Key Points
-
The retinoblastoma gene (RB) was the first tumour suppressor to be cloned, but the mechanism behind its role in tumour suppression remains unclear.
-
The retinoblastoma protein (RB) has been implicated in many cellular processes, such as regulation of the cell cycle, DNA-damage responses, DNA repair, DNA replication, protection against apoptosis, and differentiation, all of which could contribute to its function as a tumour suppressor.
-
RB is closely related to two genes in mice and humans (p107 and 130, which are not commonly mutated in tumours), which have been found to have partially redundant as well as opposing functions in genetic experiments in mice. Future studies are required to identify the similarities and differences between these proteins, which might explain the seemingly unique role of RB as a tumour suppressor.
-
As retinoblastomas do not arise as a consequence of Rb loss in mice, this model system is not ideal for studying this disease. However, this model system has provided information regarding the role of RB, p107 and p130 in development and tumorigenesis. Further experiments with conditional knockouts, as well as inter-crossing experiments, will provide a better insight into their roles in biological processes.
-
Biochemical experiments have provided information regarding the nature of various protein interactions with RB. Considering that there are more than 100 reported RB-binding proteins, much detailed structure/function mapping is required to clarify the biological relevance of all these interactions.
-
RB has also been associated with tumour growth, based on studies in which a constitutively active Rb allele gives rise to dysplasia in transgenic mice.
Abstract
Since its discovery, the retinoblastoma (RB) tumour-suppressor protein has been a focal point of cancer research. Accumulating evidence indicates a complex role for RB in cell proliferation, differentiation and survival. To further complicate matters, proteins that are related to RB have redundant as well as antagonistic functions. Recent studies of knockout mice and cells that lack one or more of these proteins have begun to clarify their various context-specific functions and the unique activity of this tumour suppressor.
Similar content being viewed by others
Main
It was first suggested more than a century ago that inherited abnormalities predispose some individuals to cancer (for review, see Ref. 1), but it is only in the past two decades, with the development of molecular cloning methodologies, that cancer-associated genes have been identified. These were found to include both oncogenes (dominant gain-of-function proteins) and tumour-suppressor genes (recessive loss-of-function proteins), and both classes of genes were discovered principally by virtue of their alteration in human cancers. Much of our current understanding of these genes has come from biochemical and functional studies in cultured cells; however, as described below, more recent clues to the mechanism by which the dysfunction of these genes contributes to human cancer has come from animal models.
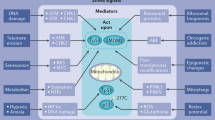
The first tumour-suppressor gene to be cloned was the retinoblastoma ( RB ) gene2,3,4,5. In familial cases of retinoblastoma, a germline mutation in the RB gene is inherited, and affected individuals develop retinal tumours in which the loss of the second RB allele is the rate-limiting step (reviewed in Ref. 6). The identification and subsequent cloning of the RB gene3,4 launched a new era in cancer genetics, and RB was subsequently found to be mutated in other human cancers, such as small-cell lung carcinoma and osteosarcoma. The RB gene product, RB, was identified as an important target of oncoproteins that are expressed by DNA tumour viruses, such as adenovirus E1A, SV40 T antigen and the E7 proteins of human papillomaviruses. These findings indicated that RB is a regulator of cell proliferation7,8,9,10 (Fig. 1). The discovery that RB overexpression caused cells to undergo arrest in the G1 phase of the cell cycle11,12,13, whereas cells that are deficient in RB show an accelerated G1 transition14,15,16, also provided support for the idea of RB as a cell-proliferation inhibitor.
The E2F transcription-factor family has been shown to regulate the expression of genes, products of which are involved in G1–S phase transition. The cell cycle is depicted at the bottom of the figure with its various phases (G0–G1–S–G/M). E2F-mediated transcription is negatively regulated by the RB family of proteins and their activity is, in turn, regulated by CDKs (CDK2, CDK4, and CDK6) and their inhibitors (p15, p16, p18, p19, p21 and p27). Molecules in this regulatory network (p16, cyclin D, CDK4/CDK6 and RB) (blue) have been found altered in a wide variety of tumours, with the exception of E2F. In addition to its ability to block S-phase progression, RB is believed to regulate a wide variety of processes, such as apoptosis, DNA replication, DNA repair, checkpoint control and differentiation (red). It is unclear whether these functions are all mediated through regulation of E2F or are dependent on one of the many (more than 100) factors with which RB can interact. Little is known about the impact the CDK regulatory network has on these alternative functions of RB.
The identification of viral oncoproteins that interact with and presumably interrupt the function of RB also raised the possibility that analogous cellular proteins could exist. Subsequent experimentation led to the identification of the E2F1 transcription factor as the first cellular target of RB17,18,19 (reviewed in Refs 20,21). Verification that E2F1 regulates cell proliferation came from studies in which it was shown that E2F1 overexpression can promote transition from the G1 phase to the S phase of the cell cycle21. Additional studies revealed that E2F1 is a member of a family of closely related proteins, several of which have been found to interact with RB22 (Fig. 2). Most current models propose that the role of RB in controlling cell-cycle progression is exerted through its repressive effects on gene expression mediated by the E2Fs21,23,24,25,26,27,28,29,31.
a | RB is believed to bind the transcription factor E2F and its associated subunit DP. RB represses E2F-mediated transcription by recruiting chromatin remodelling complexes to the promoter in resting cells. At the G1–S-phase transition, RB is thought to be phosphorylated by CDK2, CDK4 and CDK6. Hyperphosphorylated RB releases E2F, allowing it to activate transcription of its target genes. b | The mammalian genome encodes at least six different E2F proteins, listed as E2F1–6. The E2F transcription factors can be subdivided into three categories based on their transcriptional properties as well as their interaction with different pocket proteins. The first group consists of E2F1, E2F2 and E2F3, which are potent transcriptional activators that can also interact with RB. This presumably represses the activating potential of these three E2Fs. E2F4 and E2F5, which represent the second group, are fairly weak transcriptional activators that seem to more readily form complexes with p107 and p130 (although E2F4 will interact with all three pocket proteins). The third category, E2F6, does not associate with pocket proteins and seems to act as a transcriptional repressor22. The RB family of proteins (in most cases, it has only been tested for RB) has been shown to interact with chromatin remodelling complexes, such as the SWI/SNF complex, the histone deacetylases (HDACs), polycomb group (PcG) proteins as well as methylases. Recruitment of these complexes to E2F-binding sites is thought to mediate the repressive function of RB-family members32,33.
It was recently shown that the repressive action of RB in regulating gene expression occurs, in part, via the recruitment of chromatin-remodelling complexes to promoter regions. These complexes mediate chromatin condensation and subsequent inhibition of transcription32,33,34. The ability of RB to repress E2F-mediated transcription is regulated by the phosphorylation of RB by cyclin-dependent kinases (CDKs). This phosphorylation disrupts the association between E2Fs and RB35,36. RB is therefore linked directly to the intricate regulatory network that controls cell-cycle progression37 (Figs 1,2). Alterations in cell-cycle regulatory genes that encode proteins that participate in the regulation of RB function are also commonly observed in a broad spectrum of tumour types. Taken together, these findings indicate that deregulation of the normal pathway in which RB functions is a common and important event in tumorigenesis (Fig. 1).
Efforts to clearly establish the mechanism by which RB acts as a tumour suppressor have been complicated by various factors. For example, in addition to its role in cell-cycle control, RB has been implicated in regulating a wide variety of cellular processes, including DNA replication38,39,40, differentiation41 and apoptosis41,42 (Fig. 1). It is easy to imagine that a decrease in differentiation potential, in addition to an increase in proliferative rates, both of which are seen in RB-deficient cells, could contribute to tumorigenesis. However, in light of accumulating evidence indicating that inhibition of apoptosis contributes to tumorigenesis, it is more difficult to reconcile RB's role as a tumour suppressor, with reports that loss of RB leads to increased apoptosis in some circumstances. Although there is accumulating evidence that de-repression of E2F is important in both apoptosis and cell-cycle regulation43,44,45,46,47,48,49, it is unclear which aspects of RB's biological functions are achieved through its effects on E2F-mediated transcription.
Indeed, RB potentially interacts with more than 100 different cellular proteins50, and the functional relevance of most of these interactions has yet to be established. A further complication is that RB is one member of a family of related proteins that are collectively called the 'pocket proteins' (Fig. 3) and that seem to have both redundant and opposing functions, depending on the context (for review, see Refs 51,52). Finally, the fact that most tumour-associated RB mutations lead to the production of a truncated or unstable protein has made it difficult to map precisely the domains of RB that are most important for its tumour-suppressive function. In short, there are several significant obstacles that have been encountered in the pursuit of a clear understanding of the cellular function of RB.
a | RB, p107 and p130 make up the 'pocket' family of proteins. The largest region of homology between these proteins lies in a 'pocket' domain (regions A and B, separated by a spacer), which is required for their interaction with E2Fs and many other factors. Following the publication of the crystal structure of an RB molecule bound to a peptide from the human papillomavirus (HPV) E7 protein, a more defined mapping of specific interactions have been initiated (again mainly in the RB molecule)124,125,126,127,128,129,130,131 to determine the binding sites of histone deacetylase (HDAC), as well as the viral oncoproteins adenovirus E1A (E1A), SV40 T antigen (T Ag) and HPV E7 protein (E7). Regions within the pocket, as well as a domain in the carboxyl terminus of RB, have been shown to be important for E2F binding, but the specific contacts made between E2F and RB have not yet been mapped in detail. Furthermore, it is still unclear whether the requirements are the same for all the E2F-family members that bind RB (p107 and p130). Arrows represent the many regulatory phosphorylation sites that have been mapped in RB, and two acetylation sites (stars) have been mapped in the carboxyl terminus of the protein132. The p107 and p130 pocket domains are also shown (regions A and B, which are separated by a longer spacer than in Rb). The cyclin-binding site maps to the spacer that is unique to p107 and p130, as does a conserved domain that binds to viral oncoproteins (E1A, E7 and T Ag). Other protein interactions are not yet well defined. b | The pocket proteins are differentially expressed throughout the cell cycle. Whereas RB is expressed steadily throughout the cell cycle (blue), p130 (orange) expression is high in arrested cells, whereas p107 (red) expression peaks during the S phase of the cell cycle.
The pocket family of proteins
The RB protein is a member of a family of three closely related mammalian proteins that includes p107 and p130 (Fig. 3a). Together, these proteins are referred to as the 'pocket proteins' because their main sequence similarity resides in a domain (the pocket domain) that mediates interactions with the viral oncoproteins. Overexpression experiments have indicated functional similarities between these proteins in the regulation of the cell cycle. For example, when overexpressed, they can all cause arrest of some cell types in the G1 phase of the cell cycle, and they can all interact with and repress E2F-mediated gene transcription. In addition, they are all phosphorylated by CDKs (for review, see Refs 51,52).
Despite these similarities, there are significant differences between these proteins. For example, RB interacts predominantly with E2Fs 1, 2, 3 and 4, whereas p107 and p130 primarily interact with E2F4 and E2F5 (see Fig. 2b and, for review, see Refs 22,52). These associations also occur at distinct times during the cell cycle, in part due to expression patterns (Fig. 3b). Whereas RB interacts with E2Fs in both quiescent and cycling cells, p130 interacts with E2Fs primarily in quiescent cells22. However, p107 is predominantly associated with E2Fs in the S phase of the cell cycle, a phenomenon that is not well understood, particularly since overexpression of p107 results in G1 arrest in some cells53,54. In addition, p107 and p130 can recruit CDK2-containing complexes to promoters, and some experiments have indicated that p107 and p130 can act as inhibitors of CDK function55,56. Although p107 and p130 contain unique binding sites for CDK2-binding cyclins (Fig. 3), it is unclear which of the many proteins that bind to RB also interact with p107 and p130. It is also unclear whether there are other proteins that associate with p107 and p130 that do not bind to RB52.
Although numerous studies of the RB family of proteins have led to an extensive understanding of the biochemical properties of these proteins, the ability to disrupt specific genes in mice has allowed researchers to study the role of this family of proteins in embryonic development as well as during tumorigenesis. This approach has also made feasible the establishment of loss-of-function tissue-culture models. Such reagents are potentially important as it is possible that subtle determinants of specificity are lost when these proteins are overexpressed.
Phenotypes of Rb -null mice
Although initial analysis indicated that Rb is ubiquitously expressed in adult mouse tissue, levels of Rb mRNA are temporally and spatially controlled during embryogenesis. Rb is highly expressed in a few tissues, including the nervous system, blood cells, skeletal muscle and lens57. Rb−/− mice die between days 13 and 15 of gestation, with pronounced defects in erythroid, neuronal and lens development, as well as skeletal muscle defects58,59,60,61,62 (Table 1). These abnormalities are associated with a partial failure of differentiation, increased apoptosis, as well as ECTOPIC CELL CYCLES. In addition to the cell-cycle changes and insensitivity to several G1-arrest signals that have been seen in Rb-deficient cells14,15,16,63,64,65, cells established from Rb-deficient mouse embryos are defective in myogenesis as well as adipogenesis66,67,68,69,70,71. Together, these findings highlight the multitude of functions of RB in vivo.
Although it seems that changes in E2f-regulated gene expression are responsible for the shortening of the G1 phase of the cell cycle in Rb-null cells, the challenge now is to understand the mechanisms that underlie the various phenotypes seen in Rb-deficient animals and cells. Overall, these phenotypes could potentially reflect cell changes that are mediated by the deregulation of E2f-target genes, but might also be the result of changes in additional transcriptional programmes that are involved in cellular processes, such as differentiation or apoptosis. In fact, it has been suggested that Rb can indirectly or directly potentiate the activity of transcription factors that are involved in differentiation50,70,72,73.
Several lines of evidence indicate that regulation of E2F by RB is involved in apoptosis42. Ectopic expression of E2f1, E2f2 and E2f3 has been shown to induce apotosis in cultured cells as well as in transgenic mice42,43,44,45,48,49,74,75. Furthermore, mutant mouse embryos that lack Rb, and either E2f1 or E2f3, show a significant reduction in the levels of apoptosis, as well as the number of ECTOPIC S-PHASE CELLS relative to those seen in mice lacking only Rb76,77. Furthermore, the disruption of apoptotic regulatory genes that encode proteins such as Apaf1 and caspase-3, which have been shown to be activated by E2f overexpression, can rescue some of the developmental phenotypes seen in Rb-deficient embryos78,79. However, it is important to note that the loss of these individual factors does not allow Rb-deficient embryos to survive until birth. Interestingly, targeted disruption of a gene encoding another Rb-binding protein, Id2 (Ref. 80), in combination with disruption of Rb, allows mice to survive until birth81. These mice, however, show a severe reduction in muscle tissue and die shortly after birth, indicating that loss of Id2 cannot rescue the muscle differentiation defect in Rb−/− mice.
It is also interesting that CHIMERIC Rb animals in which Rb-deficient cells contribute substantially to all tissues are viable and show very mild phenotypes relative to those seen in Rb-deficient mice82,83,84,85,86. This result indicates that Rb might have an additional cell non-autonomous role in development. Experiments with conditional knockout mice, however, have shown that depletion of Rb in the TELENCEPHALON results in neurons that are able to survive and differentiate but exhibit enhanced neurogenesis, due to an increase in cell division87. Taken together, the analysis of Rb-knockout mice has highlighted the complex nature of Rb's normal function during embryonic development.
Phenotypes of p170 - or p130 -null mice
In contrast to the mice that are deficient in Rb, mice deficient in either p107 or 103 develop normally and exhibit no obvious adult phenotypes88,89. However, differences in the genetic background of mice have recently been shown to be important determinants of the developmental consequences of the genetic loss of p107 and p130. Mice with disruptions in p107 and p130 in the C57/BL6-129 genetic background show defective chondrocyte development and die, shortly after birth, of unknown causes. This reveals an apparent functional redundancy between these proteins in some developmental processes88. Mice with disruptions in p107 or p130 in a BALB/c background have more severe phenotypes90,91. Cells isolated from p107- or p130-null mouse embryos have a shortened G1 phase of the cell cycle, similar to that of Rb-deficient cells15,16. However, the genes that are deregulated as a result of p107 or p130 loss are different from those that are deregulated in Rb-deficient cells15,92,93,94. These results indicate that although Rb-family members have distinct roles in the regulation of E2f-mediated gene expression, loss of their function seems to have a similar consequence in the context of cell-cycle progression.
Moreover, there are additional phenotypic similarities between Rb and p107- or p130-deficient cells, such as the inability of these cells to undergo arrest in response to p16 overexpression95. The ability of cells that lack the various pocket proteins to respond to DNA damage is more controversial. Several studies showed that Rb-deficient cells failed to undergo cell-cycle arrest following DNA damage64,65. However, in some studies, cells lacking both p107 and p130 showed a similar failure65, whereas at least one study reported that such cells responded normally to DNA damage64. It is therefore possible that subtle differences in the handling of these cells can substantially impact on their properties in culture.
As described above, cells that lack the various Rb-family proteins have several similar properties. It has become clear, however, that the presence or absence of particular members of the pocket-protein family can markedly affect these properties. For example, in an assay of adipogenic differentiation, RB−/− 3T3 fibroblasts failed to respond to a cocktail of adipogenesis-inducing agents, whereas p107−/−/p130−/− cells differentiated at a much higher rate than wild-type cells66. This difference potentially reflects the differential ability of these proteins to interact with the various E2fs. In fact, consistent with the known specificity of E2f interactions with the various pocket proteins, it was recently reported that E2f1-deficient cells showed a reduced potential for adipogenic differentiation, whereas cells lacking E2f4 showed increased adipogenesis96. However, the possibility that Rb and p107/p130 additionally regulate other transcription factors that are involved in the differentiation programme cannot be ruled out.
The phenotypes of Rb−/−/p107−/− and Rb−/−/p130−/− double-mutant embryos further support the notion that Rb-family members show partial redundancy during development41. So, embryos of both of these genotypes have phenotypes that are similar to Rb−/− embryos; however, these embryos die 2 days earlier. Similarly, chimeric animals that are generated from Rb/p107-deficient embryonal stem (ES) cells exhibit a very low contribution of mutant cells to the developing embryo, whereas ES cells that lack only Rb can contribute approximately 50% to the development of viable embryos83,84,97. Whereas embryos that lack all three pocket proteins have not been characterized, embryo-derived fibroblasts lacking all three proteins have recently been reported98,99. Such cells were found to be unresponsive to many G1-arrest signals (such as growth-factor withdrawal and DNA damage), are impaired in their ability to arrest at confluence and do not respond to senescence-inducing signals. As these defects are substantially more severe than those seen in cells harbouring any of the double-mutant combinations, it seems that, at least in some cellular contexts, all three of these proteins can function redundantly.
Based on the results obtained from the characterization of developmental defects and the analysis of embryo-derived cells from these various knockout mice, it is clear that Rb performs various in vivo functions related to cell proliferation, apoptosis and differentiation. Moreover, it has also become clear from this analysis that the additional Rb-family members, p107 and p130, perform some overlapping functions as well as some seemingly opposing functions. It might be the differences among the biological properties of the pocket proteins that account for the fact that Rb, but not p107 and p130, is selectively lost in tumours.
Tumour phenotypes in mice
Although the predominant tumour type that is seen in human patients harbouring a germline mutation in the RB gene is retinoblastoma, targeted disruption of one allele of Rb in mice does not lead to this cancer. In addition, studies of conditional knockout mice have shown that loss of Rb in mouse photoreceptor cells does not result in retinoblastoma or any other phenotypic changes in these cells100. Instead, mice that are heterozygous for Rb show an increased predisposition to pituitary and thyroid tumours, which are associated with a loss of heterozygosity at the Rb locus62,101. Significantly, when Trp53 (which encodes p53 in mice) is deleted from Rb-heterozygous mice, the animals develop retinal dysplasia61,102. This indicates that loss of p53 could be a rate-limiting event in the development of this tumour in mice, but not in humans. As mentioned previously, loss of Rb in mouse development is associated with increased apoptosis in some tissues, supporting the notion that for loss of Rb to contribute to tumorigenesis, a block to apoptosis must arise. This is consistent with earlier observations that mice that express a T-antigen transgene in the retina develop retinoblastoma103,104. T antigen inactivates all three pocket proteins, as well as p53. Similarly, combined transgenic expression of the papillomavirus E6 and E7 proteins, which inactivate p53 and the pocket proteins, respectively, also promotes retinoblastoma formation in mice105.
Establishing the role of the pocket proteins in the development of retinoblastoma in transgenic and knockout mice has been further complicated by studies in which combinations of pocket proteins have been disrupted. Although mice with disruptions in all three pocket proteins have not been analysed in this context, chimaeras that contain cells with disruptions in both copies of the Rb and p107 genes develop retinoblastoma97. Notably, these tumours seem to develop independently of their p53 status. Loss of p130, however, does not seem to impact on the development of retinoblastoma in an Rb-heterozygous background. Moreover, the loss of p107 and/or p130 does not lead to the development of retinal dysplasia or to the development of any tumour type51.
Collectively, these studies have shown that the development of retinoblastoma in mice might require distinct genetic changes in addition to loss of the Rb gene, and furthermore, that overcoming the tendency to undergo apoptosis is essential to the development of retinoblastoma in mice. Moreover, it is clear from observations in humans, as well as those from animal studies, that loss of RB contributes to tumorigenesis in a relatively small subset of tissue types, potentially reflecting partially redundant functions among the various pocket proteins in other tissues. Alternatively, it is possible that loss of RB is actually detrimental to the development of many tumour types, and that RB expression contributes to tumorigenesis. For example, removal of E2f1 from the mouse germline results in tumour formation in ageing mice, indicating that E2f1-induced apoptosis (repressed by Rb) protects against tumour development106,107. Furthermore, studies have revealed that RB levels are substantially elevated in some cancers, such as colorectal tumours108. Significantly, reduction of RB expression in those cells leads to apoptosis and reduced cell proliferation. High levels of RB might contribute to tumour development by increasing E2f-mediated transcriptional repression, as overexpression of E2f1 has also been shown to result in apoptosis in these same colon cancer cell lines109. Furthermore, studies in transgenic mice have indicated that E2f1 overexpression, which could be analogous to loss of Rb, can, in fact, suppress the transforming ability of a constitutively active Ras allele in some tissues but not in others110,111. Along these same lines, mice that express a 'hyperactive' Rb molecule (a mutated form of Rb that cannot be inactivated by phosphorylation) from the mouse mammary tumour virus (MMTV) promoter show reduced proliferation at the end buds, as well as precocious cellular differentiation and extended survival of the mammary epithelium112. Some of these mice developed hyperplastic nodules as well as mammary adenocarcinoma, indicating that the extended survival/anti-apoptotic effect of hyperactive Rb could promote tumour progression in this setting.
Conclusion and perspectives
The importance of understanding the functional details of the RB regulatory pathway is underscored by the fact that many human tumours have deregulated some of its components. As described above, accumulating evidence from cell-culture experiments, as well as animal model studies, implicates RB and the related p107 and p130 in several cellular processes, thereby complicating the establishment of a precise role for RB as a tumour suppressor. Moreover, it is clear that these proteins perform context-specific overlapping and even opposing functions. This could account for RB's apparently unique role among the pocket proteins as a tumour suppressor, and for its selective loss in a relatively small subset of human tissues. Future studies will certainly reveal new insights into the interplay between the various RB-family members, and about the nature of downstream targets and signalling pathways that impinge on RB function. Importantly, the fruitfly Drosophila melanogaster and nematode worm Caenorhabditis elegans contain conserved pathways with fewer redundancies at each level. Specific components of these pathways have been shown to regulate cell growth, cell death, cell-fate determination and cell-cycle control113,114,115,116,117,118,119,120,121,122,123,124. Genetic studies in these model systems are likely to reveal in vivo partners, upstream regulators and downstream genes of RB in each of these processes.
Finally, with regard to targeting the RB pathway for the treatment of cancer, we might have to revise and re-examine current models in light of recent 'counterintuitive' results, which indicated that loss of RB might, in fact, be detrimental to the development of tumours in some cell types. So, it is possible that targeting RB itself might be an effective antitumour strategy in some contexts.
References
Hungerford, D. A. Some early studies of human chromosomes, 1879–1955. Cytogenet. Cell Genet 20, 1–11 (1978).
Knudson, A. G. Jr. Retinoblastoma: a prototypic hereditary neoplasm. Semin. Oncol 5, 57–60 (1978)
Friend, S. H. et al. A human DNA segment with properties of the gene that predisposes to retinoblastoma and osteosarcoma. Nature 323, 643–646 (1986).
Lee, W. H. et al. Human retinoblastoma susceptibility gene: cloning, identification, and sequence. Science 235, 1394–1399 (1987).
Fung, Y. K. et al. Structural evidence for the authenticity of the human retinoblastoma gene. Science 236, 1657–1661 (1987).
Weinberg, R. A. Tumor suppressor genes. Science 254, 1138–1146 (1991).
Whyte, P. et al. Association between an oncogene and an anti-oncogene: the adenovirus E1A proteins bind to the retinoblastoma gene product. Nature 334, 124–129 (1988).
DeCaprio, J. A. et al. SV40 large tumor antigen forms a specific complex with the product of the retinoblastoma susceptibility gene. Cell 54, 275–283 (1988). References 7 and 8 provided the first description of RB interaction with a viral oncogene – adenovirus E1A and SV40 T Ag; these findings laid the foundation for studies of RB's role in cell-cycle control.
Munger, K. et al. Complex formation of human papillomavirus E7 proteins with the retinoblastoma tumor suppressor gene product. EMBO J. 8, 4099–4105 (1989).
Dyson, N. & Harlow, E., eds. (1992). Adenovirus E1A targets key regulators of cell proliferation. (Imperial Cancer Research Fund).
Qin, X. Q., Chittenden, T., Livingston, D. M. & Kaelin, W. G. Jr. Identification of a growth suppression domain within the retinoblastoma gene product. Genes Dev. 6, 953–964 (1992).
Huang, H. J. et al. Suppression of the neoplastic phenotype by replacement of the RB gene in human cancer cells. Science 242, 1563–1566 (1988).
Goodrich, D. W., Wang, N. P., Qian, Y. -W., Lee, E. Y. -H. P. & Lee, W. -H. The retinoblastoma gene product regulates progression through the G1 phase of the cell cycle. Cell 67, 293–302 (1991).
Herrera, R. E. et al. Altered cell cycle kinetics, gene expression, and G1 restriction point regulation in Rb-deficient fibroblasts. Mol. Cell. Biol. 16, 2402–2407 (1996).
Hurford, R., Cobrinik, D., Lee, M. -H. & Dyson, N. pRB and p107/p130 are required for the regulated expression of different sets of E2F responsive genes. Genes Dev. 11, 1447–1463 (1997).
Classon, M. et al. Combinatorial roles for pRB, p107 and p130 in E2F-mediated cell cycle control. Proc. Natl Acad. Sci. USA 97, 10820–10825 (2000).
Helin, K. et al. A cDNA encoding a pRB-binding protein with properties of the transcription factor E2F. Cell 70, 337–350 (1992).
Kaelin, W. G. Jr et al. Expression cloning of a cDNA encoding a retinoblastoma-binding protein with E2F-like properties. Cell 70, 351–364 (1992).
Shan, B., Durfee, T. & Lee, W. -H. Disruption of RB/E2F-1 interaction by single point mutations in E2F-1 enhances S-phase entry and apoptosis. Proc. Natl Acad. Sci. USA 93, 679–684 (1996).
Nevins, J. R. E2F: a link between the Rb tumor suppressor protein and viral oncoproteins. Science 258, 424–429 (1992).
Dyson, N. The regulation of E2F by pRB-family proteins. Genes Dev. 12, 2245–2262 (1998).
Trimarchi, J. M. & Lees, J. A. Sibling rivalry in the E2F family. Nature Rev. Mol. Cell Biol. 3, 11–20 (2002).
Fattaey, A., Helin, K. & Harlow, E. Transcriptional inhibition by the retinoblastoma protein. – Phil. Trans. R. Soc. Lond. B 340, 333–336 (1993).
Helin, K., Harlow, E. & Fattaey, A. Inhibition of E2F-1 transactivation by direct binding of the retinoblastoma protein. Mol. Cell. Biol. 13, 6501–6508 (1993).
Johnson, D. G., Ohtani, K. & Nevins, J. R. Autoregulatory control of E2F-1 expression in response to positive and negative regulators of cell cycle expression. Genes Dev. 8, 1514–1525 (1994).
Weintraub, S. J., Prater, C. A. & Dean, D. C. Retinoblastoma protein switches the E2F site from positive to negative element. Nature 358, 259–261 (1992). The first study that suggested a role for RB as an active transcriptional repressor, later shown more directly (see reference 27).
Weintraub, S. J. et al. Mechanism of active transcriptional repression by the retinoblastoma protein. Nature 375, 812–815 (1995).
Nevins, J. Toward an understanding of the functional complexity of the E2F and retinoblastoma families. Cell Growth Differ. 9, 585–593 (1998).
Flemington, E. K., Speck, S. H. & Kaelin, J., W. G. E2F-1-mediated transactivation is inhibited by complex formation with the retinoblastoma susceptibility gene product. Proc. Natl Acad. Sci. USA 90, 6914–6918 (1993).
Bremner, R. et al. A. Direct transcriptional repression by pRB and its reversal by specific cyclins. Mol. Cell. Biol. 15, 3256–3265 (1995).
Adnane, J., Shao, Z. & Robbins, P. D. The retinoblastoma susceptibility gene product represses transcription when directly bound to the promoter. J. Biol. Chem. 270, 8837–8843 (1995).
Harbour, J. W. & Dean, D. C. Chromatin remodeling and Rb activity. Curr. Opin. Cell Biol. 12, 685–689 (2000).
Harbour, J. W. & Dean, D. C. Corepressors and retinoblastoma protein function. Curr. Top. Microbiol. Immunol. 254, 137–144 (2001).
Brehm, A. & Kouzarides, T. Retinoblastoma protein meets chromatin. Trends Biochem. Sci. 24, 142–145 (1999).
Buchkovich, K., Duffy, L. A. & Harlow, E. The retinoblastoma protein is phosphorylated during specific phases of the cell cycle. Cell 58, 1097–1105 (1989).
Cooper, S. & Shayman, J. A. Revisiting retinoblastoma protein phosphorylation during the mammalian cell cycle. Cell. Mol. Life Sci. 58 580–595 (2001).
Sherr, C. J. Cancer cell cycles. Science 274, 1672–1677 (1996).
Kennedy, B., Barbie, D., Classon, M., Dyson, N. & Harlow, E. Nuclear organization of DNA replication in primary mammalian cells. Genes Dev. 15, 2855–2868 (2000).
Royzman, I., Austin, R. J., Bosco, G., Bell, S. P. & Orr-Weaver, T. L. ORC localization in Drosophila follicle cells and the effects of mutations in dE2F and dDP. Genes Dev. 13, 827–840 (1999).
Bosco, G., Du, W. & Orr-Weaver, T. DNA replication control through interaction of E2F-RB and the origin recognition complex. Nature Cell Biol. 3, 289–295 (2001). References 38 and 40 indicate a direct role for RB in DNA replication.
Lipinski, M. M. & Jacks, T. The retinoblastoma gene family in differentiation and development. Oncogene 18, 7873–7882 (1999).
Hickman, E. S., Moroni, M. C. & Helin, K. The role of p53 and Rb in apoptosis and cancer. Curr. Opin. Genet. Dev. 12, 60–66 (2002).
Wu, X. & Levine, A. J. p53 and E2F-1 cooperate to mediate apoptosis. Proc. Natl Acad. Sci. USA 91, 3602–3606 (1994).
Qin, X. -Q., Livingston, D. M., Kaelin, W. G. J. & Adams, P. Deregulated transcription factor E2F-1 expression leads to S-phase entry and p53-mediated apoptosis. Proc. Natl Acad. Sci. USA 91, 10918–10922 (1994).
Shan, B. & Lee, W. -H. Deregulated expression of E2F-1 induces S-phase entry and leads to apoptosis. Mol. Cell. Biol. 14, 8166–8173 (1994). References 43–45 first indicated a positive role for E2F1 in inducing apoptosis, raising the possibility that RB could have anti-apoptotic functions.
Guo, Z., Yikang, S., Yoshida, H., Mak, T. W. & Zacksenhaus, E. Inactivation of the retinoblastoma tumor suppressor induces Apaf-1 dependent and independent apoptotic pathways during embryogenesis. Cancer Res. 61, 8395–8400 (2001).
Ishida, S. et al. Role for E2F in control of both DNA replication and mitotic functions as revealed from DNA microarray analysis. Mol. Cell. Biol. 21, 4684–4699 (2001).
Moroni, M. et al. Apaf-1 is a transcriptional target for E2F and p53. Nature Cell Biol. 3, 552–558 (2001).
Muller, H. et al. E2Fs regulate the expression of genes involved in differentiation, development, proliferation, and apoptosis. Genes Dev. 15, 267–285 (2001).
Morris, E. J. & Dyson, N. J. Retinoblastoma protein partners. Adv. Cancer Res. 82, 1–54 (2001).
Mulligan, G. & Jacks, J. The retinoblastoma gene family: cousins with overlapping interests. Trends Genet. 14, 223–229 (1998).
Classon, M. & Dyson, N. p107 and p130: versatile proteins with interesting pockets. Exp. Cell Res. 264, 135–147 (2001).
Zhu, L. et al. Growth suppression by members of the retinoblastoma protein family. Cold Spring Harb. Symp. Quant. Biol. 59, 75–84 (1994).
Zhu, L. et al. The pRB-related protein p107 contains two growth suppression domains: Independent interactions with E2F and cyclin/cdk complexes. EMBO J. 14, 1904–1913 (1995).
Dynlacht, B. D., Moberg, K., Lees, J. A., Harlow, E. & Zhu, L. Specific regulation of E2F family members by cyclin-dependent kinase. Mol. Cell. Biol. 17, 3867–3875 (1997).
Zhu, L., Harlow, E. & Dynlacht, B. D. p107 uses a p21CIP1- related domain to bind cyclin/cdk2 and regulate interactions with E2F. Genes Dev. 9, 1740–1752 (1995).
Jiang, Z., Guo, Z., Saad, F. A. & Zacksenhaus, E. Retinoblastoma gene promoter directs transgene expression exclusively to the nervous system. J. Biol. Chem. 276, 593–600 (2001).
Zacksenhaus, E. et al. pRb controls proliferation, differentiation, and death of skeletal muscle cells and other lineages during embryogenesis. Genes Dev. 10, 3051–3064 (1996).
Jacks, T. et al. Effects of an Rb mutation in the mouse. Nature 359, 295–300 (1992).
Lee, E. Y. et al. Mice deficient for Rb are nonviable and show defects in neurogenesis and haematopoiesis. Nature 359, 288–294 (1992). References 58–60 explore the developmental consequences of loss of Rb in mice, and indicate that Rb is important in proliferation, apototosis and differentiation during development.
Morgenbesser, S. D., Williams, B. O., Jacks, T. & De Pinho, R. A. p53-dependent apoptosis produced by Rb-deficiency in the developing lens. Nature 371, 72–74 (1994).
Jacks, T. Tumor suppressor gene mutations in mice. Annu. Rev. Genet. 30, 603–636 (1996).
Herrera, R. E., Makela, T. P. & Weinberg, R. A. TGFβ-induced growth inhibition in primary fibroblasts requires the retinoblastoma protein. Mol. Biol. Cell. 7, 1335–1342 (1996).
Harrington, E. A., Bruce, J. L., Harlow, E. & Dyson, N. pRB plays an essential role in cell cycle arrest induced by DNA damage. Proc. Natl Acad. Sci. USA 95, 11945–11950 (1998).
Brugarolas, J. et al. Inhibition of cyclin-dependent kinase 2 by p21 is necessary for retinoblastoma protein-mediated G1 arrest after γ-irradiation. Proc. Natl Acad. Sci. USA 96, 1002–1007 (1999). The experiments described in references 64 and 65 indicate a role for RB in DNA-damage-induced G1 arrest.
Classon, M., Kennedy, B. K., Mulloy, R. & Harlow, E. E. Opposing roles for pRB and p107 in adipocyte differentiation. Proc. Natl Acad. Sci. USA 97, 10826–10831 (2000).
Chen, P. -L., Riley, D. J., Chen, Y. & Lee, W. -H. Retinoblastoma protein positively regulates terminal adipocyte differentiation through direct interaction with C/EBPs. Genes Dev. 10, 2794–2804 (1996).
Novitch, B. G., Mulligan, G. J., Jacks, T. & Lassar, A. B. Skeletal muscle cells lacking the retinoblastoma protein display defects in muscle gene expression and accumulate in S and G2 phases of the cell cycle. J. Cell Biol. 135, 441–456 (1996).
Novitch, B., Spicer, D., Kim, P., Cheung, W. & Lassar, A. pRb is required for MEF2-dependent gene expression as well as cell-cycle arrest during skeletal muscle differentiation. Curr. Biol. 9, 449–459 (1999).
Gu, W. et al. Interaction of myogenic factors and the retinoblastoma protein mediates muscle cell commitment and differentiation. Cell 72, 309–324 (1993).
Puri, P. L. et al. Class I histone deacetylases sequentially interact with MyoD and pRB during skeletal myogenesis. Mol. Cell. 8, 885–897 (2001).
Chen, P. L., Riley, D. J., Chen-Kiang, S. & Lee, W. H. Retinoblastoma protein directly interacts with and activates the transcription factor NF-IL6. Proc. Natl Acad. Sci. USA 93, 465–469 (1996).
Sellers, W. R. et al. Stable binding to E2F is not required for the retinoblastoma protein to activate transcription, promote differentiation, and suppress tumor cell growth. Genes Dev. 12, 95–106 (1998).
Kowalik, T. F., DeGregori, J., Schwarz, J. K. & Nevins, J. R. E2F1 overexpression in quiescent fibroblasts leads to induction of cellular DNA synthesis and apoptosis. J. Virol. 69, 2491–2500 (1995).
DeGregori, J., Leone, G., Miron, A., Jakoi, L. & Nevins, J. R. Distinct roles for E2F proteins in cell growth control and apoptosis. Proc. Natl Acad. Sci. USA 94, 7245–7250 (1997).
Ziebold, U., Reza, T., Caron, A. & Lees, J. A. E2F3 contributes both to the inappropriate proliferation and to the apoptosis arising in Rb mutant embryos. Genes Dev. 15, 386–391 (2001).
Tsai, K. Y. et al. Mutation of E2F1 suppresses apoptosis and inappropriate S-phase entry and extends survival of Rb-deficient mouse embryos. Mol. Cell. 2, 293–304 (1998).
Guo, Z., Yikang, S., Yoshida, H., Mak, T. & Zacksenhaus, E. Inactivation of the retinoblastoma tumor suppressor induces Apaf-1 dependent and independent apoptotic pathways during embryogenesis. Cancer Res. 61, 8395–8400 (2001).
Simpson, M. T. W. et al. Caspase 3 deficiency rescues peripheral nervous system defect in retinoblastom nullizygous mice. J. Neurosci. 21, 7089–7098 (2001).
Iavarone, A., Garg, P., Lasorella, A., Hsu, J. & Israel, M. A. The helix-loop-helix protein Id-2 enhances cell proliferation and binds to the retinoblastoma protein. Genes Dev. 8, 1270–1284 (1994).
Lasorella, A., Noseda, M., Beyna, M. & Iavarone, A. Id2 is a retinoblastoma protein target and mediates signalling by Myc oncoproteins. Nature 407, 592–598 (2000).
Lipinski, M., MacLeod, K., Williams, B., Mullaney, T., Crowley, D. & Jacks, T. Cell-autononomous and non-cell-autonomous functions of the Rb tumor suppressor in developing central nervous system. EMBO J. 20, 3402–3413 (2001).
Williams, B. O. et al. Extensive contribution of Rb-deficient cells to adult chimeric mice with limited histopathological consequences. EMBO J. 13, 4251–4259 (1994).
Maandag, E. C. R. et al. Developmental rescue of an embryonic-lethal mutation in the retinoblastoma gene in chimeric mice. EMBO J. 13, 4260–4268 (1994).
Voojis, M. & Berns, A. Developmental defects and tumor predisposition in Rb mutant mice. Oncogene 18, 5293–5303 (1999).
Whyatt, D. & Grosveld, F. Cell-nonautonomous function of the retinoblastoma tumor suppressor protein: new interpretations of old phenotypes. EMBO J. 3, 130–135 (2002).
Ferguson, K. L. et al. Telencephalon-specific Rb knockouts reveal enhanced neurogenesis, survival and abnormal cortical development. EMBO J. 21, 3337–3346 (2002).
Cobrinik, D. et al. Shared role of the pRB-related p130 and p107 proteins in limb development. Genes Dev. 10, 1633–1644 (1996).
Lee, M. -H. et al. Targeted disruption of p107: functional overlap between p107 and Rb. Genes Dev. 10, 1621–1632 (1996).
LeCouter, J. E, Kablar B., Whyte P. F., Ying C., Rudnicki M. A. Strain-dependent embryonic lethality in mice lacking the retinoblastoma-related p130 gene. Development 125, 4669–4679 (1998).
LeCouter, J. E. et al. A. Strain-dependent myeloid hyperplasia, growth deficiency, and accelerated cell cycle in mice lacking the pRB-related p107 gene. Mol. Cell. Biol. 18, 7455–7465 (1998).
Mulligan, G. J., Wong, J. & Jacks, T. p130 is dispensable in peripheral T lymphocytes: evidence for functional compensation by p107 and p130. Mol. Cell. Biol. 18, 206–220 (1998).
Takahashi, Y., Rayman, J. & Dynlacht, B. Analysis of promoter binding by the E2F and pRB families in vivo: distinct E2F proteins mediate activation and repression. Genes Dev. 14, 804–816 (2000).
Wells, J., Boyd, K., Fry, C., Bartley, S. & Farnham, P. Target gene specificity of E2F and pocket protein family members in living cells. Mol. Cell. Biol. 20, 5797–5807 (2000).
Bruce, J., Hurford, R., Classon, M., Koh, J. & Dyson, N. Requirements for cell cycle arrest by p16. Mol. Cell. 6, 737–742 (2000).
Fajas, L. et al. E2Fs regulate adipocyte differentiation. Dev. Cell. 3, 39–49 (2002).
Robanus-Maandag, E. et al. p107 is a suppressor of retinoblastoma development in pRB-deficient mice. Genes Dev. 12, 1599–1609 (1998). References 88, 89 and 97 initiated analyses of mouse strains that are deficient in Rb, p107 and p130, showing separate and overlapping functions of the proteins in this family.
Sage, J. et al. Targeted disruption of the three Rb-related genes leads to loss of G1 control and immortalization. Genes Dev. 14, 3037–3050 (2000).
Dannenberg, J. -H., van Rossum, A., Schuijff, L. & te Riele, H. Ablation of the retinoblastoma gene family deregulates G1 control causing immortalization and increased cell turnover under growth-restricting conditions. Genes Dev. 14, 3051–3064 (2000). References 98 and 99 analyse the phenotype of cells deficient in Rb, p107 and p130 isolated from chimeric animals generated using embryonic stem cells that are deficient in all three proteins. These studies indicate that these cells have deregulation of the cell cycle, but it is presently unknown what the contribution of these cells to different tissues is.
Vooijs, M., te Riele, H., van der Valk, M. & Berns, A. Tumor formation in mice with somatic inactivation of the retinoblastoma gene in interphotoreceptor retinol binding protein-expressing cells. Oncogene 21, 4635–4645 (2002).
Vooijs, M. & Berns, A. Developmental defects and tumor predisposition in Rb mutant mice. Oncogene 18, 5293–5303 (1999).
Williams, B. O. et al. Cooperative tumorigenic effects of germline mutations in Rb and p53. Nature Genet. 7, 480–484 (1994).
Windle, J. J. et al. Retinoblastoma in transgenic mice. Nature 343, 665–669 (1990).
Saenz Robles, M. T., Symonds, H., Chen, J. & van Dyke, T. Induction versus progression of brain tumor development: differential functions for the pRB and p53-targeting domains of SV40 T antigen. Mol. Cell Biol. 14, 2686–2698 (1994).
Howes, K. A. et al. Apoptosis or retinoblastoma: alternative fates of photoreceptors expressing the HPV-16 E7 gene in the presence or absence of p53. Genes Dev. 8, 1300–1310 (1994).
Yamasaki, L. et al. Tumor induction and tissue atrophy in mice lacking E2F-1. Cell 85, 537–548 (1996).
Field, S. J. et al. E2F-1 functions in mice to promote apoptosis and suppress proliferation. Cell 85, 549–561 (1996). The surprising finding in references 106 and 107 was that ageing mice lacking E2F1 displayed a wide range of tumours.
Yamamoto, H. et al. Paradoxical increase in retinoblastoma protein in colorectal carcinomas may protect cells from apoptosis. Clin. Cancer Res. 5, 1805–1815 (1999).
Elliott, M. J., Dong, Y. B., Yang, H. & McMasters, K. M. E2F-1 upregulates c-Myc and p14ARF and induces apoptosis in colon cancer cells. Clin. Cancer Res. 7, 3590–3597 (2001).
Pierce, A. M. et al. E2F1 has both oncogenic and tumor-suppressive properties in a transgenic mouse model. Mol. Cell Biol. 19, 6408–6414 (1999).
Pierce, A. M., Fisher, S. M., Conti, C. J. & Johnson, D. G. Deregulated expression of E2F1 induces hyperplasia and cooperates with ras in skin tumor development. Oncogene 12, 1267–1276 (1998).
Jiang, Z. & Zacksenhaus, E. Activation of retinoblastoma protein in mammary gland leads to ductal growth suppression, precocious differentiation, and adenocarcinoma. J. Cell Biol. 156, 185–198 (2002). The studies in this paper found a counterintuitive role for a constitutively active (phosphorylation-site mutations) Rb molecule that was found to induce adenocarcinoma in transgenic mice, indicating that too much Rb might prevent apoptotic responses that normally safeguard against tumour formation.
Lu, X. & Horvitz, H. R. lin-35 and lin-53, two genes that antagonize a C. elegans Ras pathway, encode proteins similar to Rb and its binding protein RbAp48. Cell 95, 981–991 (1998).
Ceol, C. J. & Horvitz, H. R. dpl-1 DP and efl1 E2F act with lin-35 Rb to antagonize Ras signaling in C. elegans vulval development. Mol Cell, 7, 461–473 (2001).
Boxem, M. & van den Heuvel, S. lin-35 RB and cki-1 Cip/Kip cooperate in developmental regulation of G1 progression in C. elegans. Development 128, 4349–4359 (2001).
Boxem, M. & van den Heuvel, S. C. elegans Class B synthetic multivulva genes act in G(1) regulation. Curr. Biol. 12, 906–911 (2002).
Duronio, R. J., O'Farrell, P. H., Xie, J. -E., Brook, A. & Dyson, N. The transcription factor E2F is required for S phase during Drosophila embryogenesis. Genes Dev. 9, 1445–1455 (1995).
Du, W., Vidal, M., Xie, J. -E. & Dyson, N. RBF, a novel RB-related gene that regulates E2F activity and interacts with cyclin E in Drosophila. Genes Dev. 10, 1206–1218 (1996).
Du, W. Suppression of the rbf null mutants by a de2f1 allele that lacks transactivation domain. Development 127, 367–379 (2000).
Frolov, M. V. et al. Functional antagonism between E2F family members. Genes Dev. 15, 2146–2160 (2001).
Myster, D. L., Bonnette, P. C. & Duronio, R. J. A role for the DP subunit of the E2F transcription factor in axis determination during Drosophila oogenesis. Development 127, 3249–3261 (2000).
Cayirlioglu, P., Bonnette, P. C., Dickson, M. R. & Duronio, R. J. Drosophila E2f2 promotes the conversion from genomic DNA replication to gene amplification in ovarian follicle cells. Development 128, 5085–5098 (2001).
Stevaux, O., D., D., Frolov, M. V., Taylor-Harding, B., Morris, E. & Dyson, N. Distinct mechanisms of E2F regulation by Drosophila RBF1 and RBF2. EMBO J. 21, 4297–4937 (2002).
Neufeld, T. P., de la Cruz, A. F. A., Johnston, L. A. & Edgar, B. A. Coordination of cell growth and division by Drosophila E2F. Cell 93, 1183–1193 (1998).
Lee, J. -O., Russo, A. A. & Pavletich, N. P. Structure of the retinoblastoma tumour-suppressor pocket domain bound to a peptide from HPV E7. Nature 391, 859–865 (1998). Describes the crystal structure of the RB pocket bound to E7 and, as such, laid the foundation for many mutational studies of this domain.
Dick, F. A., Sailhamer, E. & Dyson, N. J. Mutagenesis of the pRB pocket domain reveals that cell cycle arrest functions are separable from binding to viral oncoproteins. Mol. Cell Biol. 20, 3715–3727 (2000).
Dick, F. A. & Dyson, N. J. Three regions of the pRB pocket domain affect its inactivation by human papillomavirus E7 proteins. J. Virol. 76, 6224–6234 (2002).
Dahiya, A., Gavin, M. R., Luo, R. X. & Dean, D. C. Role of the LXCXE binding site in Rb function. Mol. Cell Biol. 20, 6799–6805 (2000).
Chen, T. -T. & Wang, J. Y. J. Establishment of irreversible growth arrest in myogenic differentiation requires the RB LXCXE-binding function. Mol. Cell Biol. 20, 5571–5580 (2000).
Knudsen, K. E. et al. RB-dependent S-phase response to DNA damage. Mol. Cell Biol. 20, 7751–7763 (2000).
Chestukhin, A., Litovchick, L., Rudich, K. & De Caprio, J. A. Nucleocytoplasmic shuttling of p130: novel regulatory mechanism. Mol. Cell Biol. 22, 453–468 (2002).
Chan, H. M., Krstic-Demonacos, M., Smith, L., Demonacos, C. & N. B., L. Acetylation control of the retinoblastoma tumour suppressor protein. Nature Cell Biol. 3, 667–674 (2001).
Acknowledgements
We thank J. Settleman, S. van den Heuvel, N. Dyson, F. Dick, E. Morris and D. Dimova for helpful comments and/or reading of this manuscript. We also thank G. Klein and all previous and present members of the Harlow and Dyson laboratories for helpful discussions over the years. Finally, we apologize to all our colleagues whose interesting work we could not cite, due to space restrains.
Author information
Authors and Affiliations
Glossary
- ECTOPIC CELL CYCLE
-
Unscheduled or inappropriate entry into the cell cycle.
- ECTOPIC S-PHASE CELLS
-
Unscheduled or inappropriate entry into S phase.
- CHIMAERA
-
An organism with a genetic mixture of cells, such as wild type and mutant.
- TELENCEPHALON
-
Evolutionarily, the most recently evolved part of the brain. It is the forebrain, which is involved in cognition, learning and memory.
Rights and permissions
About this article
Cite this article
Classon, M., Harlow, E. The retinoblastoma tumour suppressor in development and cancer. Nat Rev Cancer 2, 910–917 (2002). https://doi.org/10.1038/nrc950
Issue Date:
DOI: https://doi.org/10.1038/nrc950
This article is cited by
-
Retinoblastoma: present scenario and future challenges
Cell Communication and Signaling (2023)
-
Post-translational modifications on the retinoblastoma protein
Journal of Biomedical Science (2022)
-
Cellular senescence or stemness: hypoxia flips the coin
Journal of Experimental & Clinical Cancer Research (2021)
-
Decreased PGC1β expression results in disrupted human erythroid differentiation, impaired hemoglobinization and cell cycle exit
Scientific Reports (2021)
-
Inhibiting CDK4/6 in Breast Cancer with Palbociclib, Ribociclib, and Abemaciclib: Similarities and Differences
Drugs (2021)