Key Points
-
Conservation biology is considered to be a 'crisis discipline', and therefore requires an arsenal of approaches and techniques to ensure that educated, rapid decisions about endangered species and threatened areas are made.
-
The expansion of genomic technologies in data acquisition, storage and analysis has assisted in increasing the efficiency of the field of conservation genetics and in helping conservation decision-making.
-
Conservation genetics leads us to a better picture of pattern and process in endangered species, and allows the use of pattern and process in conservation decision-making.
-
The most important applications of conservation genetics in the future will be those that incorporate both pattern and process into a cohesive approach to decision-making.
-
A wide range of unconventional sources for DNA is used in conservation work; this includes faeces, feathers, fur, sloughed-off skin, plants from herbarium sheets and clever direct-biopsy approaches that minimize invasiveness.
-
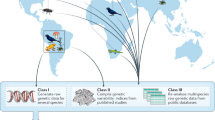
One approach that will be increasingly important in the expansion of conservation genetics concerns recent efforts to form centralized DNA barcode databases and DNA registries.
-
Conservation genetics itself needs to be placed in the context of the difficulties of working across political boundaries, amidst economic challenges and in the face of the complexity of using science to inform management decisions.
Abstract
The 'crisis discipline' of conservation biology has voraciously incorporated many technologies to speed up and increase the accuracy of conservation decision-making. Genetic approaches to characterizing endangered species or areas that contain endangered species are prime examples of this. Technical advances in areas such as high-throughput sequencing, microsatellite analysis and non-invasive DNA sampling have led to a much-expanded role for genetics in conservation. Such expansion will allow for more precise conservation decisions to be made and, more importantly, will allow conservation genetics to contribute to area- and landscape-based decision-making processes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Jablonski, D. Background and mass extinctions: the alternation of macroevolutionary regimes. Science, 231, 129–133 (1986).
Lande, R. Genetics and demography in biological conservation. Science 241, 1455–1460 (1988). This landmark paper was one of the first to discuss the relevance of genetics and demography in conservation biology.
Soulé, M. E. Conservation biology: the science of scarcity and diversity (Sinauer Associates, Sunderland, Massachusetts, 1986).
Schlotterer, C. The evolution of molecular markers — just a matter of fashion? Nature Rev. Genet. 5, 63–69 (2004).
Godwin, I. D., Aitken, E. A. & Smith, L. W. Application of inter-simple sequence repeat (ISSR) markers to plant genetics. Electrophoresis 18, 1524–1528 (1997).
Zhang, D. X. & Hewitt, G. M. Nuclear DNA analyses in genetic studies of populations: practice, problems and prospects. Mol. Ecol. 12, 563–584 (2003).
Aitken, N., Smith, S., Schwarz, C. & Morin, P. A. Single nucleotide polymorphism (SNP) discovery in mammals: a targeted-gene approach. Mol. Ecol. 13, 1423–1431 (2004).
Brumfield, R., Nickerson, D., Beerli, P. & Edwards, S. V. The utility of single nucleotide polymorphisms in inferences of population history. Trends Ecol. Evol. 18, 249–256 (2003).
Gibson, G. Microarrays in ecology and evolution: a preview. Mol. Ecol. 11, 17–24 (2002).
Purugganan, M. & Gibson, G. Merging ecology, molecular evolution, and functional genetics. Mol. Ecol. 12, 1109–1112 (2003).
Bouchier, C., Boyle, T. & Young, A. Forest Conservation Genetics (CSIRO, Australia, 2000).
Avise, J. & Hamrick, P. Conservation Genetics: Case Histories From Nature (Chapman and Hall, New York, 1996).
Spellerberg, I. Conservation Biology (Longman Group, Ltd, Harlow, 1997).
Avise, J. C. Molecular Markers, Natural History and Evolution (Sinauer Associates, Sunderland, Massachusetts, 2004).
Frankham, R., Ballou, J. D. & Briscoe, D. A. Introduction to Conservation Genetics (Cambridge Univ. Press, Cambridge, United Kingdom, 2003). A comprehensive overview of the state of modern conservation genetics.
Ryder, O. A. Species conservation and systematics: the dilemma of the subspecies. Trends Ecol. Evol. 1, 9–10 (1986).
O'Brien, S. J. & Mayr, E. Bureaucratic mischief: recognizing endangered species and subspecies. Science 251, 1187–1188 (1991).
Amato, G. Species hybridization and protection of endangered animals. Science 253, 250–251 (1991).
Waples, R. Pacific salmon, Oncorhynchus spp., and the definition of 'species' under the Endangered Species Act. Marine Fisheries Rev. 53, 2–11 (1991).
Caughley, G. Directions in conservation biology. J. Animal Ecol. 63, 215–244. (1994).
Hedrick, P. W., Lacy, R. C. Allendorf, F. W. & Soulé, M. E. Directions in conservation biology. Comments on Caughley. Conserv. Biol. 10, 1312–1320 (1996).
Allendorf, F. W. & Leary, R. F. in Conservation Biology (ed. Soule, M. E.), 57–76 (Sinauer Associates, Sunderland, Massachusetts, 1986).
Waples, R. S. A generalized approach for estimating effective population size from temporal changes in allele frequency. Genetics 121, 379–391 (1989).
Templeton, A. R. & Read, B. Factors eliminating inbreeding depression in a captive herd of Speke's gazelle. Zoo Biol. 3, 177–199 (1984).
Ralls, K., Ballou, J. D. & Templeton, A. Estimates of lethal equivalents and the cost of inbreeding in mammals. Conserv. Biol. 2, 40–56 (1988).
Goodnight, K. F. & Quellar, D. C. Computer software for performing likelihood tests of pedigree relationship using genetic markers. Mol. Ecol. 8, 1231–1234 (1999).
Ritland, K. Marker-inferred relatedness as a tool for detecting heritability in nature. Mol. Ecol. 9, 1195–1204 (2000).
Blouin, M. S. DNA-based methods for pedigree reconstruction and kinship analysis in natural populations. Trends Ecol. Evol. 18, 503–511 (2003).
Glaubitz, J. C., Rhodes, E. & Dewoody, J. E. Prospects for inferring pairwise relationships with single nucleotide polymorphisms. Mol. Ecol. 12, 1039–1047 (2003).
Petit, E., Balloux, F. & Excoffier, L. Mammalian population genetics: why not Y? Trends Ecol. Evol. 17, 28–35 (2002).
Vekemens, X. & Hardy, O. J. New insights from fine-scale spatial genetic structure analyses in plant populations. Mol. Ecol. 13, 921–935 (2004).
Lacy, R. C. Should we select genetic alleles in our conservation breeding programs? Zoo Biol. 19, 279–282 (2000).
Miller, C. R., Adams, J. R. & Waits, L. P. Pedigree-based assignment tests for reversing coyote (Canis latrans) introgression into the wild red wolf (Canis rufus) population. Mol. Ecol. 12, 3287–3299 (2003).
Geyer, C. J., Ryder, O. A, Chemnick, L. G. & Thompson, E. A. Analysis of relatedness in the California condors, from DNA fingerprints. Mol. Biol. Evol. 10, 571–589 (1993).
Russello, M. & Amato, G. Ex situ management in the absence of pedigree information: integration of microsatellite-based estimates of relatedness into a captive breeding strategy for the St. Vincent Amazon Parrot (Amazona guildingii). Mol. Ecol. (in the press).
Russello, M. & Amato, G. Application of a non-invasive, PCR-based test for sex identification in an endangered parrot, Amazona guildingii. Zoo Biol. 20, 41–45 (2001).
Templeton, A. R. Nested clade analyses of phylogeographic data: testing hypotheses about gene flow and population history. Mol. Ecol. 7, 381–397 (1998). A clear description of the nested clade approach and how it can be applied to a better understanding of ecological, demographic and genetic aspects of populations.
Neigel, J. E. Is FST obsolete? Conserv. Genet. 3, 167–173 (2002).
Excoffier, L., Smouse, P. E. & Quattro, J. M. Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131, 479–491 (1992).
Schneider, S., Roessli, D. & Excoffier, L. Arlequin: A Software for Population Genetics Data Analysis. Ver 2.000 (Genetics and Biometry Laboratory, Department of Anthropology, Univ. Geneva, 2000).
Goldstein, P. Z. & DeSalle, R. Phylogenetic species, nested hierarchies, and character fixation. Cladistics 16, 364–384 (2000).
Davis, J. I. & Nixon, K. C. Populations, genetic variation, and the delimitation of phylogenetic species. Systematic Biol. 41, 421–435 (1992).
Sites, J. W. & Crandall, K. A. Testing species boundaries in biodiversity studies. Conserv. Biol. 11, 1289–1297 (1997).
Goldstein, P. Z., DeSalle, R., Amato, G. & Vogler, A. Conservation genetics at the species boundary. Conserv. Biol. 14, 120–131 (2000).
Sites, J. & Marshall, J. C. Delimiting species: a renaissance issue in systematic biology. Trends Ecol. Evol. 18, 461–471 (2003). An excellent review of the methods and approaches used to delimit species, describing character-based and tree-based methods of inference.
Losos, J. B. & Glor, R. E. Phylogenetic comparative methods and the geography of speciation. Trends Ecol. Evol. 18, 220–227 (2003).
Moritz, C. Defining 'evolutionarily significant units' for conservation. Trends Ecol. Evol. 9, 373–375 (1994).
Posada, D., Crandall, K. A. & Templeton, A. R. GeoDis: a program for the cladistic nested analysis of the geographical distribution of genetic haplotypes. Mol. Ecol. 9, 487–488 (2000).
Clement, M., Posada, D. & Crandall, K. A. TCS: a computer program to estimate gene genealogies. Mol. Ecol. 9, 1657–1659 (2000).
Templeton, A. R. Using phylogeographic analyses of gene trees to test species status and processes. Mol. Ecol. 10, 779–791 (2001).
Templeton, A. R. Statistical phylogeography: methods of evaluating and minimizing inference errors. Mol. Ecol. 13, 789–809 (2004).
Crandall, K. A. Conservation phylogenetics of Ozark crayfishes: assigning priorities for aquatic habitat protection. Biol. Conserv. 84, 107–117 (1998).
Nixon, K. C. & Wheeler, Q. D. in Extinction and Phylogeny (eds Novacek, M. J. & Wheeler, Q. D.) 216–234 (Columbia Univ. Press, New York, 1992). Clearly and concisely describes the role of systematics in delimiting measures of cladistic diversity that might be useful in conservation biology.
Faith, D. P. Conservation evaluation and phylogenetic diversity. Biol. Conserv. 61, 1–10 (1992).
Faith, D. P. Systematics and conservation: on predicting the feature diversity of subsets of taxa. Cladistics 8, 361–373 (1992).
Faith, D. P. Quantifying biodiversity: a phylogenetic perspective. Conserv. Biol. 16, 248–252 (2002).
Crozier, R. H. Preserving the information content of species: genetic diversity, phylogeny, and conservation worth. Annu. Rev. Ecol. Systematics 28, 243–268 (1997).
Neel, M. & Cummings, M. P. Genetic consequences of ecological reserve design guidelines: an empirical investigation. Conserv. Genet. 4, 427–439 (2003).
Palumbi, S. R. in Marine Community Ecology (eds Bertness, M. D., Gaines, S. D. & Hay, M. E.) 509–530 (Sinauer Associates, Sunderland, Massachusetts, 2001).
Moritz, C., Patton, J. L., Schneider, C. J. & Smith, T. B. Diversification of rainforest faunas: an integrated molecular approach. Ann. Rev. Ecol. Syst. 31, 533–563 (2000).
Gibbs, J. P. & Amato, G. in Turtle Conservation (ed. Klemens, M. W) 207–217 (Smithsonian Institution Press, Washington and London, 2000).
Roemer, G. W. & Wayne, R. K. Conservation in conflict: a tale of two endangered species. Conserv. Biol. 17, 1251–1260 (2003).
Brook, B. W., Tonkyn, D. W., O'Grady, J. J. & Frankham, R. Contribution of inbreeding to extinction risk in threatened species. Conserv. Ecol. 6, 6–28 (2002).
Roman, J. & Palumbi, S. R. Whales before whaling in the North Atlantic. Science 301, 508–510 (2003).
Rosenbaum, H. C. et al. The effect of different reproductive success on population genetic structures: correlations of life history with matrilines in humpback whales of the Gulf of Maine. J. Hered. 93, 389–399 (2002).
Lacy, R. C. Structure of the VORTEX simulation model for population viability analysis. Ecol. Bull. 48, 191–203 (2000).
Templeton, A. R. in Conservation Biology: the Science of Scarcity and Diversity (ed. Soulé, M. E.) 105–116 (Sinauer Associates, Sunderland, Massachusetts, 1986).
Ballou, J. & Lacy, R. C. in Population Management for Survival and Recovery (eds Ballou, J., Gilpin, M. & Foose, T. J.) 76–111 (Columbia Univ. Press, New York, 1995).
Ruggiero, L. F., Hayward, G. D. & Squires, J. R. Viability analysis in biological evaluations: concepts of population viability analysis, biological population, and ecological scale. Conserv. Biol. 8, 364–372 (1984).
Beissinger, S. R. & McCullough, D. R. Population Viability Analysis (Univ. Chicago Press, Chicago, 2002).
Reed, D. H., O'Grady, J. J., Brook, B. W., Ballou, J. D. & Frankham, R. Estimates of minimum viable population size for vertebrates and factors affecting those estimates. Biological Conserv. 113, 23–34 (2003).
Wayne, R. K. & Jenks, S. M. Mitochondrial DNA analysis implying extensive hybridization of the endangered red wolf Canis rufus. Nature 351, 565–568 (1991).
Roy, M. S., Girman, D. J., Taylor, A. C. & Wayne, R. K. The use of museum specimens to reconstruct the genetic variability and relationships of extinct populations. Experientia 50, 551–557 (1994).
Garcia-Moreno, J., Roy, M. S., Geffen, E. & Wayne, R. K. Relationships and genetic purity of the endangered Mexican Wolf based on analysis of microsatellite loci. Conserv. Biol. 10, 376–389 (1996).
Duagherty, C. H., Cree, A., Hay, J. M. & Thompson, M. B. Neglected taxonomy and continuing extinctions of tuatara (Sphenodon) Nature 347, 177–179 (1990).
DeSalle, R. & Birstein, V. PCR analysis of black caviar. Nature 381, 197–198 (1996).
Birstein, V. J., Doukakis, P., Sorkin, B. & DeSalle, R. Population aggregation analysis of three caviar-producing species of sturgeons and implications for the species identification of black caviar. Conserv. Biol. 12, 766–775 (1998).
Ginsberg, J. CITES at 30, or 40. Conserv. Biol. 12, 1184–1191 (2002).
Birstein, V. J., Doukakis, P. & DeSalle, R. Caviar species identification and polyphyletic structure of Russian sturgeon. Conserv. Genet. 1, 81–88 (2000).
Hedrick, P. W. Gene flow and genetic restoration: the Florida panther; a case study. Conserv. Biol. 9, 996–1007 (1995).
DeMauro, M. M. Relationship of breeding system to rarity in the lakeside daisy (Hymenoxys acaulis var. glabra). Conserv. Biol. 7, 542–550 (1993).
Payne, R. B. & Sorensen, M. D. Museum collections as sources of genetic data. Bonner Zoologische Beiträge Band 51, 97–104 (2002).
Hofreiter, M., Serre, D., Poinar, H. N., Kuch, M. & Paabo, S. Ancient DNA. Nature Rev. Genet. 2, 353–359 (2001).
Rosenbaum, H. C. et al. The utility of museum specimens in right whale conservation genetics. Conserv. Biol. 14, 1837–1842 (2000). The authors used century-old baleen samples to elucidate the genetic status of north Atlantic populations of right whales at the turn of the nineteenth century.
Goldstein, P. Z. & DeSalle, R. Calibrating phylogenetic species formation in a threatened species using DNA from historical specimens. Mol. Ecol. 12, 1993–1998 (2003). The authors used century-old pinned tiger beetles to assess the phylogeographic patterns in this endangered insect along the north-east Atlantic coast.
Bouzat, J. L., Lewin, H. A. & Paige, K. N. The ghost of genetic diversity past: historical DNA analysis of the greater prairie chicken. American Naturalist 152, 1–6 (1998).
Bellinger, M. R., Johnson, J. A., Toepfer, J. & Dunn, P. Loss of genetic variation in greater prairie chickens following a population bottleneck in Wisconsin USA. Conserv. Biol. 17, 717–724 (2003).
Amato, G. et al. The rediscovery of Roosevelt's barking deer (Muntiacus rooseveltorum). J. Mammalogy 80, 639–643 (1999).
Fleischer, R. C., Olson, S. L., James, H. F. & Cooper, A. C. Identification of the extinct Hawaiian eagle (Haliaeetus) by mtDNA sequence analysis. Auk 117, 1051–1056 (2000).
Fleischer, R. C., Tarr, C. L., James, H. F., Slikas, B. & Mcintosh, C. E. Phylogenetic placement of the Po'ouli, Melamprosops phaeosoma, based on mitochondrial DNA sequence data and osteological characters. Studies Avian Biol. 22, 98–103 (2001).
Glenn, T. C., Stephan, W. & Braun, M. J. Effects of a population bottleneck on Whooping Crane mitochondrial DNA variation. Conserv. Biol. 13, 1097–1107 (1999).
Nielsen, E. E., Hansen, M. M. & Loeschcke, V. Genetic variation in time and space: microsatellite analysis of extinct and extant populations of Atlantic salmon. Evolution 53, 261–268 (1999).
Hoelzel, A. R. et al. Elephant seal genetic variation and the use of simulation models to investigate historical population bottlenecks. J. Hered. 84, 443–449 (1993).
Leonard, J. A., Wayne, R. K. & Cooper, A. Population genetics of Ice Age brown bears. Proc. Natl Acad. Sci. USA 97, 1651–1654 (2000).
Rogers, S. O. & Bendich, A. J. Extraction of DNA from milligram amounts of fresh, herbarium and mummified plant tissues. Plant Mol. Biol. 5, 69–76 (1985).
Paabo, S. Of bear conservation genetics and the value of time travel. Proc. Natl Acad. Sci. USA 97, 1320–1321 (2000).
Cooper, A. et al. Ancient DNA and island endemics. Nature 381, 484 (1996).
Höss, M., Dilling, A., Currant, A. & Pääbo, S. Molecular phylogeny of the extinct ground sloth Mylodon darwinii. Proc. Natl Acad. Sci. USA 93, 181–185 (1996).
Poinar, H., Kuch, M., McDonald, G., Martin, P. & Paabo, S. Nuclear gene sequences from a late pleistocene sloth coprolite. Curr. Biol. 13, 1150–1152 (2003).
Lanza, R. P. et al. Cloning of an endangered species (Bos gaurus) using interspecies nuclear transfer. Cloning 2, 79–90 (2000).
Hebert, P. D., Cywinska, A., Ball, S. L. & deWaard, J. R. Biological identifications through DNA barcodes. Proc. R. Soc. Lond. B 270, 313–322 (2003). Articulates the usefulness of DNA sequences for identifying species, and suggests that a single gene — which encodes cytochrome oxidase 1 — will be adequate for 'barcoding' all animal life.
Stoeckle, M. Taxonomy, DNA, and the barcode of life. BioScience 53, 2–3 (2003).
Baker, S., Dalebout, M. L., Lavery, S. & Ross, H. A. www.DNA-surveillance: applied molecular taxonomy for species conservation and discovery. Trends Ecol. Evol. 18, 271–272 (2003). A description of the construction and public availability of a web site for whale surveillance using DNA-sequence variation as data to establish relatedness.
Tautz, D., Arctander, P., Minelli, A., Thomas, R. H. & Vogler, A. P. A plea for DNA taxonomy. Trends Ecol. Evol. 18, 70–74 (2003).
Dunn, C. P. Keeping taxonomy based in morphology. Trends Ecol. Evol. 18, 270–271 (2003).
Seberg, O. et al. Shortcuts in systematics? A commentary on DNA-based taxonomy. Trends Ecol. Systematics 18, 63–64 (2003).
Lipscomb, D., Platnick, N. & Wheeler, Q. The intellectual content of taxonomy: a comment on DNA taxonomy. Trends Ecol. Systematics 18, 64–65 (2003).
Cipriano, F. & Palumbi, S. R. Genetic tracking of a protected whale. Nature 397, 307–308 (1999).
Angermeier, P. L. The natural imperative for biological conservation. Conserv. Biol. 14, 373–381 (2000).
Hunter, M. Benchmarks for managing ecosystems: are human activities natural? Conserv. Biol. 10, 695–697 (1996). The context of what is 'natural' is an important aspect of conservation biology, as discussed in this article. The article goes on to define the important aspects of what is natural.
Haila, Y., Comer, P. J. & Hunter, M. A 'natural' benchmark for ecosystem function. Conserv. Biol. 11, 300–305 (1997).
Morin, P. A. Genetic resources: opportunities and perspectives for the new century. Conserv. Genet. 1, 271–275 (2000).
Moran, P. Current conservation genetics: building an ecological approach to the synthesis of molecular and quantitative genetic methods. Ecol. Freshwater Fish 11, 30–55 (2002).
Bertotorelle, G., Bruford, M., Chemini, C., Vernesi, C. & Hauffe, H. C. New flexible Bayesian methods to revolutionize conservation genetics. Conserv. Biol. 18, 584–585 (2004).
Freyfogle, E. T. & Lutz Newton, J. Putting science in its place. Conserv. Biol. 16, 863–873 (2002).
Sanderson, E. W. et al. The human footprint and the last of the wild. Bioscience 52, 891–904 (2002).
Acknowledgements
The authors thank M. Russello for assistance in preparing figure 3e and acknowledge the continued support to the Conservation Genetics Program from the American Museum of Natural History (AMNH) Center for Biodiversity Conservation. R.D. thanks the Louis and Dorothy Cullman Program for Molecular Systematics at the AMNH.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Glossary
- SYSTEMATICS
-
The field of biology concerned with the diversity of life on Earth. Systematics is usually viewed as having two components — phylogenetics and taxonomy.
- RANDOMLY AMPLIFIED POLYMORPHIC DNA
-
(RAPD). DNA fragments that are generated by PCR using one or two randomly selected oligonucleotides or primers and are polymorphic in size.
- MINISATELLITES
-
Regions of DNA in which repeat units of 6–100 bp are arranged in tandem arrays that are 0.5–30 kb in length
- RESTRICTION-FRAGMENT-LENGTH POLYMORPHISM
-
(RFLP). A fragment-length variant of a DNA sequence that is generated through the gain or loss of a restriction site due to a DNA substitution.
- ALLOZYMES
-
Co-dominant nuclear DNA markers that consist of enzymes that differ in their mobility on a charged gel.
- PARAPHYLETIC
-
A term applied to a clade of organisms that includes the most recent common ancestor of all of its members, but not all of the descendants of that most recent common ancestor.
- AMPLIFIED-FRAGMENT-LENGTH POLYMORPHISM
-
(AFLP). A DNA marker generated by digestion of genomic DNA with two restriction enzymes to create many DNA fragments, followed by ligation of specific sequences of DNA (adaptors) to the ends of these fragments, amplification by PCR (using primers corresponding to the adaptors plus random combinations of three additional bases at the end) and visualization of the fragments by gel electrophoresis.
- MICROSATELLITES
-
Co-dominant nuclear DNA markers that consist of sets of short, repeated nucleotide sequences.
- INTER-SIMPLE-SEQUENCE REPEAT
-
(ISSR). DNA fragments found between adjacent, oppositely oriented microsatellites. DNA is amplified by PCR, separated by gel electrophoresis and scored for the presence or absence of fragments.
- SUBSPECIES
-
A physically distinct subunit of a species.
- EVOLUTIONARILY SIGNIFICANT UNIT
-
(ESU). A population of organisms that is reproductively isolated from other populations of the same species, and represents an important component in the evolutionary legacy of the species.
- INBREEDING DEPRESSION
-
Reduction in fitness or vigour caused by one or more generations of inbreeding.
- BREEDING EFFECTIVE POPULATION SIZE
-
The number of individuals that make up the breeding population in an idealized population.
- MINIMUM VIABLE POPULATION SIZE
-
An estimate of the smallest number of individuals in a population that is capable of maintaining that population without significant manipulation.
- WRIGHT'S INBREEDING COEFFICIENTS
-
Measures of inbreeding in a sub-population first devised by Sewall Wright to describe the amount of homozygosity in a population due to inbreeding. Measures of inbreeding at different hierarchical levels of comparison can be obtained using this approach.
- COALESCENCE-THEORY-BASED ANALYSIS
-
A means of investigating the shared genealogical history of genes. A genealogy is constructed backwards in time, starting with the present-day sample. Lineages coalesce when they have a common ancestor.
- POPULATION-VIABILITY ANALYSIS
-
(PVA). The process of identifying threats faced by a species and incorporating these threats into an estimation of the likelihood of persistence of a species for a given time in the future.
- POPULATION-AGGREGATION ANALYSIS
-
(PAA). Provides a straightforward criterion for demarcating a phylogenetic break between aggregates of individuals. An aggregate is said to be distinct when an attribute or combination of attributes in one aggregate is fixed and different from an attribute or combination of attributes in another aggregate.
- CLADISTIC DIVERSITY
-
A measurement that ranks areas for biodiversity-conservation priorities based on information in cladograms or phylogenetic trees. Also called phylogenetic diversity.
- AREAS OF ENDEMISM
-
Geographical areas where maximal numbers of endemic species exist.
- CETACEAN
-
An aquatic mammal of the order Cetacea, including whales, porpoises and dolphins.
- MITOCHONDRIAL D-LOOP
-
The highly variable, non-coding, portion of the mammalian mitochondrial genome. A D-loop is the configuration found during DNA replication of chloroplast and mitochondrial chromosomes wherein the origin of replication is different on the two strands. The first structure formed is a displacement loop or D-loop.
- SPECIES-SURVIVAL PROGRAMS
-
Programmes established by the American Zoo and Aquarium Association to ensure the survival of selected wild or captive species through cooperative genetic and demographic management.
- CONVENTION ON IMPORTATION AND TRADE OF ENDANGERED SPECIES OF WILD FLORA AND FAUNA
-
(CITES). A convention first started in 1973 and established amongst participating nations to restrict the international commerce of plant and animal species harmed by international trade.
- MANAGEMENT UNIT
-
(MU). A population, stock or group of stocks of a species that are aggregated for the purposes of achieving a desired conservation objective.
- SEMISPECIES
-
An emerging species in the early stages of speciation.
- INCIPIENT SPECIES
-
A newly formed species pair.
- ENDANGERED SPECIES ACT
-
(ESA). A US congressional act established in 1973 that articulates guidelines and rules for the protection of species on the brink of extinction.
- NATURAL; NATURALNESS
-
A thing is natural if it is not made by humans. Naturalness is the degree to which something is natural.
- NATURAL IMPERATIVES
-
An essential and urgent duty to restore some aspect of the environment to its 'natural' state.
- CONVENTION OF THE PARTIES
-
Also known as the Conference of the Parties to the Convention on Biological Diversity. An international organization dedicated to fostering conservation, sustainable use and equitable benefit-sharing concerning biological diversity.
Rights and permissions
About this article
Cite this article
DeSalle, R., Amato, G. The expansion of conservation genetics. Nat Rev Genet 5, 702–712 (2004). https://doi.org/10.1038/nrg1425
Issue Date:
DOI: https://doi.org/10.1038/nrg1425
This article is cited by
-
Ecological niche modelling and population genetic analysis of Indian temperate bamboo Drepanostachyum falcatum in the western Himalayas
Journal of Plant Research (2023)
-
Population genetic analysis illustrated a high gene diversity and genetic heterogeneity in Himalayacalamus falconeri: a socio-economically important Indian temperate woody bamboo taxon
Journal of Plant Biochemistry and Biotechnology (2023)
-
Islands in the desert: assessing fine scale population genomic variation of a group of imperiled desert fishes
Conservation Genetics (2022)
-
Population genetics of three at-risk tiger beetles Habroscelimorpha dorsalis dorsalis, H. d. media, and Ellipsoptera puritana
Conservation Genetics (2022)
-
Phylogeography of the capybara, Hydrochoerus hydrochaeris, in a large portion of its distribution area in South America
Journal of Mammalian Evolution (2022)