Key Points
-
Protein kinases are a large family of regulatory enzymes that change multiple processes in cells by addition of inorganic phosphate to other proteins.
-
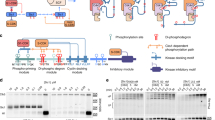
All eukaryotic protein kinases share a conserved catalytic core that provides effective phosphotransfer from a molecule of ATP to the recipient protein substrate. Despite this similarity, each kinase phosphorylates only its own substrate or a set of substrates. Understanding how this specificity is achieved is a major challenge.
-
Cyclic AMP-dependent kinase (PKA) is one of the most important protein kinases, as it is the major sensor of cAMP in cells. cAMP is an important second messenger that transmits various signals in living cells.
-
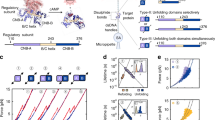
PKA has four major isoforms that are very similar at the level of their catalytic domains or cAMP-binding domains. However, these isoforms differ in their function, sensitivity to cAMP and cell localization.
-
Specificity of different PKA isoforms is achieved by assembling similar tertiary structures into different quaternary complexes. Such packing is driven by highly diverse and highly flexible linkers that are located in the regulatory subunits of PKA.
-
A kinase anchoring proteins (AKAPs), a large family of scaffolding proteins, bind PKA quaternary complexes and incorporate them into even bigger macromolecular complexes, which provide specificity of PKA function in space and time.
Abstract
Protein kinases are dynamic molecular switches that have evolved to be only transiently activated. Kinase activity is embedded within a conserved kinase core, which is typically regulated by associated domains, linkers and interacting proteins. Moreover, protein kinases are often tethered to large macromolecular complexes to provide tighter spatiotemporal control. Thus, structural characterization of kinase domains alone is insufficient to explain protein kinase function and regulation in vivo. Recent progress in structural characterization of cyclic AMP-dependent protein kinase (PKA) exemplifies how our knowledge of kinase signalling has evolved by shifting the focus of structural studies from single kinase subunits to macromolecular complexes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Krebs, E. G., Graves, D. J. & Fischer, E. H. Factors affecting the activity of muscle phosphorylase B kinase. J. Biol. Chem. 234, 2867–2873 (1959).
Shoji, S. et al. Complete amino acid sequence of the catalytic subunit of bovine cardiac-muscle cyclic AMP-dependent protein-kinase. Proc. Natl Acad. Sci. USA 78, 848–851 (1981).
Barker, W. C. & Dayhoff, M. O. Viral src gene products are related to the catalytic chain of mammalian cAMP-dependent protein kinase. Proc. Natl Acad. Sci. USA 79, 2836–2839 (1982).
Kannan, N. et al. Evolution of allostery in the cyclic nucleotide binding module. Genome Biol. 8, R264 (2007).
Pidoux, G. & Tasken, K. Specificity and spatial dynamics of protein kinase A signaling organized by A-kinase-anchoring proteins. J. Mol. Endocrinol. 44, 271–284 (2010).
Wong, W. & Scott, J. D. AKAP signalling complexes: focal points in space and time. Nature Rev. Mol. Cell Biol. 5, 959–970 (2004).
Newlon, M. G. et al. The molecular basis for protein kinase A anchoring revealed by solution NMR. Nature Struct. Biol. 6, 222–227 (1999).
Sarma, G. N. et al. Structure of D-AKAP2:PKA RI complex: insights into AKAP specificity and selectivity. Structure 18, 155–166 (2010).
Knighton, D. R. et al. Crystal structure of the catalytic subunit of cyclic adenosine monophosphate-dependent protein kinase. Science 253, 407–414 (1991).
Knighton, D. R. et al. Structure of a peptide inhibitor bound to the catalytic subunit of cyclic adenosine monophosphate-dependent protein kinase. Science 253, 414–420 (1991).
Hanks, S. K., Quinn, A. M. & Hunter, T. The protein kinase family: conserved features and deduced phylogeny of the catalytic domains. Science 241, 42–52 (1988).
Kornev, A. P., Haste, N. M., Taylor, S. S. & Ten Eyck, L. F. Surface comparison of active and inactive protein kinases identifies a conserved activation mechanism. Proc. Natl Acad. Sci. USA 103, 17783–17788 (2006).
Kornev, A. P., Taylor, S. S. & Ten Eyck, L. F. A helix scaffold for the assembly of active protein kinases. Proc. Natl Acad. Sci. USA 105, 14377–14382 (2008).
Taylor, S. S. & Kornev, A. P. Protein kinases: evolution of dynamic regulatory proteins. Trends Biochem. Sci. 36, 65–77 (2011). Comprehensive review on eukaryotic protein kinases, their conserved core and kinase-specific regions. Compares eukaryotic protein kinases to their evolutionary predecessors, which are eukaryote-like kinases.
Johnson, L. N. & Lewis, R. J. Structural basis for control by phosphorylation. Chem. Rev. 101, 2209–2242 (2001).
Nolen, B., Taylor, S. & Ghosh, G. Regulation of protein kinases; controlling activity through activation segment conformation. Mol. Cell 15, 661–675 (2004).
Iyer, G. H., Garrod, S., Woods, V. L. Jr & Taylor, S. S. Catalytic independent functions of a protein kinase as revealed by a kinase-dead mutant: study of the Lys72His mutant of cAMP-dependent kinase. J. Mol. Biol. 351, 1110–1122 (2005).
Keshwani, M. M. et al. Cotranslational cis-phosphorylation of the COOH-terminal tail is a key priming step in the maturation of cAMP-dependent protein kinase. Proc. Natl Acad. Sci. USA 9 Apr 2012 (doi:10.1073/pnas.1202741109).
Steichen, J. M. et al. Global consequences of activation loop phosphorylation on protein kinase A. J. Biol. Chem. 285, 3825–3832 (2010).
Steichen, J. M. et al. Structural basis for the regulation of protein kinase A by activation loop phosphorylation. J. Biol. Chem. 287, 14672–14680 (2012).
Kannan, N., Haste, N., Taylor, S. S. & Neuwald, A. F. The hallmark of AGC kinase functional divergence is its C-terminal tail, a cis-acting regulatory module. Proc. Natl Acad. Sci. USA 104, 1272–1277 (2007).
Romano, R. A., Kannan, N., Kornev, A. P., Allison, C. J. & Taylor, S. S. A chimeric mechanism for polyvalent trans-phosphorylation of PKA by PDK1. Protein Sci. 18, 1486–1497 (2009).
Bastidas, A. C. et al. Role of N-terminal myristylation in the structure and regulation of cAMP-dependent protein kinase. J. Mol. Biol. 422, 215–29 (2012).
Herberg, F. W., Zimmermann, B., McGlone, M. & Taylor, S. S. Importance of the A-helix of the catalytic subunit of cAMP-dependent protein kinase for stability and for orienting subdomains at the cleft interface. Protein Sci. 6, 569–579 (1997).
Sastri, M., Barraclough, D. M., Carmichael, P. T. & Taylor, S. S. A-kinase-interacting protein localizes protein kinase A in the nucleus. Proc. Natl Acad. Sci. USA 102, 349–354 (2005).
King, C. C., Sastri, M., Chang, P., Pennypacker, J. & Taylor, S. S. The rate of NF-κB nuclear translocation is regulated by PKA and A kinase interacting protein 1. PLoS ONE 6, e18713 (2011).
Diskar, M., Zenn, H. M., Kaupisch, A., Prinz, A. & Herberg, F. W. Molecular basis for isoform-specific autoregulation of protein kinase A. Cell. Signal. 19, 2024–2034 (2007).
Martin, B. R., Deerinck, T. J., Ellisman, M. H., Taylor, S. S. & Tsien, R. Y. Isoform-specific PKA dynamics revealed by dye-triggered aggregation and DAKAP1α-mediated localization in living cells. Chem. Biol. 14, 1031–1042 (2007). Using live cells, this study shows that two PKA isoforms have different localization and reaction to cAMP.
Berman, H. M. et al. The cAMP binding domain: an ancient signaling module. Proc. Natl Acad. Sci. USA 102, 45–50 (2005).
Weber, I. T., Steitz, T. A., Bubis, J. & Taylor, S. S. Predicted structures of cAMP binding domains of type I and II regulatory subunits of cAMP-dependent protein kinase. Biochemistry 26, 343–351 (1987).
Humphries, K. M., Deal, M. S. & Taylor, S. S. Enhanced dephosphorylation of cAMP-dependent protein kinase by oxidation and thiol modification. J. Biol. Chem. 280, 2750–2758 (2005).
Kannan, N., Taylor, S. S., Zhai, Y., Venter, J. C. & Manning, G. Structural and functional diversity of the microbial kinome. PLoS Biol. 5, e17 (2007).
Kornev, A. P., Taylor, S. S. & Ten Eyck, L. F. A generalized allosteric mechanism for cis-regulated cyclic nucleotide binding domains. PLoS Comput. Biol. 4, e1000056 (2008).
Jeffrey, P. D. et al. Mechanism of CDK activation revealed by the structure of a cyclinA–CDK2 complex. Nature 376, 313–320 (1995).
Kim, C., Xuong, N. H. & Taylor, S. S. Crystal structure of a complex between the catalytic and regulatory (RIα) subunits of PKA. Science 307, 690–696 (2005).
Kim, C., Cheng, C. Y., Saldanha, S. A. & Taylor, S. S. PKA-I holoenzyme structure reveals a mechanism for cAMP-dependent activation. Cell 130, 1032–1043 (2007).
Wu, J., Brown, S. H., von Daake, S. & Taylor, S. S. PKA type IIα holoenzyme reveals a combinatorial strategy for isoform diversity. Science 318, 274–279 (2007).
Crivici, A. & Ikura, M. Molecular and structural basis of target recognition by calmodulin. Annu. Rev. Biophys. Biomol. Struct. 24, 85–116 (1995).
Bos, J. L. Epac: a new cAMP target and new avenues in cAMP research. Nature Rev. Mol. Cell Biol. 4, 733–738 (2003).
Li, S. et al. Mechanism of intracellular cAMP sensor Epac2 activation: cAMP-induced conformational changes identified by amide hydrogen/deuterium exchange mass spectrometry (DXMS). J. Biol. Chem. 286, 17889–17897 (2011).
Brown, S. H., Wu, J., Kim, C., Alberto, K. & Taylor, S. S. Novel isoform-specific interfaces revealed by PKA RIIβ holoenzyme structures. J. Mol. Biol. 393, 1070–1082 (2009).
Dunker, A. K. et al. Intrinsically disordered protein. J. Mol. Graph. Model. 19, 26–59 (2001).
Boettcher, A. J. et al. Realizing the allosteric potential of the tetrameric protein kinase A RIα holoenzyme. Structure 19, 265–276 (2011).
Heller, W. T. et al. C subunits binding to the protein kinase A RI α dimer induce a large conformational change. J. Biol. Chem. 279, 19084–19090 (2004).
Vigil, D. et al. Conformational differences among solution structures of the type Iα, IIα and IIβ protein kinase A regulatory subunit homodimers: role of the linker regions. J. Mol. Biol. 337, 1183–1194 (2004).
Vigil, D., Blumenthal, D. K., Taylor, S. S. & Trewhella, J. Solution scattering reveals large differences in the global structures of type II protein kinase A isoforms. J. Mol. Biol. 357, 880–889 (2006).
Amieux, P. S. et al. Compensatory regulation of RIa protein levels in protein kinase A mutant mice. J. Biol. Chem. 272, 3993–3998 (1997).
Veugelers, M. et al. Comparative PRKAR1A genotype–phenotype analyses in humans with Carney complex and prkar1a haploinsufficient mice. Proc. Natl Acad. Sci. USA 101, 14222–14227 (2004).
Brandon, E. P. et al. Hippocampal long-term depression and depotentiation are defective in mice carrying a targeted disruption of the gene encoding the RIβ subunit of cAMP-dependent protein kinase. Proc. Natl Acad. Sci. USA 92, 8851–8855 (1995).
Huang, Y. Y. et al. A genetic test of the effects of mutations in PKA on mossy fiber LTP and its relation to spatial and contextual learning. Cell 83, 1211–1222 (1995).
Skalhegg, B. S. et al. Location of cAMP-dependent protein kinase type I with the TCR–CD3 complex. Science 263, 84–87 (1994).
Ilouz, R. et al. Localization and quaternary structure of PKA RIb holoenzyme. Proc. Natl Acad. Sci. USA 109, 12443–12448 (2012). Shows how small intrinsically disordered linkers in the full length PKA RIβ holoenzyme can drive packing of large unique macromolecular complexes by using almost identical building blocks.
Cadd, G. & McKnight, G. S. Distinct patterns of cAMP-dependent protein kinase gene expression in mouse brain. Neuron 3, 71–79 (1989).
Clegg, C. H., Cadd, G. G. & McKnight, G. S. Genetic characterization of a brain-specific form of the type I regulatory subunit of cAMP-dependent protein kinase. Proc. Natl Acad. Sci. USA 85, 3703–3707 (1988).
Herberg, F. W. & Taylor, S. S. Physiological inhibitors of the catalytic subunit of cAMP-dependent protein kinase: effect of MgATP on protein–protein interactions. Biochemistry 32, 14015–14022 (1993).
Bubis, J., Vedvick, T. S. & Taylor, S. S. Antiparallel alignment of the two protomers of the regulatory subunit dimer of cAMP-dependent protein kinase I. J. Biol. Chem. 262, 14961–14966 (1987).
Brennan, J. P. et al. Oxidant-induced activation of type I protein kinase A is mediated by RI subunit interprotein disulfide bond formation. J. Biol. Chem. 281, 21827–21836 (2006).
McConnachie, G., Langeberg, L. K. & Scott, J. D. AKAP signaling complexes: getting to the heart of the matter. Trends Mol. Med. 12, 317–323 (2006). Reviews large signalling complexes formed with different AKAPs that provide spatial and temporal specificity of cAMP signals.
Cummings, D. E. et al. Genetically lean mice result from targeted disruption of the RIIβ subunit of protein kinase A. Nature 382, 622–626 (1996).
Schreyer, S. A., Cummings, D. E., McKnight, G. S. & LeBoeuf, R. C. Mutation of the RIIβ subunit of protein kinase A prevents diet-induced insulin resistance and dyslipidemia in mice. Diabetes 50, 2555–2562 (2001).
Czyzyk, T. A., Sikorski, M. A., Yang, L. & McKnight, G. S. Disruption of the RIIβ subunit of PKA reverses the obesity syndrome of agouti lethal yellow mice. Proc. Natl Acad. Sci. USA 105, 276–281 (2008).
Zhang, P. et al. Structure and allostery of the PKA RIIβ tetrameric holoenzyme. Science 335, 712–716 (2012). Shows the first structure of the full length PKA tetrameric holoenzyme. This structure demonstrates how the tertiary structure can define the cooperative features of PKA activation.
Oliveria, S. F., Dell'Acqua, M. L. & Sather, W. A. AKAP79/150 anchoring of calcineurin controls neuronal L-type Ca2+ channel activity and nuclear signaling. Neuron 55, 261–275 (2007).
Violin, J. D., Zhang, J., Tsien, R. Y. & Newton, A. C. A genetically encoded fluorescent reporter reveals oscillatory phosphorylation by protein kinase C. J. Cell Biol. 161, 899–909 (2003).
Conti, M. & Beavo, J. Biochemistry and physiology of cyclic nucleotide phosphodiesterases: essential components in cyclic nucleotide signaling. Annu. Rev. Biochem. 76, 481–511 (2007).
Ni, Q. et al. Signaling diversity of PKA achieved via a Ca2+–cAMP–PKA oscillatory circuit. Nature Chem. Biol. 7, 34–40 (2011).
Stefan, E. et al. PKA regulatory subunits mediate synergy among conserved G-protein-coupled receptor cascades. Nature Commun. 2, 598 (2011).
Krebs, E. G. & Fischer, E. H. The phosphorylase-b to bhosphorylase-a converting enzyme of rabbit skeletal muscle. Biochim. Biophys. Acta 20, 150–157 (1956).
Manning, G., Whyte, D. B., Martinez, R., Hunter, T. & Sudarsanam, S. The protein kinase complement of the human genome. Science 298, 1912–1934 (2002).
Rall, T. W. & Sutherland, E. W. Formation of a cyclic adenine ribonucleotide by tissue particles. J. Biol. Chem. 232, 1065–1076 (1958).
Northup, J. K. et al. Purification of the regulatory component of adenylate cyclase. Proc. Natl Acad. Sci. USA 77, 6516–6520 (1980).
Walsh, D. A., Perkins, J. P. & Krebs, E. G. An adenosine 3′,5′-monophosphate-dependant protein kinase from rabbit skeletal muscle. J. Biol. Chem. 243, 3763–3765 (1968).
Brostrom, C. O., Corbin, J. D., King, C. A. & Krebs, E. G. Interaction of the subunits of adenosine 3′:5′-cyclic monophosphate-dependent protein kinase of muscle. Proc. Natl Acad. Sci. USA 68, 2444–2447 (1971).
Gill, G. N. & Garren, L. D. A cyclic-3′,5′-adenosine monophosphate dependent protein kinase from the adrenal cortex: comparison with a cyclic AMP binding protein. Biochem. Biophys. Res. Commun. 39, 335–343 (1970).
Tao, M., Salas, M. L. & Lipmann, F. Mechanism of activation by adenosine 3′:5′-cyclic monophosphate of a protein phosphokinase from rabbit reticulocytes. Proc. Natl Acad. Sci. USA 67, 408–414 (1970).
Haste, N. M. et al. Exploring the Plasmodium falciparum cyclic-adenosine monophosphate (cAMP)-dependent protein kinase (PfPKA) as a therapeutic target. Microbes Infect. 14, 838–850 (2012).
Gangal, M. et al. Mobilization of the A-kinase N-myristate through an isoform-specific intermolecular switch. Proc. Natl Acad. Sci. USA 96, 12394–12399 (1999).
Gisler, S. M. et al. PDZK1: II. an anchoring site for the PKA-binding protein D-AKAP2 in renal proximal tubular cells. Kidney Int. 64, 1746–1754 (2003).
Acknowledgements
Work in the authors' laboratory is supported by the Howard Hughes Medical Institute and grants from the US National Institutes of Health (NIH) (GM19301, GM34921) and DK54441 to S. S. T.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Glossary
- Phosphodiesterases
-
(PDEs). A family of proteins that break down cyclic AMP to AMP.
- AGC subfamily
-
A group of about 60 Ser/Thr kinases that share several sequence, structural and functional similarities. The name is derived from three main representatives of this kinase family, namely cyclic AMP-dependent protein kinase (PKA), PKG and PKC.
- Cyclic nucleotide-binding domains
-
(CNB domains). Conserved protein domains that bind a single molecule of cyclic AMP. Binding of cAMP causes a substantial change of the domain tertiary structure, thus providing a sensor for cAMP levels in the solution.
- A kinase anchoring proteins
-
(AKAPs). A diverse family of scaffolding proteins that have a binding site for the dimerization and docking domain in cyclic AMP-dependent protein kinase regulatory subunits.
- Cyclin-dependent kinases
-
(CDKs). Kinases that regulate the cell cycle. They are activated by a cyclin molecule, which is a small protein that is generated inside the nucleus during mitosis.
- EPAC
-
(Exchange protein directly activated by cyclic AMP). A protein that regulates the activity of RAP1, which is an important cellular regulator. EPAC has a crucial role in cell proliferation and survival.
- Intrinsically disordered regions
-
(IDRs). Parts of protein sequences that cannot accept stable secondary or tertiary structures. Disordered regions are very flexible and often serve as binding sites for other proteins.
- Small-angle X-ray scattering
-
(SAXS). Experimental technique that studies scattering of X-rays by proteins in solution (or protein solutions). Scattering profiles in the small angle range (0.1–10 degrees) provide information about general, low-resolution (50–250Å) forms of the protein.
- Small-angle neutron scattering
-
(SANS). Experimental technique that studies scattering of neutron beams by proteins in solution or protein solutions. It is similar to small-angle X-ray scattering (SAXS) but allows the use of isotope labelling of the protein sample, thus enhancing the technique.
- Nonsense-mediated mRNA decay
-
(NMD). A cellular process that controls the correct synthesis of mRNA. It detects nonsense mutations that insert an erroneous stop codon in the mRNA and thus prevents its folding through degradation.
- Long-term depression
-
(LTD). Weakening of the ability of a neuron to transmit a signal. LTD can last for hours or longer and is one of the most fundamental processes in neurophysiology.
- Long-term potentiation
-
(LTP). Strengthening of the ability of a neuron to transmit signal. This form of synaptic plasticity is thought to underlie memory formation and is characterized by synapses using leads for their long-term strengthening. LTP can last for hours or longer.
- PDZ domain
-
Common domain in a family of signalling proteins. It is named after the three first discovered representatives of this family (Postsynaptic density of 95 kDa, Discs large and zonula occludens 1).
- Agouti mice
-
Heterozygous mice that carry a mutation in the gene encoding agouti. This mutation leads to the yellow obese mouse syndrome, which is characterized by obesity, insulin resistance, hyperglycaemia, increased growth and yellow coat colour.
- Apparent activation constant
-
K¬a(cAMP). The concentration of cAMP that is required for 50% activation of cyclic AMP-dependent protein kinase.
Rights and permissions
About this article
Cite this article
Taylor, S., Ilouz, R., Zhang, P. et al. Assembly of allosteric macromolecular switches: lessons from PKA. Nat Rev Mol Cell Biol 13, 646–658 (2012). https://doi.org/10.1038/nrm3432
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm3432
This article is cited by
-
ATP-dependent conformational dynamics in a photoactivated adenylate cyclase revealed by fluorescence spectroscopy and small-angle X-ray scattering
Communications Biology (2024)
-
Differential effects of mutations of POPDC proteins on heteromeric interaction and membrane trafficking
Acta Neuropathologica Communications (2023)
-
Synaptic retrograde regulation of the PKA-induced SNAP-25 and Synapsin-1 phosphorylation
Cellular & Molecular Biology Letters (2023)
-
Proteomic analysis of neonatal mouse hearts shows PKA functions as a cardiomyocyte replication regulator
Proteome Science (2023)
-
Membranes prime the RapGEF EPAC1 to transduce cAMP signaling
Nature Communications (2023)