori, C. et al. c-Myc binds to human ribosomal DNA and stimulates transcription of rRNA genes by RNA polymerase I. Nature Cell Biol. 7, 311–318 (2005).
Grewal, S. S., Li, L., Orian, A., Eisenman, R. N. & Edgar, B. A. Myc-dependent regulation of ribosomal RNA synthesis during Drosophila development. Nature Cell Biol. 7, 295–302 (2005).
Dai, M. S. & Lu, H. Crosstalk between c-Myc and ribosome in ribosomal biogenesis and cancer. J. Cell Biochem. 105, 670–677 (2008).
Arabi, A. et al. c-Myc associates with ribosomal DNA and activates RNA polymerase I transcription. Nature Cell Biol. 7, 303–310 (2005).
Zeller, K. I. et al. Global mapping of c-Myc binding sites and target gene networks in human B cells. Proc. Natl Acad. Sci. USA 103, 17834–17839 (2006).
Fernandez, P. C. et al. Genomic targets of the human c-Myc protein. Genes Dev. 17, 1115–1129 (2003).
Orian, A. et al. Genomic binding by the Drosophila Myc, Max, Mad/Mnt transcription factor network. Genes Dev. 17, 1101–1114 (2003).
Blackwood, E. M. & Eisenman, R. N. Max: a helix-loop-helix zipper protein that forms a sequence-specific DNA-binding complex with Myc. Science 251, 1211–1217 (1991).
Blackwell, T. K., Kretzner, L., Blackwood, E. M., Eisenman, R. N. & Weintraub, H. Sequence-specific DNA binding by the c-Myc protein. Science 250, 1149–1151 (1990).
Papoulas, O., Williams, N. G. & Kingston, R. E. DNA binding activities of c-Myc purified from eukaryotic cells. J. Biol. Chem. 267, 10470–10480 (1992).
Frank, S. R. et al. MYC recruits the TIP60 histone acetyltransferase complex to chromatin. EMBO Rep. 4, 575–580 (2003).
McMahon, S. B., Wood, M. A. & Cole, M. D. The essential cofactor TRRAP recruits the histone acetyltransferase hGCN5 to c-Myc. Mol. Cell. Biol. 20, 556–562 (2000).
Ayer, D. E., Lawrence, Q. A. & Eisenman, R. N. Mad–Max transcriptional repression is mediated by ternary complex formation with mammalian homologs of yeast repressor Sin3. Cell 80, 767–776 (1995).
Schreiber-Agus, N. et al. An amino-terminal domain of Mxi1 mediates anti-Myc oncogenic activity and interacts with a homolog of the yeast transcriptional repressor SIN3. Cell 80, 777–786 (1995).
Seoane, J. et al. TGFβ influences Myc, Miz-1 and Smad to control the CDK inhibitor p15INK4b. Nature Cell Biol. 3, 400–408 (2001).
Staller, P. et al. Repression of p15INK4b expression by Myc through association with Miz-1. Nature Cell Biol. 3, 392–399 (2001).
Brenner, C. et al. Myc represses transcription through recruitment of DNA methyltransferase corepressor. EMBO J. 24, 336–346 (2005).
Wu, S. et al. Myc represses differentiation-induced p21CIP1 expression via Miz-1-dependent interaction with the p21 core promoter. Oncogene 22, 351–360 (2003).
Steiger, D., Furrer, M., Schwinkendorf, D. & Gallant, P. Max-independent functions of Myc in Drosophila melanogaster. Nature Genet. 40, 1084–1091 (2008).
Gomez-Roman, N., Grandori, C., Eisenman, R. N. & White, R. J. Direct activation of RNA polymerase III transcription by c-Myc. Nature 421, 290–294 (2003).
Knoepfler, P. S. Myc goes global: new tricks for an old oncogene. Cancer Res. 67, 5061–5063 (2007).
Wu, C. H. et al. Cellular senescence is an important mechanism of tumor regression upon c-Myc inactivation. Proc. Natl Acad. Sci. USA 104, 13028–13033 (2007).
Lindstrom, M. S. Emerging functions of ribosomal proteins in gene-specific transcription and translation. Biochem. Biophys. Res. Commun. 379, 167–170 (2009).
Rodnina, M. V. & Wintermeyer, W. Recent mechanistic insights into eukaryotic ribosomes. Curr. Opin. Cell Biol. 21, 435–443 (2009).
Schlosser, I. et al. A role for c-Myc in the regulation of ribosomal RNA processing. Nucleic Acids Res. 31, 6148–6156 (2003).
Tschochner, H. & Hurt, E. Pre-ribosomes on the road from the nucleolus to the cytoplasm. Trends Cell Biol. 13, 255–263 (2003).
Uechi, T., Tanaka, T. & Kenmochi, N. A complete map of the human ribosomal protein genes: assignment of 80 genes to the cytogenetic map and implications for human disorders. Genomics 72, 223–230 (2001).
Roeder, R. G. & Rutter, W. J. Specific nucleolar and nucleoplasmic RNA polymerases. Proc. Natl Acad. Sci. USA 65, 675–682 (1970).
Savkur, R. S. & Olson, M. O. Preferential cleavage in pre-ribosomal RNA byprotein B23 endoribonuclease. Nucleic Acids Res. 26, 4508–4515 (1998).
Bowman, L. H., Rabin, B. & Schlessinger, D. Multiple ribosomal RNA cleavage pathways in mammalian cells. Nucleic Acids Res. 9, 4951–4966 (1981).
Weinmann, R. & Roeder, R. G. Role of DNA-dependent RNA polymerase 3 in the transcription of the tRNA and 5S RNA genes. Proc. Natl Acad. Sci. USA 71, 1790–1794 (1974).
Gebauer, F. & Hentze, M. W. Molecular mechanisms of translational control. Nature Rev. Mol. Cell Biol. 5, 827–835 (2004).
Asano, K., Clayton, J., Shalev, A. & Hinnebusch, A. G. A multifactor complex of eukaryotic initiation factors, eIF1, eIF2, eIF3, eIF5, and initiator tRNA (Met) is an important translation initiation intermediate in vivo. Genes Dev. 14, 2534–2546 (2000).
Scheper, G. C. et al. Phosphorylation of eukaryotic initiation factor 4E markedly reduces its affinity for capped mRNA. J. Biol. Chem. 277, 3303–3309 (2002).
Hernandez, G. & Vazquez-Pianzola, P. Functional diversity of the eukaryotic translation initiation factors belonging to eIF4 families. Mech. Dev. 122, 865–876 (2005).
Kaiser, C. et al. Activation of cap-independent translation by variant eukaryotic initiation factor 4G in vivo. RNA 14, 2170–2182 (2008).
Qin, X. & Sarnow, P. Preferential translation of internal ribosome entry site-containing mRNAs during the mitotic cycle in mammalian cells. J. Biol. Chem. 279, 13721–13728 (2004).
Volarevic, S. et al. Proliferation, but not growth, blocked by conditional deletion of 40S ribosomal protein S6. Science 288, 2045–2047 (2000).
Sulic, S. et al. Inactivation of S6 ribosomal protein gene in T lymphocytes activates a p53-dependent checkpoint response. Genes Dev. 19, 3070–3082 (2005).
Panic, L. et al. Ribosomal protein S6 gene haploinsufficiency is associated with activation of a p53-dependent checkpoint during gastrulation. Mol. Cell. Biol. 26, 8880–8891 (2006).
Barkic, M. et al. The p53 tumor suppressor causes congenital malformations in Rpl24-deficient mice and promotes their survival. Mol. Cell. Biol. 29, 2489–2504 (2009).
Kirn-Safran, C. B. et al. Global growth deficiencies in mice lacking the ribosomal protein HIP/RPL29. Dev. Dyn. 236, 447–460 (2007).
Liu, J. J., Huang, B. H., Zhang, J., Carson, D. D. & Hooi, S. C. Repression of HIP/RPL29 expression induces differentiation in colon cancer cells. J. Cell. Physiol. 207, 287–292 (2006).
Wang, Y., Cheong, D., Chan, S. & Hooi, S. C. Heparin/heparan sulfate interacting protein gene expression is up-regulated in human colorectal carcinoma and correlated with differentiation status and metastasis. Cancer Res. 59, 2989–2994 (1999).
Anderson, S. J. et al. Ablation of ribosomal protein L22 selectively impairs αβ T cell development by activation of a p53-dependent checkpoint. Immunity 26, 759–772 (2007).
Ruggero, D. et al. The translation factor eIF-4E promotes tumor formation and cooperates with c-Myc in lymphomagenesis. Nature Med. 10, 484–6 (2004).
Ruggero, D. & Pandolfi, P. P. Does the ribosome translate cancer? Nature Rev. Cancer 3, 179–192 (2003).
Naora, H., Takai, I. & Adachi, M. Altered cellular responses by varying expression of a ribosomal protein gene: sequential coordination of enhancement and suppression of ribosomal protein S3a gene expression induces apoptosis. J. Cell Biol. 141, 741–753 (1998).
Kondoh, N. et al. Enhanced expression of S8, L12, L23a, L27 and L30 ribosomal protein mRNAs in human hepatocellular carcinoma. Anticancer Res. 21, 2429–2433 (2001).
Dai, M. S., Arnold, H., Sun, X. X., Sears, R. & Lu, H. Inhibition of c-Myc activity by ribosomal protein L11. EMBO J. 26, 3332–3345 (2007).
Lohrum, M. A., Ludwig, R. L., Kubbutat, M. H., Hanlon, M. & Vousden, K. H. Regulation of HDM2 activity by the ribosomal protein L11. Cancer Cell 3, 577–587 (2003).
Zhang, Y. et al. Ribosomal protein L11 negatively regulates oncoprotein MDM2 and mediates a p53-dependent ribosomal-stress checkpoint pathway. Mol. Cell. Biol. 23, 8902–8912 (2003).
Bhat, K. P., Itahana, K., Jin, A. & Zhang, Y. Essential role of ribosomal protein L11 in mediating growth inhibition-induced p53 activation. EMBO J. 23, 2402–2412 (2004).
Dai, M. S., Sears, R. & Lu, H. Feedback regulation of c-Myc by ribosomal protein L11. Cell Cycle 6, 2735–2741 (2007).
Schuhmacher, M. et al. Control of cell growth by c-Myc in the absence of cell division. Curr. Biol. 9, 1255–1258 (1999).
Iritani, B. M. & Eisenman, R. N. c-Myc enhances protein synthesis and cell size during B lymphocyte development. Proc. Natl Acad. Sci. USA 96, 13180–13185 (1999).
Johnston, L. A., Prober, D. A., Edgar, B. A., Eisenman, R. N. & Gallant, P. Drosophila myc regulates cellular growth during development. Cell 98, 779–790 (1999).
Saeboe-Larssen, S., Lyamouri, M., Merriam, J., Oksvold, M. P. & Lambertsson, A. Ribosomal protein insufficiency and the minute syndrome in Drosophila: a dose-response relationship. Genetics 148, 1215–1224 (1998).
Pierce, S. B. et al. dMyc is required for larval growth and endoreplication in Drosophila. Development 131, 2317–2327 (2004).
Schuhmacher, M. et al. The transcriptional program of a human B cell line in response to Myc. Nucleic Acids Res. 29, 397–406 (2001).
Kim, S., Li, Q., Dang, C. V. & Lee, L. A. Induction of ribosomal genes and hepatocyte hypertrophy by adenovirus-mediated expression of c-Myc in vivo. Proc. Natl Acad. Sci. USA 97, 11198–11202 (2000).
Beer, S. et al. Developmental context determines latency of MYC-induced tumorigenesis. PLoS Biol. 2, e332 (2004).
Trumpp, A. et al. c-Myc regulates mammalian body size by controlling cell number but not cell size. Nature 414, 768–773 (2001).
Mateyak, M. K., Obaya, A. J., Adachi, S. & Sedivy, J. M. Phenotypes of c-Myc-deficient rat fibroblasts isolated by targeted homologous recombination. Cell Growth Differ. 8, 1039–1048 (1997).
Boon, K. et al. N-myc enhances the expression of a large set of genes functioning in ribosome biogenesis and protein synthesis. EMBO J. 20, 1383–1393 (2001).
Zeller, K. I. et al. Characterization of nucleophosmin (B23) as a Myc target by scanning chromatin immunoprecipitation. J. Biol. Chem. 276, 48285–48291 (2001).
Friedrich, J. K., Panov, K. I., Cabart, P., Russell, J. & Zomerdijk, J. C. TBP–TAF complex SL1 directs RNA polymerase I pre-initiation complex formation and stabilizes upstream binding factor at the rDNA promoter. J. Biol. Chem. 280, 29551–29558 (2005).
Poortinga, G. et al. MAD1 and c-MYC regulate UBF and rDNA transcription during granulocyte differentiation. EMBO J. 23, 3325–3335 (2004).
Oskarsson, T. & Trumpp, A. The Myc trilogy: lord of RNA polymerases. Nature Cell Biol. 7, 215–217 (2005).
Shiue, C. N., Berkson, R. G. & Wright, A. P. c-Myc induces changes in higher order rDNA structure on stimulation of quiescent cells. Oncogene 28, 1833–1842 (2009).
Wu, C. H. et al. Combined analysis of murine and human microarrays and ChIP analysis reveals genes associated with the ability of MYC to maintain tumorigenesis. PLoS Genet. 4, e1000090 (2008).
Maggi, L. B. Jr et al. Nucleophosmin serves as a rate-limiting nuclear export chaperone for the mammalian ribosome. Mol. Cell. Biol. 28, 7050–7065 (2008).
De Benedetti, A. & Graff, J. R. eIF-4E expression and its role in malignancies and metastases. Oncogene 23, 3189–99 (2004).
Russell, J. & Zomerdijk, J. C. The RNA polymerase I transcription machinery. Biochem. Soc. Symp. 73, 203–216 (2006).
Brown, S. E. & Szyf, M. Dynamic epigenetic states of ribosomal RNA promoters during the cell cycle. Cell Cycle 7, 382–390 (2008).
Stefanovsky, V., Langlois, F., Gagnon-Kugler, T., Rothblum, L. I. & Moss, T. Growth factor signaling regulates elongation of RNA polymerase I transcription in mammals via UBF phosphorylation and r-chromatin remodeling. Mol. Cell 21, 629–639 (2006).
Mais, C., Wright, J. E., Prieto, J. L., Raggett, S. L. & McStay, B. UBF-binding site arrays form pseudo-NORs and sequester the RNA polymerase I transcription machinery. Genes Dev. 19, 50–64 (2005).
Chen, D., Belmont, A. S. & Huang, S. Upstream binding factor association induces large-scale chromatin decondensation. Proc. Natl Acad. Sci. USA 101, 15106–15111 (2004).
Panov, K. I., Friedrich, J. K., Russell, J. & Zomerdijk, J. C. UBF activates RNA polymerase I transcription by stimulating promoter escape. EMBO J. 25, 3310–3322 (2006).
Coller, H. A. et al. Expression analysis with oligonucleotide microarrays reveals that MYC regulates genes involved in growth, cell cycle, signaling, and adhesion. Proc. Natl Acad. Sci. USA 97, 3260–3265 (2000).
Thoma, C., Fraterman, S., Gentzel, M., Wilm, M. & Hentze, M. W. Translation initiation by the c-myc mRNA internal ribosome entry sequence and the poly(A) tail. RNA 14, 1579–1589 (2008).
Watson, J. D., Oster, S. K., Shago, M., Khosravi, F. & Penn, L. Z. Identifying genes regulated in a Myc-dependent manner. J. Biol. Chem. 277, 36921–36930 (2002).
Ginisty, H., Amalric, F. & Bouvet, P. Nucleolin functions in the first step of ribosomal RNA processing. EMBO J. 17, 1476–1486 (1998).
Herrera, J. E., Savkur, R. & Olson, M. O. The ribonuclease activity of nucleolar protein B23. Nucleic Acids Res. 23, 3974–3979 (1995).
Szebeni, A. & Olson, M. O. Nucleolar protein B23 has molecular chaperone activities. Protein Sci. 8, 905–912 (1999).
Okuwaki, M., Tsujimoto, M. & Nagata, K. The RNA binding activity of a ribosome biogenesis factor, nucleophosmin/B23, is modulated by phosphorylation with a cell cycle-dependent kinase and by association with its subtype. Mol. Biol. Cell 13, 2016–2030 (2002).
Li, Z., Boone, D. & Hann, S. R. Nucleophosmin interacts directly with c-Myc and controls c-Myc-induced hyperproliferation and transformation. Proc. Natl Acad. Sci. USA 105, 18794–18799 (2008).
Grisendi, S. et al. Role of nucleophosmin in embryonic development and tumorigenesis. Nature 437, 147–153 (2005).
Li, J., Sejas, D. P., Burma, S., Chen, D. J. & Pang, Q. Nucleophosmin suppresses oncogene-induced apoptosis and senescence and enhances oncogenic cooperation in cells with genomic instability. Carcinogenesis 28, 1163–1170 (2007).
Wanzel, M. et al. A ribosomal protein L23-nucleophosmin circuit coordinates Miz1 function with cell growth. Nature Cell Biol. 10, 1051–1061 (2008).
Jin, A., Itahana, K., O'Keefe, K. & Zhang, Y. Inhibition of HDM2 and activation of p53 by ribosomal protein L23. Mol. Cell. Biol. 24, 7669–7680 (2004).
Schmidt, E. V. The role of c-myc in regulation of translation initiation. Oncogene 23, 3217–3221 (2004).
Cole, M. D. & Cowling, V. H. Specific regulation of mRNA cap methylation by the c-Myc and E2F1 transcription factors. Oncogene 28, 1169–1175 (2009).
Shachaf, C. M. & Felsher, D. W. Tumor dormancy and MYC inactivation: pushing cancer to the brink of normalcy. Cancer Res. 65, 4471–4474 (2005).
Barna, M. et al. Suppression of Myc oncogenic activity by ribosomal protein haploinsufficiency. Nature 456, 971–975 (2008).
Draptchinskaia, N. et al. The gene encoding ribosomal protein S19 is mutated in Diamond–Blackfan anaemia. Nature Genet. 21, 169–175 (1999).
Watanabe, M., Zinn, A. R., Page, D. C. & Nishimoto, T. Functional equivalence of human X- and Y-encoded isoforms of ribosomal protein S4 consistent with a role in Turner syndrome. Nature Genet. 4, 268–271 (1993).
Kenmochi, N., Yoshihama, M., Higa, S. & Tanaka, T. The human ribosomal protein L6 gene in a critical region for Noonan syndrome. J. Hum. Genet. 45, 290–293 (2000).
Henry, J. L., Coggin, D. L. & King, C. R. High-level expression of the ribosomal protein L19 in human breast tumors that overexpress erbB-2. Cancer Res. 53, 1403–1408 (1993).
Vaarala, M. H. et al. Several genes encoding ribosomal proteins are over-expressed in prostate-cancer cell lines: confirmation of L7a and L37 over-expression in prostate-cancer tissue samples. Int. J. Cancer 78, 27–32 (1998).
Cheng, Q. et al. Identification of molecular markers for the early detection of human squamous cell carcinoma of the uterine cervix. Br. J. Cancer 86, 274–281 (2002).
Wang, Q. et al. Cloning and characterization of full-length human ribosomal protein L15 cDNA which was overexpressed in esophageal cancer. Gene 263, 205–209 (2001).
Kim, J. H. et al. Over-expression of the ribosomal protein L36a gene is associated with cellular proliferation in hepatocellular carcinoma. Hepatology 39, 129–138 (2004).
Wilson, A. et al. c-Myc controls the balance between hematopoietic stem cell self-renewal and differentiation. Genes Dev. 18, 2747–2763 (2004).
Pardal, R., Molofsky, A. V., He, S. & Morrison, S. J. Stem cell self-renewal and cancer cell proliferation are regulated by common networks that balance the activation of proto-oncogenes and tumor suppressors. Cold Spring Harb. Symp. Quant. Biol. 70, 177–185 (2005).
Felsher, D. W. & Bishop, J. M. Reversible tumorigenesis by MYC in hematopoietic lineages. Mol. Cell 4, 199–207 (1999).
Jain, M. et al. Sustained loss of a neoplastic phenotype by brief inactivation of MYC. Science 297, 102–104 (2002).
Shachaf, C. M. et al. MYC inactivation uncovers pluripotent differentiation and tumour dormancy in hepatocellular cancer. Nature 431, 1112–1117 (2004).
Li, Z. & Hann, S. R. The Myc–nucleophosmin–ARF network: a complex web unveiled. Cell Cycle 8, 2703–2707 (2009).
Weinstein, I. B. & Joe, A. Oncogene addiction. Cancer Res. 68, 3077–3080 (2008).
Jenuwein, T. & Allis, C. D. Translating the histone code. Science 293, 1074–1080 (2001).
Strahl, B. D. & Allis, C. D. The language of covalent histone modifications. Nature 403, 41–45 (2000).
Kobayashi, T. et al. Activation of the ribosomal protein L13 gene in human gastrointestinal cancer. Int. J. Mol. Med. 18, 161–170 (2006).
Pogue-Geile, K. et al. Ribosomal protein genes are overexpressed in colorectal cancer: isolation of a cDNA clone encoding the human S3 ribosomal protein. Mol. Cell. Biol. 11, 3842–3849 (1991).
Acknowledgements
We thank the Felsher laboratory for critical reading of the manuscript. Our work is supported by the National Institutes of Health, the Leukemia and Lymphoma Society and the Burroughs Wellcome Fund.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
DATABASES
OMIM
FURTHER INFORMATION
Rights and permissions
About this article
Cite this article
van Riggelen, J., Yetil, A. & Felsher, D. MYC as a regulator of ribosome biogenesis and protein synthesis. Nat Rev Cancer 10, 301–309 (2010). https://doi.org/10.1038/nrc2819
Issue Date:
DOI: https://doi.org/10.1038/nrc2819
This article is cited by
-
OST-01, a natural product from Baccharis coridifolia, targets c-Myc-dependent ribogenesis in acute myeloid leukemia
Leukemia (2024)
-
Identification of glucose-independent and reversible metabolic pathways associated with anti-proliferative effect of metformin in liver cancer cells
Metabolomics (2024)
-
The Plateau in Muscle Growth with Resistance Training: An Exploration of Possible Mechanisms
Sports Medicine (2024)
-
MYC overexpression and SMARCA4 loss cooperate to drive medulloblastoma formation in mice
Acta Neuropathologica Communications (2023)
-
Effects of sustained hyperprolactinemia in late gestation on the mammary parenchymal tissue transcriptome of gilts
BMC Genomics (2023)