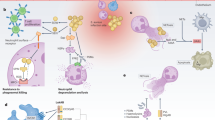
Abstract
Despite progress in our understanding of infectious disease biology and prevention, the conditions that select for the establishment and maintenance of microbial virulence remain enigmatic. To address this aspect of pathogen biology, we focus on two members of the Staphylococcus genus — Staphylococcus aureus and Staphylococcus epidermidis — and consider why S. aureus has evolved to become more virulent than S. epidermidis. Several hypotheses to explain this phenomenon are discussed and a mathematical model is used to argue that a complex transmission pathway is the key factor in explaining the evolution and maintenance of virulence in S. aureus. In the case of S. epidermidis, where skin contact affords easier transmission between hosts, high levels of virulence do not offer an advantage to this pathogen.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Gill, S. R. et al. Insights on evolution of virulence and resistance from the complete genome analysis of an early methicillin-resistant Staphylococcus aureus strain and a biofilm-producing methicillin-resistant Staphylococcus epidermidis strain. J. Bacteriol. 187, 2426–2438 (2005).
Freney, J. et al. Recommended minimal standards for description of new staphylococcal species. Subcommittee on the taxonomy of staphylococci and streptococci of the International Committee on Systematic Bacteriology. Int. J. Syst. Bacteriol. 49, 489–502 (1999).
Lowy, F. D. Staphylococcus aureus infections. N. Engl. J. Med. 339, 520–532 (1998).
von Eiff, C., Peters, G. & Heilmann, C. Pathogenesis of infections due to coagulase-negative staphylococci. Lancet Infect. Dis. 2, 677–685 (2002).
Emmerson, A. M., Enstone, J. E., Griffin, M., Kelsey, M. C. & Smyth, E. T. The second national prevalence survey of infection in hospitals — overview of the results. J. Hosp. Infect. 32, 175–190 (1996).
Jones, R. N. Global epidemiology of antimicrobial resistance among community-acquired and nosocomial pathogens: a five-year summary from the SENTRY Antimicrobial Surveillance Program (1997–2001). Semin. Respir. Crit. Care Med. 24, 121–134 (2003).
Fluit, A. C., Verhoef, J. & Schmitz, F. J. Frequency of isolation and antimicrobial resistance of Gram-negative and Gram-positive bacteria from patients in intensive care units of 25 European university hospitals participating in the European arm of the SENTRY Antimicrobial Surveillance Program 1997–1998. Eur. J. Clin. Microbiol. Infect. Dis. 20, 617–625 (2001).
Tiemersma, E. W. et al. Methicillin-resistant Staphylococcus aureus in Europe, 1999–2002. Emerg. Infect. Dis. 10, 1627–1634 (2004).
Diep, B. A. et al. Complete genome sequence of USA300, an epidemic clone of community-acquired meticillin-resistant Staphylococcus aureus. Lancet. 367, 731–739 (2006).
Takeuchi, F. et al. Whole-genome sequencing of Staphylococcus haemolyticus uncovers the extreme plasticity of its genome and the evolution of human-colonizing staphylococcal species. J. Bacteriol. 187, 7292–7308 (2005).
Kuroda, M. et al. Whole genome sequence of Staphylococcus saprophyticus reveals the pathogenesis of uncomplicated urinary tract infection. Proc. Natl. Acad. Sci. USA. 102, 13272–13277 (2005).
Baba, T. et al. Genome and virulence determinants of high virulence community-acquired MRSA. Lancet. 359, 1819–1827 (2002).
Kuroda, M. et al. Whole genome sequencing of meticillin-resistant Staphylococcus aureus. Lancet. 357, 1225–1240 (2001).
Holden, M. T. et al. Complete genomes of two clinical Staphylococcus aureus strains: evidence for the rapid evolution of virulence and drug resistance. Proc. Natl. Acad. Sci. USA. 101, 9786–9791 (2004).
Foster, T. J. Immune evasion by staphylococci. Nature Rev. Microbiol. 3, 948–958 (2005).
Lindsay, J. A. & Holden, M. T. Understanding the rise of the superbug: investigation of the evolution and genomic variation of Staphylococcus aureus. Funct. Integr. Genomics 2, 1–16 (2006).
Flannagan, S. E. & Clewell, D. B. Identification and characterization of genes encoding sex pheromone cAM373 activity in Enterococcus faecalis and Staphylococcus aureus. Mol. Microbiol. 44, 803–817 (2002).
Nakayama, J. et al. Isolation and structure of staph-cAM373 produced by Staphylococcus aureus that induces conjugal transfer of Enterococcus faecalis plasmid pAM373. Biosci. Biotechnol. Biochem. 60, 1038–1039 (1996).
Fokkens, W. J. & Scheeren, R. A. Upper airway defence mechanisms. Paediatr. Respir. Rev. 1, 336–341 (2000).
Rooijakkers, S. H., van Kessel, K. P. & van Strijp, J. A. Staphylococcal innate immune evasion. Trends. Microbiol. 13, 596–601 (2005).
Ewald, P. W. & De Leo, G. in Adaptive Dynamics of Infectious Disease (eds Dieckmann, U., Metz, J. A. J., Sabelis, M. W. & Sigmund, K.) 10–25 (Cambridge University Press, 2002).
Quinn, T. C. et al. Viral load and heterosexual transmission of Human Immunodeficiency Virus type 1. N. Engl. J. Med. 342, 921–929 (2000).
Anderson, R. M. & May, R. M. Infectious Diseases of Humans (Oxford University Press, UK, 1991).
Anderson, R. M. & May, R. M. Regulation and stability of host–parasite population interactions. J. Anim. Ecol. 47, 219–247 (1978).
Anderson, R. M. & May, R. M. The population dynamics of microparasites and their invertebrate hosts. Phil. Trans. R. Soc. 291, 451–524 (1981).
Anderson, R. M. & May, R. M. Epidemiology and genetics in the coevolution of parasites and hosts. Proc. R. Soc. Lond. B Biol. Sci. 219, 281–313 (1983).
Levin, S. A. & Pimentel, D. Selection of intermediate rates of increase in parasite–host systems. Am. Nat. 78, 308–315 (1981).
Levin, B. R. et al. in Population Biology of Infectious Disease (eds Anderson, R. M. & May, R. M.) 212–43 (Springer, New York, 1982).
Nowak, M. & May, R. M. Superinfection and the evolution of parasite virulence. Proc. Biol. Sci. 255, 81–89 (1994).
Frank, S. A. Models of parasite virulence. Q. Rev. Biol. 71, 37–78 (1996).
Day, T. Parasite transmission modes and the evolution of virulence. Evolution Int. J. Org. Evolution 55, 2389–2400 (2001).
Ewald, P. W. Host–parasite relations, vectors, and the evolution of disease severity. Ann. Rev. Ecol. Sys. 14, 465–485 (1983).
Ewald, P. W. et al. Evolutionary control of infectious disease: prospects for vectorborne and waterborne pathogens. Mem. Inst. Oswaldo Cruz. 93, 567—576 (1998).
Mackinnon, M. J. & Read, A. F. Virulence in malaria: an evolutionary viewpoint. Philos. Trans. R. Soc. Lond. B Biol. Sci. 359, 965–986 (2004).
Paul, R. E. et al. Experimental evaluation of the relationship between lethal or non-lethal virulence and transmission success in malaria parasite infections. BMC Evol. Biol. 4, (2004).
Lorange, E. A., Race, B. L., Sebbane, F. & Hinnebusch, B. J. Poor vector competence of fleas and the evolution of hypervirulence in Yersinia pestis. J. Infect. Dis. 191, 1907–1912 (2005).
Kluytmans, J., van Belkum, A. & Verbrugh, H. Nasal carriage of Staphylococcus aureus: epidemiology, underlying mechanisms, and associated risks. Clin. Microbiol. Rev. 10, 505–520 (1997).
Van den Akker, E. L. et al. Staphylococcus aureus nasal carriage is associated with glucocorticoid receptor gene polymorphisms. J. Infect. Dis. 194, 814–818 (2006).
Bauer, T. M., Ofner, E., Just, H. M., Just, H. & Daschner, F. D. An epidemiological study assessing the relative importance of airborne and direct contact transmission of microorganisms in a medical intensive care unit. J. Hosp. Infect. 15, 301–309 (1990).
Ji, G., Beavis, R. & Novick, R. P. Bacterial interference caused by autoinducing peptide variants. Science 276, 2027–2030 (1997).
Jarraud, S. et al. Exfoliatin-producing strains define a fourth agr specificity group in Staphylococcus aureus. J. Bacteriol. 182, 6517–6522 (2000).
Fleming, V. et al. Agr interference between clinical Staphylococcus aureus strains in an insect model of virulence. J. Bacteriol. 21, 7686–7688 (2006).
Dufour, P. et al. High genetic variability of the agr locus in Staphylococcus species. J. Bacteriol. 184, 1180–1186 (2002).
Lina, G. et al. Bacterial competition for human nasal cavity colonization: role of staphylococcal agr alleles. Appl. Environ. Microbiol. 69, 18–23 (2003).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
DATABASES
Entrez Genome Project
Rights and permissions
About this article
Cite this article
Massey, R., Horsburgh, M., Lina, G. et al. The evolution and maintenance of virulence in Staphylococcus aureus: a role for host-to-host transmission?. Nat Rev Microbiol 4, 953–958 (2006). https://doi.org/10.1038/nrmicro1551
Issue Date:
DOI: https://doi.org/10.1038/nrmicro1551
This article is cited by
-
Spatial disease dynamics of free-living pathogens under pathogen predation
Scientific Reports (2017)
-
DNA microarray analysis of Staphylococcus aureus causing bloodstream infection: bacterial genes associated with mortality?
European Journal of Clinical Microbiology & Infectious Diseases (2016)
-
IsaB Inhibits Autophagic Flux to Promote Host Transmission of Methicillin-Resistant Staphylococcus aureus
Journal of Investigative Dermatology (2015)
-
Molecular basis of Staphylococcus epidermidis infections
Seminars in Immunopathology (2012)
-
Staphylococcus epidermidis — wie ein Hautkeim zum Problem wird
BIOspektrum (2011)