Key Points
-
Microbial-community abundance distributions have a long 'tail' of low-abundance organisms, referred to as the rare biosphere, which often comprises the large majority of species.
-
Rare-biosphere microorganisms display specific and sometimes unique ecology and biogeography that can differ substantially from that of more abundant representatives.
-
The rare biosphere contributes to a persistent microbial seed bank, contrasting the influence of local microbial extinction and immigration.
-
Recruitment from the rare biosphere provides a broad reservoir of ecological function and resiliency (redundancy and flexibility).
-
Broad time-series studies with rich metadata will improve the study of rare-biosphere dynamics and conditionally rare taxa.
-
Study of the rare biosphere is prone to experimental artefacts (sequencing noise) and biological artefacts (dormancy and taphonomic gradients).
-
The majority of microbial diversity exists, at least transiently, in the rare biosphere.
-
Novel rare-biosphere members can be studied through cultivation, targeted phylogenetic mining and single-cell genomics.
Abstract
The profound influence of microorganisms on human life and global biogeochemical cycles underlines the value of studying the biogeography of microorganisms, exploring microbial genomes and expanding our understanding of most microbial species on Earth: that is, those present at low relative abundance. The detection and subsequent analysis of low-abundance microbial populations — the 'rare biosphere' — have demonstrated the persistence, population dynamics, dispersion and predation of these microbial species. We discuss the ecology of rare microbial populations, and highlight molecular and computational methods for targeting taxonomic 'blind spots' within the rare biosphere of complex microbial communities.
Similar content being viewed by others
Main
For most of the twentieth century, our understanding of microbial diversity relied on microscopy, cultivation and experimentation to identify the genetic and phenotypic characteristics of isolated microorganisms. Although these approaches led to many foundational discoveries, the magnitude of microbial diversity and the phylogenetic relationships among microorganisms were poorly understood. Extending from the groundbreaking work of researchers such as Woese and Fox1, small subunit ribosomal RNA (SSU rRNA) was established as a marker to survey the diversity of natural microbial communities2,3, originally by a comparison to ∼25 published full-length reference sequences4. This new molecular view of the microbial world led Norman Pace to recommend that “we undertake a representative survey of microbial diversity in the environment” (Ref. 5) on the grounds that this would answer many questions about the microbial biosphere.
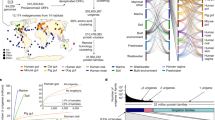
Cost and labour considerations limited the scope of early molecular studies of microbial communities (Fig. 1), and the sample sizes involved were inadequate6. Nevertheless, traditional assessments of microbial diversity were identified as being analogous to sampling of community members in traditional (macro)ecological analyses7, enabling the adoption of non-parametric estimators of diversity. This provided an empirical target for the sequencing effort that was required to evaluate and compare community diversity, which was well timed for the introduction of Sanger-based 'tag' sequencing8,9,10 and the subsequent advent of high-throughput sequencing (HTS) technologies.
The graph on the left shows a schematic rank–abundance curve, which indicates the coverage depth ranking of different experimental techniques used in taxonomic surveys of microorganisms: gel fingerprinting (including denaturing gradient gel electrophoresis), clone libraries, metagenomics and high-throughput sequencing (HTS) of PCR amplicons. The approximate threshold for the detection of rare-biosphere organisms is indicated. The graphs on the right show typical rank–abundance curves for high-diversity environments (such as soil) and low-diversity environments (such as faeces), demonstrating the differences in the size of the long 'tail' that corresponds to the rare biosphere. OTU, operational taxonomic unit.
The initial discovery of vast sequence diversity at low relative abundance involved an analysis of marine water column and sediment samples by Sogin et al.11. They generated thousands of short rRNA gene sequences (tags) and showed that even when these reads were clustered into broad categories, grouping sequences with identities that varied by as much as 10%, the number of unique sequences in each sample remained in the thousands. Such astounding genetic diversity, and presumably corresponding phenotypic diversity, is consistent with the evolution of unicellular organisms over billions of years. The absence of suitable methods for sampling high-diversity microbial habitats explained why these organisms had largely eluded detection before this work.
The discovery of a rare biosphere implicated low-abundance taxa as important contributors to assessments of α-diversity (local species diversity within sites or habitats) and β-diversity (the differentiation of species diversity across sites or habitats; see Box 1). However, because little was known about the distributions of and controls on rare-biosphere taxa, the initial recognition of such high diversity in the rare biosphere left unanswered questions about the global distributions of rare microorganisms and whether rare-biosphere taxa distribute according to Baas Becking's dictum: “everything is everywhere, but, the environment selects” (Ref. 12; see also Ref. 13).
Many rare-biosphere taxa have important ecological roles, as described below, serving as nearly limitless reservoirs of genetic and functional diversity14,15. A greater understanding of species that are adapted to life at low relative abundance will improve our understanding of microbial ecology, help to identify and resolve taxonomic 'blind spots' in the tree of life and inform emerging applications, such as gene family discovery and bioprospecting. In this Review, we discuss the ecology of rare microbial populations, the factors influencing their abundance and distribution patterns, and the methods that are used to target and explore this largely unrecognized genomic and phylogenetic diversity.
Defining the rare biosphere
Several HTS methods that were originally developed for SSU rRNA gene sequencing11,16,17,18,19 have revealed an enormous complement of low-abundance microbial taxa (Fig. 1). Defining the rare biosphere in such data has been arbitrary, and the methods used include relative abundance cut-offs (approximately 0.1%20 or 0.01%21), sequence counts in generated data sets (for example, two sequences per sample22) and empirical thresholds23,24. In practice, it is common to remove low-abundance sequences from analyses at a specific numerical or ecological threshold, such as the contribution to community dissimilarity23. Because rare species are substantial contributors to community ecology — especially temporal variability (for example, conditionally rare taxa25) — and to ecological resilience, excluding some subsets of rare taxa from analyses can blunt the interpretation of the data set. Indeed, increased weighting of the rare biosphere in β-diversity analyses (for example, by including the incidence of conditionally rare taxa25 or by using unweighted UniFrac26) has provided novel insights into microbial communities25,27,28.
Regardless of the threshold used for defining the rare biosphere, microbial community abundance distributions based on marker gene surveys (Fig. 1) typically show a long 'tail' of low-relative-abundance operational taxonomic units (OTUs). The length and shape of this tail differ depending on the diversity of the sampled community (see the graphs on the right in Fig. 1) and on the underlying species–abundance distribution29,30,31,32, which is poorly understood for most microbial communities33 (Box 1). The relative contributions of PCR and sequencing artefacts to assessments of rare microbial taxa have also been subject to debate34,35, because such artefacts can manifest, superficially, as rare OTUs, which also affects the length and shape of the long tail. Methods for filtering and analysing bona fide rare-biosphere sequences are required to understand how these artefacts manifest and to model these errors (Box 2).
Distribution, ecology and phylogeny
Microbial abundance is subject to factors that influence growth and immigration versus those that influence death and emigration36,37 (Fig. 2). Previous descriptions of the ecological relevance of rare microbial taxa indicated that rare microorganisms are susceptible to local elimination through environmental or biological processes that select for the survival of abundant organisms (as reviewed by Pedrós-Alió in Refs 20, 38), but these selective pressures are in contrast to the potential role of rare taxa as a 'seed bank' for eventual colonization under optimal conditions11,20,25. Indeed, various biotic and abiotic factors influence the abundance of rare microorganisms over time (Fig. 2). In this section, we explore the factors that contribute to the known distributions and phylogenies of rare taxa.
The species that are present in the rare biosphere may have a specific ecology and exert stronger influences on their local environment than their relative abundance would suggest. The schematic shows hypothetical temporal abundance profiles for various different members of the rare biosphere. The r-selected microorganisms that are periodically recruited from the rare biosphere (blue line) can switch between abundant and rare, depending on periodic environmental conditions such as temperature and seasonality. Such periodic resource exploitation is associated with increased susceptibility to predation. The r-selected microorganisms that are occasionally recruited from the rare biosphere (red line) persist with relatively rare abundances, responding to occasional episodic or stochastic cues such as precipitation and stress. Again, such periodic resource exploitation is associated with increased susceptibility to predation. Microbial taxa that show periodic increases in abundance but are permanently rare (green line) are adapted to exist at low relative abundance and respond with K-selected shifts in abundance, persistently avoiding predation and occupying a narrow niche. This lifestyle is associated with increased susceptibility to starvation. Permanently rare taxa (yellow line), which include possible keystone species, show persistent low-abundance distributions, occupying a narrow niche, and can escape from predation, although there is increased susceptibility to starvation. Transiently rare taxa (purple line) are occasionally rare owing to immigration. Their persistence depends on suitable conditions for survival and reproduction. Taxa exhibiting periodic or occasional recruitment from the rare biosphere can be considered to be conditionally rare taxa, as recently proposed25. The grey shading represents seasonal variations.
Variable patterns of abundance, distribution and activity. The abundance of rare microbial taxa can reflect environmental selection, similarly to the situation for dominant taxa, such as dynamics observed during a disturbance from a source of off-shore oil39 and environmental filtering for factors such as season and water depth in the study of bacterioplankton populations40. Rare-biosphere organisms can also exhibit a specific biogeography, which is distinct from that of abundant taxa and contributes to the discrimination of environmental samples. For example, an analysis of fine-scale (1 mm) diversity of sulfidic sediment samples demonstrated that maximal β-diversity was associated with rare OTUs41. Similarly, Gobet et al.42 identified non-random distributions of rare bacterial taxa in coastal sands in response to nutrient stress, although these rare fractions were not predicted to contribute to key biogeochemical processes. Bowen et al.43 demonstrated consistent rare-biosphere composition within multiple replicate salt marsh sediment samples that were highly distinct from a nearby water column sample. Recently, rare marine microeukaryotic taxa were found to display unique biogeographical distributions and were probably active and responsive to environmental change44. Often, deviations of rare taxa from broad environmental filtering are related to a narrow ecological niche, such as that suitable for copiotrophic taxa, which may be adapted to less frequent disturbances40.
In addition to a unique biogeography, rare microbial taxa can be associated with unique abundance distributions and phylogenetic compositions. Galand et al.21 sampled marine bacterial and archaeal communities from the Arctic Ocean, which revealed biogeographical patterns, higher dissimilarity to database sequences and unique species–abundance distributions associated with the rare biosphere, comprising a log-normal distribution for abundant taxa and a log-series distribution for rare taxa. This study reinforced the finding that abundance distributions can vary with ecosystem diversity and α-diversity (Box 1), influencing the size and composition of the rare biosphere. Understanding this relationship, as well as the taxonomic profiles of rare-biosphere organisms, is fundamental for their ecological study. Although the relative abundance of rare microbial taxa will not necessarily correlate with phylogenetic novelty, several recent studies have demonstrated unique phylogenetic characteristics associated with low-abundance members of microbial communities15,21,45,46.
Abundance changes in dominant OTUs can obscure population dynamics of low-abundance taxa, even though differential distributions of rare taxa may be relevant to β-diversity analyses. Discriminating life-history strategies among persistent low-abundance OTUs and organisms undergoing cyclical changes in abundance can help to identify factors that contribute to population dynamics in the rare biosphere (Fig. 2). The study of such ecological patterns requires long-term and repeated HTS of samples, coincident with the collection of appropriate metadata47, and could be undertaken within existing human microbiome and environmental monitoring projects such as the Human Microbiome Project48, the Earth Microbiome Project49 and the International Census of Marine Microbes (ICoMM).
The importance of time-series sampling and large data sets was demonstrated in a recent study of rare-biosphere dynamics. Shade et al.25 analysed 3,237 samples across 42 time-series data sets, identifying conditionally rare taxa. Contributing up to 28% of community membership, these taxa were also largely responsible for the discrimination of communities at different time points. This study also observed that variations in the abundance of rare organisms correlated with specific events, such as disturbance. Rare-biosphere organisms, which are low contributors to total community abundance but important contributors to α-diversity, represent a substantial amount of ecological potential. To best identify the causes of rare-to-prevalent community dynamics, both deep sequencing and deep sampling are recommended for uncovering the controls on rare microbial taxon abundances, reinforcing the utility of long-term sample collections. Furthermore, investigations of conditionally rare taxa coupled with rich sample metadata will help to elucidate the dynamics of rare-biosphere members.
Among collected metadata, abiotic factors — such as experimental scale and stochastic events — can influence rare-biosphere dynamics. Species observed as rare may be abundant within their microenvironment, or microniche, but be rare overall owing to a paucity of suitable microenvironments50. Such an observation is consistent with a study of bacterioplankton40, which suggested that rare community members exploit fine variations in conditions. Rare-species dynamics can be disproportionately influenced by local random events owing to low abundance and low evenness caused by such microniches. This contrast between random and deterministic processes greatly influences the ecological development and potential of local ecosystems, largely affecting recruitment from the rare biosphere (Fig. 2). This influence on ecological resiliency is negatively proportional to the incidence of guilds (groups of organisms with ecological redundancy but differing optimal conditions50).
Rare taxa may also be disproportionately active in some ecosystems, as seen in a study of marine waters proximal to Delaware Bay51. In this study, deviations from ratios of equal rRNA to rDNA (that is, the transcript to the rRNA gene) suggested either increased (>1) or decreased (<1) activity. The majority of detected OTUs displayed an approximate ratio of 1/1. Nonetheless, threefold more rare OTUs contributed to the set of high-activity OTUs, whereas a similar proportion of abundant OTUs contributed to the low-activity set51. A previous study of lake ecosystems also demonstrated this inverse relationship between relative abundance and the ratio of rRNA transcripts to rRNA genes52. Combined with observations of highly active rare-biosphere members from stream biofilm communities53 and from benthic biofilms of glacier-fed streams54, it seems as though microorganisms represented by long tails in rank–abundance plots (Fig. 1) are more active and dynamic (for example, see Fig. 2) than expected based on their abundance in single time-point DNA-based analyses.
Predation and prey avoidance. Predation and viral lysis are factors that benefit low-abundance organisms when these negative interactions are indiscriminate (Fig. 2). Viral lysis depends on encounter probabilities38,55, which are low for rare species. One suggested advantage to subsisting at low relative abundance has therefore been escape from predation, which is reflected by the killing-the-winner hypothesis56. This hypothesis also provides at least a partial explanation for the 'paradox of the plankton', which refers to the fact that plankton diversity is higher than expected given the comparatively uncomplicated resource limitations observed in aquatic environments57. Similarly, selective predation by bacterivores is a function of size, activity and abundance58,59. In such cases, organisms that are specifically adapted to persistent low abundance may have distinct strategies for predator avoidance, in contrast to transient low-abundance organisms. Predation is one factor that sustains high diversity in communities that would otherwise be shaped by competitive exclusion.
Host-specific predation and parasitism also contribute to increased bacterial species diversity. The phage–bacteria relationship is often highly specific. Analogous to the Janzen–Connell hypothesis60,61, which proposes predation as a mechanism for promoting coexistence and maintenance of high plant diversity in tropical ecosystems, phages may alter competition among bacterial species to increase coexistence and overall diversity62. However, the dynamics of this interaction are not well studied, and evidence suggests that environmental conditions are likely to play a substantial part44. Parallel studies of bacteria and the associated phage or phages will contribute to the understanding of microbial diversity and dynamics of the rare biosphere, which is important given the relationship between bacterial diversity and ecosystem function63.
Dormancy, the microbial seed bank and stored ecological potential. The existence of stable resting stages could have a substantial impact on measures of the rare biosphere52. The proportion of viable microorganisms with greatly reduced metabolic activity is variable, ranging from ∼20% in host-associated environments, such as the human gut or the rumen, to potentially the majority of microorganisms in environments such as soil64. The recovery of viable thermophilic organisms from Arctic tundra65 and Arctic marine sediment66 showed that there is a widespread oceanic distribution of bacterial endospores. Specifically, Hubert et al.66 sampled Arctic sediments and identified active fermentation and sulfate-reducing activity in sediment slurries incubated between 41 °C and 62 °C, which reflected germination and activity of endospores of the Firmicutes, including members of the sulfate-reducing genus Desulfotomaculum. The authors estimated that, based on known cell-specific sulfate-reduction rates and depth-specific activity profiles, the accumulation of hyperthermophilic spores settling at the Svalbard sediment surface from oceanic dispersal was 2 × 108 m−2 year−1, which is equivalent to approximately 55 spores cm−2 day−1. The authors of another study67 estimated similar rates for Aarhus Bay sediment and suggested that sulfate-reducing thermophiles have a viability half-life of ∼300 years. This indicates that these particular thermophilic members of the rare biosphere contribute to a taphonomic gradient of preservation and gradual decomposition (Box 2). These inactive rare-biosphere cells can be used as tracers of microbial dispersion by oceanic currents and show non-ubiquitous distributions resulting from passive dispersal68. Notably, in contrast to organisms in the Archaea or Bacteria domains, dormancy does not play as important a part in microbial eukaryotes, especially in planktonic species44.
Dormancy, when not part of a taphonomic gradient, is also an important factor in microbial seed bank dynamics69,70,71. Phases of low abundance or dormancy are often cyclical or occasional, persisting until populations are presented with suitable ecological conditions (Fig. 2). Rare-biosphere organisms can therefore respond to environmental change by acting as an active microbial seed bank with a pool of ecological potential. As part of the microbial seed bank, low-abundance organisms can contribute to myriad ecosystem dynamics and become dominant under favourable conditions25,72,73, sometimes undergoing rapid changes in abundance, such as those occuring in harmful algal blooms or during human disease.
One important implication for aquatic environments is the apparent cosmopolitan nature of phytoplankton that contribute to harmful algal blooms in response to nutrient supply. Phytoplankton populations can persist for long periods of time and initiate r-selected growth responses, or 'bloom' proliferation, under favourable conditions74. A similar assessment of rapid abundance changes, which are characteristic of algal blooms in response to specific environmental triggers, will be valuable in predicting recruitment from a rare-biosphere seed bank and will guide environmental management.
One of the most deeply sequenced global sites is the L4 marine station at the Western Channel Observatory, which has been subjected to both time-series (2003–2008) and single-site deep sequencing69,71,75. More than 95% of OTUs observed over 6 years of sampling were recovered at a single time point, when that single sample was sequenced to a depth of more than 10 million sequences71. The recent discovery of thaumarchaeotal vitamin B12 production in aquatic environments also demonstrated winter archaeal abundance at station L4, in contrast to the low relative abundance of related genes at other sampled time points181. Together, these results provided evidence that the microbial rare biosphere can serve as a recruitment pool for periodic population size increases. Furthermore, comparing a deep-sequenced L4 library to 356 diverse sample data sets from the globally distributed ICoMM data revealed substantial (44% of all taxa) cross-sample OTU overlap75, which is consistent with the idea that “everything is everywhere, but, the environment selects” (Ref. 12; see also Ref. 13). It is unclear whether this broad OTU recovery at a single site by extensive sequencing would be observed at other locations or be restricted to certain marine environments.
A persistent functional microbial seed bank was identified in studies of marine sponges of the phylum Porifera, wherein symbionts of the candidate phylum Poribacteria, among others, are abundant in host sponge tissues. HTS approaches identified Poribacteria and other putative sponge symbionts at low abundance in surrounding seawater76, implicating the rare biosphere in environmental transmission of these sponge microorganisms. In a subsequent survey of >12 million 16S rRNA gene sequences associated with ICoMM, almost half of 'sponge-specific' bacteria were detected in seawater and additional non-sponge environmental samples77. Importantly, the analysis also demonstrated the presence of sponge-specific bacteria in RNA-based sequence data, suggesting that some of these microorganisms survive and are viable outside their sponge host. Regardless, the presence of at least some bacterial species outside the host indicates possible host acquisition of the symbiont through colonization by rare-biosphere taxa rather than vertical transmission. Similarly, skin from amphibians (Rana catesbeiana and Notophthalmus viridescens) may recruit rare microorganisms from the surrounding environment in a host-specific manner78. Many skin organisms can be mutualistic in amphibians, suggesting a notable role for rare-biosphere organisms in ecosystem health.
Recruitment of rare taxa from the microbial seed bank has also been observed in freshwater environments. A microbial survey of Toolik Lake (Alaska, USA) identified a hydrological connection between upslope terrestrial communities and the lake, via headwater streams, as a source of microorganisms for surface-water communities79. In fact, four out of ten dominant taxa in Toolik Lake were present as rare-biosphere members in upslope soil samples (analogous to the red line in Fig. 2). The observation of abundant lake taxa that are potentially seeded from the surrounding catchment suggests that these taxa have at least transient roles in lake ecology and establishes a clear taxonomic interaction between terrestrial and freshwater ecosystems. As the survey's authors suggest, such terrestrial–aquatic links are perhaps not surprising given that soils contain aquatic microniches.
Despite evidence for the recruitment of rare-biosphere microorganisms from the microbial seed bank, other microorganisms may be persistent rare-biosphere members (for example, see Fig. 2). For example, a survey of Western Arctic Ocean bacteria revealed persistent interseasonal rarity of marine bacterial OTUs80. This persistent rarity suggests a K-selected growth strategy for organisms with low fecundity that occupy a narrow niche but contribute substantially to α-diversity. More recently, Hugoni et al.81 monitored archaeal communities in surface coastal waters and showed that several taxa exhibited a cyclical abundance with occasional dominance, similar to temporal distributions observed at station L4 (Refs 69, 70, 71). Other rare-biosphere archaeal taxa may have been inactive because these OTUs were less abundant, were distinct from database sequences and displayed low overall representation in corresponding RNA-based assessments. A third group of taxa were persistent rare-biosphere members that showed high activity based on RNA transcript abundances. The third group may be considered active rare-biosphere archaea, which are arguably adapted to low relative abundance subsistence. The taxonomic affiliations and seasonal dynamics of these persistently rare archaea were consistent with more abundant organisms81, suggesting that their life-history strategy is different from cyclical seed bank dynamics. The observation that rare microorganisms are phylogenetically similar to those that are abundant is intriguing.
Functional cache and ecosystem stability. Disturbance events are powerful structural influences on microbial ecosystems and community stability82. The resilience of an ecosystem — that is, the ability of a community to stabilize and regain its pre-disturbance properties — is largely influenced by species diversity. Greater biodiversity increases the likelihood that functions will be maintained by ensuring functional redundancy (that is, the insurance hypothesis83, which states that biodiversity maintains or enhances ecosystem function during environmental change). The more genetically diverse the community, the more likely it is to contain functions that counteract the disturbance event. This directly implicates low-abundance organisms in the maintenance of ecosystem health as they can act as a diverse source pool that responds to such disturbance events.
Community abundance profiles change over time as a response to disturbance25,82. A controlled water column mixing experiment, investigating oxygen and nutrient exchange in stratified water, demonstrated ecosystem resilience when communities experiencing changes in nutrient and oxygen levels recovered to control conditions within 10 days84. Similarly, in an ecosystem-level manipulation with artificial mixing of a temperate lake during peak summer stratification, microbial communities recovered in both the epilimnion and hypolimnion82. Importantly, the abundance of some rare groups (for example, Gammaproteobacteria in the hypolimnion) increased substantially in response to the disturbance event. In the same lake-level manipulation, conditionally rare taxa responded to a natural baseline disturbance (for example, seasonal mixing), a forced disturbance or both, further establishing that rare-biosphere organisms have a role in ecosystem resilience25. Some rare-biosphere members were able to exploit the forced manipulation only — that is, conditions that would not naturally occur in the lake (such as increased anoxic conditions in the hypolimnion).
The rare biosphere establishes a functional cache or resource pool for responding to disturbance events, thereby providing a mechanism for a diverse assemblage of organisms to persist in a community, independent of acquisition probabilities (that is, the reintroduction of organisms to the community). This reduces the lag in ecosystem recovery (resilience). The rare biosphere can harbour a persistent functional pool of ecological potential, which, through recruitment, may be broadly useful in promoting ecological stability. Indeed, dormancy and related seed banks are strategies for responding to temporally variable environments, contributing to community resilience52.
Keystone species. Microbial communities are core contributors to ecosystem function, with important roles in biogeochemical processes, including nutrient turnover and fixation85,86,87. Keystone species have a disproportionately large effect on their environment relative to their abundance, often serving as gatekeepers to ecosystem functions, as is the case for specialist degraders88. Inferring ecological roles from microbial community surveys poses a substantial challenge; however, co-occurrence networks show promise in identifying keystone species in HTS data89,90. The effects of keystone species can also be rooted in co-evolution through reductive genome evolution in free-living organisms, in which some members of the community maintain otherwise lost functions, supplementing the community as a whole, resulting in dependent and helper community members (described in the Black Queen hypothesis)91.
Species abundance is not always the best predictor of community contributions; rare populations can serve as keystone microbial species with disproportionately large effects on ecosystem services92. Many common or occasional members of the rare biosphere contribute to gatekeeper functions, including autotrophic organisms, such as nitrifiers; microorganisms that may have relatively narrow substrate preferences, such as methanotrophs and methylotrophs; and those that degrade low concentrations of specific chemical substrates93. Importantly, rare-biosphere members may be undetected when using traditional low-throughput approaches for linking rare-biosphere taxa with aspects of biogeochemistry94 such as culturing and molecular fingerprinting (Fig. 1).
The biogeochemical niche of keystone organisms, which is defined by specific biotic and abiotic characteristics, can be extremely rare owing to microenvironmental heterogeneity, especially in soils. For example, sulfate-reducing Desulfosporosinus sp. represented only a small proportion (0.006%) of the community in a peatland soil in Bavaria95. Nevertheless, high cell-specific reduction rates and carbon assimilation, implicated through DNA stable-isotope probing (DNA-SIP), suggested that this rare bacterium is responsible for a substantial proportion of the observed sulfate reduction and carbon flow95. These observations were consistent with previous DNA-SIP research demonstrating that presumed low-abundance Methylophaga spp. and novel uncultured Gammaproteobacteria were responsible for methanol assimilation in marine surface-water samples96.
Rare-biosphere members may also act as keystone species, with important contributions to biogeochemical cycling, but contribute substantial biomass to an environmental sample. Specifically, giant sulfur bacteria of the Beggiatoa97, Thioploca98 and Thiomargarita99 genera represent a substantial proportion of sediment biomass and sulfur cycling because of their large-diameter (0.1–0.7 mm) cells, but they have a relatively low numerical abundance100.
Microbial keystone species also have interactions that are not related to nutrient dynamics. In one experiment, the reduction of rare species in soil increased plant biomass and plant quality, as measured by nutrient concentration101. This suggests that some rare species may have a detrimental effect on plant production. However, the effect may not be exclusively negative, as there was some evidence that when all of the microorganisms were present, the levels of plant defensive compounds were higher and the nutrient concentration was lower, making the plant less suitable for herbivore predation and less prone to disease. Additionally, when rare microorganisms were present, aphid body size was decreased101.
Keystone species can also influence β-diversity disproportionately. Researchers observed that two moderately abundant species, Bacteroides fragilis and Bacteroides stercoris, had a disproportionate influence on the structure of the human gut microbiome102. They hypothesized that the abundance of keystone species contributed substantially to individual human gut microbiome uniqueness, which is consistent with contributions of conditionally rare taxa to community dissimilarity25.
The ecological potential present in species that are persistent in the rare biosphere, especially those that fill keystone roles, enables a flexible and responsive taxonomic profile that minimizes lag in ecosystems: gatekeeper species can be recruited from the rare biosphere rather than the ecosystem relying on immigration. It is easy to envision how this behaviour might be selected at the community level. Theoretically, rare microbial organisms have multiple potential strategies — some of which are unique — for responding to their environments (Fig. 2). Studying these strategies and related ecological factors will improve our understanding of microbial ecology.
Targeted exploration of the rare biosphere
Capturing microbial diversity through sequencing and culturing necessarily abstracts the underlying community — both approaches are subject to bias and artefacts. Because rare microbial species are difficult to distinguish from the background noise of sequencing error and false discovery103, what techniques can we use to survey the long tail of diversity?
Many different techniques can be used to filter signal from noise in marker gene surveys (Box 2), including interpreting error profiles in amplicon libraries, removing incorrect sequences such as chimaeras, and understanding the relationship between sequence diversity and OTUs. Much like the search for truffles104, indiscriminate surveys offer little benefit and the use of tools for directed targeting increases the probability of finding a specimen. In this section, we discuss approaches for identifying and studying the 'truffles' of microbial ecology — namely, rare-biosphere species that represent unexplored branches of microbial life ('dark matter')105, taxonomic blind spots (specific taxonomic novelty)45,46 and the potential fourth domain of life106.
Cultivation. Although HTS has supplanted clone libraries and Sanger sequencing for microbial community surveys, cultivation remains highly relevant for recovering rare microbial taxa. In contrast to direct sequencing, recovering taxa by culturing is not solely dictated by community abundance; traditional culture techniques can be useful for enriching and recovering low-abundance microorganisms. Recently, combinations of culture-dependent and culture-independent analyses uncovered members of the rare biosphere from a deep-sea coral reef107 and an apple orchard soil108. All of the isolates obtained from the reef sample were rare-biosphere members, based on the corresponding HTS data. From the apple orchard, only ∼40% of the 1,054 culture-based OTUs were recovered by pyrosequencing and most culture-based OTUs corresponded to rare members of the pyrosequencing library. Note that culturing studies do not necessarily disproportionately recover rare-biosphere members; cultivation also recovers abundant organisms109,110. Generally, the recovery of rare species by cultivation is a function of numbers. If it is correct that most bacterial species cannot be readily cultured and the rare fraction has many more microbial taxa, then cultures would be more likely to be derived from the rare fraction. As a result, cultivating novel rare-biosphere microorganisms can be particularly valuable for investigating highly diverse environments, such as soils111,112.
Although culturing provides access to rare microbial species, the specific benefit of a culture-based approach is that rare species obtained in culture enable specific physiological experiments113,114. Similar to the value of cultivating and characterizing Chthonomonas calidirosea of the phylum Armatimonadetes (formerly known as OP10)115, much can be learned about the physiology and biochemistry of rare-biosphere taxa by pure culture studies. Recovering representatives of prominent bacterial divisions and clades without cultivated representatives will benefit from focusing culturing efforts on environments harbouring those taxa; this is analogous to recent targeted single-cell sorting efforts that expanded the taxonomic and genomic breadth of the bacterial tree from selected aquatic habitat samples105. These approaches to studying rare species are convergent on the practice of microbial screening for the recovery of medical compounds.
Taxonomic blind spots: phylogenetic novelty and taxonomic dark matter. Rare and novel microorganisms provide challenging targets but are an exciting source of ecological, taxonomic and genomic discoveries. In addition to expanding the known breadth of microbial diversity, directed investigations into phylogenetic novelty will affect gene discovery and bioprospecting. A large fraction of HTS marker gene libraries is unclassified16, and most OTUs belong to the rare-biosphere fraction. Targeting taxonomic novelty in HTS data sets has uncovered new bacterial lineages45,46 and characterized microbial dark matter105. The Eukarya rare biosphere also contains substantial phylogenetic novelty. Logares et al.44 identified several distantly related clades that seemed to be exclusively or transiently rare. Similarly, targeting divergent sequences in existing data helped to identify a novel, uncultured, plastid-bearing eukaryotic lineage, the rappemonads28. These novel clades, potentially even phyla, are drawn from both poorly characterized and well-studied lineages in the tree of life. The high sequence divergence observed in some of these clades implies considerable genomic novelty, indicating that we are only scratching the surface of existing genomic diversity105. Furthermore, novel deep branches in phylogenetic analysis of metagenomic and marker gene surveys inform our understanding of early evolution and investigations into the potential fourth domain of life106. Only a small fraction of existing HTS libraries have been investigated for phylogenetic novelty and this is therefore a fruitful avenue for further taxonomic and phylogenetic research.
HTS marker gene surveys require PCR amplification, and the primers used are biased towards common or well-studied taxa. This results in taxonomic blind spots, where groups are either not amplified or amplified to artefactually low levels16. Even though analysing the rare biosphere in its entirety poses substantial challenges for discriminating signal from noise, individual low-abundance clades and phylogenetic novelty can be investigated directly. Many sequences recovered in HTS marker gene studies cannot be attributed to known taxa. Maintaining these sequences as 'unclassified' can blunt ecological inferences. Some novel and unclassified short-sequence fragments do represent legitimate microbial species that can, in some cases, significantly diverge from known lineages. Such taxa may have influential roles in an ecosystem, functioning as keystone species, or represent uncharacterized evolutionary novelty as part of biological dark matter105, despite not being major contributors to α-diversity and β-diversity.
Targeted analysis of such unclassified members of HTS surveys, which are often members of the rare biosphere, is challenging because primers must be sufficiently lineage-specific. The SSU rRNA regions targeted in HTS marker gene studies are highly variable and can contain short synapomorphic nucleotide segments, especially in unclassified groups. These regions can be exploited when targeting rare or novel lineages, which would expand on early work using data sets from serial analysis of ribosomal sequence tags116. In a pilot study using an Arctic tundra soil sample (from Alert, Nunavut), appreciable phylogenetic novelty was recovered using primers that exploit this specific mapping46. Phylogenetic targeting of this tundra V3 SSU rRNA gene library recovered near full-length sequences corresponding to seven novel taxonomic lineages. Two lineages were especially notable: a previously unknown clade within the brown rice cluster 1 (BRC1) candidate phylum117, and two clades that were both sisters to the very early branching cyanobacterium Gloeobacter, a genus containing multiple plesiomorphic traits. One observed novel cyanobacterial clade diverged by ∼10% from Gloeobacter spp. across 70% of the 16S rRNA gene and was even more distantly related to remaining Cyanobacteria, suggesting that Gloeobacteraceae are much more diverse than previously recognized. Interestingly, the sequence was nearly identical to only one other sequence in public databases, a cyanobacterium observed in Antarctica118, implying a bipolar distribution or anthropogenic dispersal. Combined with the recent Candidatus Gloeomargarita lithophora119 and non-photosynthetic candidate phylum Melainabacteria120, which is a sibling to Cyanobacteria, a more complicated evolutionary picture of the Cyanobacteria is emerging through targeted rare-biosphere analyses.
PCR introduces amplification biases121, which obscure the representation of microorganisms in the environment122,123. One way to circumvent PCR biases is by filtering marker genes from randomly sequenced environmental DNA that is generated in metagenomic studies124,125. The increasing capacity of HTS also facilitates genome assembly from metagenomic data. This is relatively trivial for common organisms with highly developed, pre-existing scaffolds, although noise would still exist in the data depending on experimental procedures (for example, there can be chimerism among closely related organisms). Efficient algorithms for genome assembly from metagenomes without closely related reference genomes will enable more thorough study of uncultivated and rare organisms. One such approach, differential coverage binning126, used multiple metagenomes generated from a single site to assemble high-quality genomes. This study assembled 31 bacterial genomes from a sequencing batch reactor, 12 of which were essentially complete, containing ≥99% of essential genes. Notably, four of the complete genomes were from the candidate phylum TM7, representing low relative abundances of 0.06–1.58% of the original metagenomic reads. As sequencing depth continues to increase, this technique may be increasingly relevant for assembling genomes from truly rare organisms.
Although it is challenging to remove artefacts from legitimate sequence data, it is clear that the rare biosphere contains sequences that represent genuine taxonomic blind spots. Mining of existing sequence data sets for such novel OTUs may reveal a wealth of taxonomic and phylogenetic information. Not only can such data-mining efforts provide useful insights into ecological phenomena, but expanding the taxonomic breadth of the tree of life will also improve sequence alignment and taxonomic assignment algorithms, thus providing increased accuracy in future studies.
Future of the rare biosphere
Studying the microbial ecology of the rare biosphere requires clear and consistent models of baseline dynamics across multiple ecosystems and time points. Correlating specific taxonomic characteristics of the rare biosphere with comprehensive sample metadata will go a long way towards identifying keystone species and understanding microbial seed bank dynamics, dormancy and biogeography. Large-scale studies involving deep sequencing75 and time-series designs72, combined with targeting novelty45,46,105, provide a useful framework for studying low-abundance organisms.
Although Baas Becking's dictum that “everything is everywhere, but, the environment selects” (Ref. 12) is not confirmed by existing data, and cannot be falsified, HTS studies have demonstrated that rare-biosphere taxa are both broadly distributed and under environmental selection. There is broad but not ubiquitous distribution and evidence of extensive worldwide passive dispersal of thermophilic endospores in marine sediments68. Additionally, this dictum suggests that broadly distributed and highly selective environments would be expected to have more similar microbial communities than adjacent sites. Indeed, Caron and Countway50 observed that deep-water (>150 m) communities of microbial eukaryotes in the North Pacific and North Atlantic were more similar across basins than to adjacent euphotic zone samples. A rare species of Gloeobacterales observed at only Arctic (Alert, NU) and Antarctic sites46 provides another example from recent research. In light of these contrasting findings from HTS surveys, perhaps it would be fitting to modify Becking's dictum in moving forward: everything can distribute everywhere, especially as rare microbial taxa, but the environment selects.
Our understanding of the metabolic and genomic potential of rare and uncharacterized organisms is limited by technical challenges: most notably, the extent of environmental metagenomes127 and the estimated sequencing coverage required to access rare organisms in some habitats128,129. Such large data sets require substantial investment in computational infrastructure and analysis techniques. Supplementing HTS technologies with existing and developing methods — such as cell sorting130,131, genome recovery132, multiple displacement amplification133, stable-isotope probing134, functional metagenomics135, targeted amplification of rare taxa136, cultivation111, co-occurrence networks137 and indicator species analysis138 — can provide a more comprehensive understanding of the roles and distributions of rare-biosphere organisms within microbial communities. Such investigations hold promise for future groundbreaking discoveries akin to pursuits of the fourth domain of life106 or to the sequencing of the large number of unknown lineages representing microbial dark matter105, in which rare-biosphere organisms have a considerable role.
Recent developments in genomics have demonstrated the feasibility of sequencing genomes from single cells139 as well as assembling rare genomes from metagenomes126. Focusing such efforts on understudied areas of the bacterial tree has increased our understanding of microbial genomic diversity105 and improved anchoring of metagenomic reads. Additionally, targeted genomics enables the comparison of persistently rare, conditionally rare and abundant taxa. Are there genomic features that delineate each group — for example, dormancy, rapid growth or predator avoidance in persistently rare species? Phylogenetic novelty correlates with the rare biosphere; therefore, our understanding of the taxonomy, genomics and ecology of rare organisms will continue to progress. We are still only scratching the surface of the rare biosphere, which is very likely to contain much more diversity than has been accessed so far. Furthermore, cultivation-based approaches have led to the recovery of many taxa140 and these approaches will continue to serve as useful tools for the recovery of organisms, genes and products for human benefit from many different environments. Indeed, the long tail of microbial diversity is more active and dynamic than previously recognized and harbours a substantial amount of ecological, taxonomic and genomic potential.
Change history
06 March 2015
On page 6 the paper by Doxey, A. C. et al. entitled 'Aquatic metagenomes implicate Thaumarchaeota in global cobalamin production'. ISME J. 9, 461–471 (2015) should have been cited at the end of the sentence ‘The recent discovery of thaumarchaeotal vitamin B12 production in aquatic environments also demonstrated winter archaeal abundance at station L4, in contrast to the low relative abundance of related genes at other sampled time points’ instead of reference 75. We have added this new reference as Ref 181 in the reference list.
References
Woese, C. R. & Fox, G. E. Phylogenetic structure of the prokaryotic domain: the primary kingdoms. Proc. Natl Acad. Sci. USA 74, 5088–5090 (1977).
Giovannoni, S. J., Britschgi, T. B., Moyer, C. L. & Field, K. G. Genetic diversity in Sargasso Sea bacterioplankton. Nature 345, 60–63 (1990).
Ward, D. M., Weller, R. & Bateson, M. M. 16S rRNA sequences reveal numerous uncultured inhabitants in a natural community. Nature 345, 63–65 (1990).
Pace, N. R., Stahl, D. A., Lane, D. J. & Olsen, G. J. The analysis of natural microbial populations by ribosomal RNA sequences. Adv. Microbial Ecol. 9, 1–55 (1986).
Pace, N. R. A molecular view of microbial diversity and the biosphere. Science 276, 734–740 (1997). An early review summarizing molecular phylogenetic methods for studying microbial ecosystems.
Neufeld, J. D. & Mohn, W. W. in Molecular Identification, Systematics, and Population Structure of Prokaryotes Ch. 7 (ed. Stackebrandt, E.) (Springer, 2006).
Hughes, J. B., Hellmann, J. J., Ricketts, T. H. & Bohannan, B. J. M. Counting the uncountable: statistical approaches to estimating microbial diversity. Appl. Environ. Microbiol. 67, 4399–4406 (2001).
Neufeld, J. D., Yu, Z., Lam, W. & Mohn, W. W. Serial analysis of ribosomal sequence tags (SARST): a high-throughput method for profiling complex microbial communities. Environ. Microbiol. 6, 131–144 (2004).
Ashby, M. N., Rine, J., Mongodin, E. F., Nelson, K. E. & Dimster-Denk, D. Serial analysis of rRNA genes and the unexpected dominance of rare members of microbial communities. Appl. Environ. Microbiol. 73, 4532–4542 (2007).
Neufeld, J. D. & Mohn, W. W. Unexpectedly high bacterial diversity in Arctic tundra relative to boreal forest soils revealed with serial analysis of ribosomal sequence tags (SARST). Appl. Environ. Microbiol. 71, 5710–5718 (2005).
Sogin, M. L. et al. Microbial diversity in the deep sea and the underexplored “rare biosphere”. Proc. Natl Acad. Sci. USA 103, 12115–12120 (2006). This study was the first to recognize and describe the rare biosphere with the use of HTS.
Baas Becking, L. B. Geobiologie of Inleiding Tot de Milieukunde (in Dutch) (WP Van Stockum & Zoon, 1934).
De Wit, R. & Bouvier, T. 'Everything is everywhere, but, the environment selects'; what did Baas Becking and Beijerinck really say? Environ. Microbiol. 8, 755–758 (2006).
Reid, A. & Buckley, M. The Rare Biosphere (American Academy of Microbiology, 2011).
Elshahed, M. S. et al. Novelty and uniqueness patterns of rare members of the soil biosphere. Appl. Environ. Microbiol. 74, 5422–5428 (2008).
Bartram, A. K., Lynch, M. D. J., Stearns, J. C., Moreno-Hagelsieb, G. & Neufeld, J. D. Generation of multi-million 16S rRNA gene libraries from complex microbial communities by assembling paired-end Illumina reads. Appl. Environ. Microbiol. 77, 3846–3852 (2011).
Lazarevic, V. et al. Metagenomic study of the oral microbiota by Illumina high-throughput sequencing. J. Microbiol. Methods 79, 266–271 (2009).
Caporaso, J. et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc. Natl Acad. Sci. USA 108 (Suppl 1), 4516–4522 (2010).
Gloor, G. et al. Microbiome profiling by Illumina sequencing of combinatorial sequence-tagged PCR products. PLoS ONE 5, e15406 (2010).
Pedrós-Alió, C. The rare bacterial biosphere. Annu. Rev. Mar. Sci. 4, 449–466 (2012).
Galand, P. E., Casamayor, E. O., Kirchman, D. L. & Lovejoy, C. Ecology of the rare microbial biosphere of the Arctic Ocean. Proc. Natl Acad. Sci. USA 106, 22427–22432 (2009). This study explores the ecology, taxonomy and biogeographic trends of rare marine microorganisms.
Qin, J. et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65 (2010).
Gobet, A., Quince, C. & Ramette, A. Multivariate Cutoff Level Analysis (MultiCoLA) of large community data sets. Nucleic Acids Res. 38, e155 (2010).
Haegeman, B. et al. Robust estimation of microbial diversity in theory and in practice. ISME J. 7, 1092–1101 (2013).
Shade, A. et al. Conditionally rare taxa disproportionately contribute to temporal changes in microbial diversity. mBio 5, e01371–14 (2014). This study identifies conditionally rare taxa and investigates their role in temporal abundance dynamics.
Lozupone, C., Lladser, M. E., Knights, D., Stombaugh, J. & Knight, R. UniFrac: an effective distance metric for microbial community comparison. ISME J. 5, 169–172 (2010).
Caporaso, J. G. et al. Moving pictures of the human microbiome. Genome Biol. 12, R50 (2011).
Kim, E. et al. Newly identified and diverse plastid-bearing branch on the eukaryotic tree of life. Proc. Natl Acad. Sci. USA 108, 1496–1500 (2011).
Curtis, T. P., Sloan, W. T. & Scannell, J. W. Estimating prokaryotic diversity and its limits. Proc. Natl Acad. Sci. USA 99, 10494–10499 (2002).
Curtis, T. P. & Sloan, W. T. Prokaryotic diversity and its limits: microbial community structure in nature and implications for microbial ecology. Curr. Opin. Microbiol. 7, 221–226 (2004).
Curtis, T. P. et al. What is the extent of prokaryotic diversity? Phil. Trans. R. Soc. B 361, 2023–2037 (2006).
Sloan, W. T., Woodcock, S., Lunn, M., Head, I. M. & Curtis, T. P. Modeling taxa-abundance distributions in microbial communities using environmental sequence data. Microb. Ecol. 53, 443–455 (2006).
Bent, S. J. & Forney, L. J. The tragedy of the uncommon: understanding limitations in the analysis of microbial diversity. ISME J. 2, 689–695 (2008).
Kunin, V., Engelbrektson, A., Ochman, H. & Hugenholtz, P. Wrinkles in the rare biosphere: pyrosequencing errors can lead to artificial inflation of diversity estimates. Environ. Microbiol. 12, 118–123 (2010).
Huse, S., Welch, D., Morrison, H. & Sogin, M. Ironing out the wrinkles in the rare biosphere through improved OTU clustering. Environ. Microbiol. 12, 1889–1898 (2010).
Barberán, A., Casamayor, E. O. & Fierer, N. The microbial contribution to macroecology. Front. Microbiol. 5, 203 (2014).
Fierer, N. & Lennon, J. T. The generation and maintenance of diversity in microbial communities. Am. J. Bot. 98, 439–448 (2011).
Pedrós-Alió, C. Dipping into the rare biosphere. Science 315, 192–193 (2007). A perspective article introducing the study of rare-biosphere organisms at the beginning of the HTS era.
Newton, R. J. et al. Shifts in the microbial community composition of Gulf Coast beaches following beach oiling. PLoS ONE 8, e74265 (2013).
Vergin, K. L., Done, B., Carlson, C. A. & Giovannoni, S. J. Spatiotemporal distributions of rare bacterioplankton populations indicate adaptive strategies in the oligotrophic ocean. Aquat. Microb. Ecol. 71, 1–13 (2013).
Youssef, N. et al. Comparison of species richness estimates obtained using nearly complete fragments and simulated pyrosequencing-generated fragments in 16S rRNA gene-based environmental surveys. Appl. Environ. Microbiol. 75, 5227–5236 (2009).
Gobet, A. et al. Diversity and dynamics of rare and of resident bacterial populations in coastal sands. ISME J. 6, 542–553 (2011).
Bowen, J. L., Crump, B. C., Deegan, L. A. & Hobbie, J. E. Salt marsh sediment bacteria: their distribution and response to external nutrient inputs. ISME J. 3, 924–934 (2009).
Logares, R. et al. Patterns of rare and abundant marine microbial eukaryotes. Curr. Biol. 24, 813–821 (2014).
Youssef, N., Steidley, B. L. & Elshahed, M. S. Novel high-rank phylogenetic lineages within a sulfur spring (Zodletone spring, Oklahoma, USA) revealed using a combined pyrosequencing/Sanger approach. Appl. Environ. Microbiol. 78, 2677–2688 (2012).
Lynch, M. D., Bartram, A. K. & Neufeld, J. D. Targeted recovery of novel phylogenetic diversity from next-generation sequence data. ISME J. 6, 2067–2077 (2012). This study used phylogenetic mining of HTS data to identify taxonomic blind spots through targeted gene recovery.
Yilmaz, P. et al. Minimum information about a marker gene sequence (MIMARKS) and minimum information about any (x) sequence (MIxS) specifications. Nature Biotech. 29, 415–420 (2011).
Turnbaugh, P. J. et al. The human microbiome project. Nature 449, 804–810 (2007).
Gilbert, J. A. et al. The Earth Microbiome Project: meeting report of the 1st EMP meeting on sample selection and acquisition at Argonne National Laboratory October 6th 2010. Stand. Genomic Sci. 3, 249–253 (2010).
Caron, D. A. & Countway, P. D. Hypotheses on the role of the protistan rare biosphere in a changing world. Aquat. Microb. Ecol. 57, 227–238 (2009).
Campbell, B. J., Yu, L., Heidelberg, J. F. & Kirchman, D. L. Activity of abundant and rare bacteria in a coastal ocean. Proc. Natl Acad. Sci. USA 108, 12776–12781 (2011). Discovery of disproportionate activity among rare marine bacteria.
Jones, S. E. & Lennon, J. T. Dormancy contributes to the maintenance of microbial diversity. Proc. Natl Acad. Sci. USA 107, 5881–5886 (2010).
Besemer, K. et al. Unraveling assembly of stream biofilm communities. ISME J. 6, 1459–1468 (2012).
Wilhelm, L. et al. Rare but active taxa contribute to community dynamics of benthic biofilms in glacier-fed streams. Environ. Microbiol. 16, 2514–2524 (2014).
Wilcox, R. M. & Fuhrman, J. A. Bacterial viruses in coastal seawater: lytic rather than lysogenic production. Mar. Ecol. Prog. Ser. 114, 35–45 (1994).
Winter, C., Bouvier, T., Weinbauer, M. G. & Thingstad, T. F. Trade-offs between competition and defense specialists among unicellular planktonic organisms: the “killing the winner” hypothesis revisited. Microbiol. Mol. Biol. Rev. 74, 42–57 (2010).
Hutchinson, G. E. The paradox of the plankton. Am. Nat. 95, 137–145 (1961).
Jürgens, K. & Matz, C. Predation as a shaping force for the phenotypic and genotypic composition of planktonic bacteria. Antonie van Leeuwenhoek 81, 413–434 (2002).
Pernthaler, J. Predation on prokaryotes in the water column and its ecological implications. Nature Rev. Microbiol. 3, 537–546 (2005).
Janzen, D. H. Herbivores and the number of tree species in tropical forests. Am. Nat. 104, 501–528 (1970).
Connell, J. H. in Dynamics of Populations (eds den Boer, P. J. & Gradwell, G.) 298–312 (Wageningen, 1971).
Brockhurst, M. A., Fenton, A., Roulston, B. & Rainey, P. B. The impact of phages on interspecific competition in experimental populations of bacteria. BMC Ecol. 6, 19 (2006).
Bell, T., Newman, J. A., Silverman, B. W., Turner, S. L. & Lilley, A. K. The contribution of species richness and composition to bacterial services. Nature 436, 1157–1160 (2005).
Lennon, J. T. & Jones, S. E. Microbial seed banks: the ecological and evolutionary implications of dormancy. Nature Rev. Microbiol. 9, 119–130 (2011).
Marchant, R., Banat, I. M., Rahman, T. J. & Berzano, M. The frequency and characteristics of highly thermophilic bacteria in cool soil environments. Environ. Microbiol. 4, 595–602 (2002).
Hubert, C. et al. A constant flux of diverse thermophilic bacteria into the cold Arctic seabed. Science 325, 1541–1544 (2009). This study demonstrates thermophilic bacterial immigration and preservation within Arctic sediment and identifies influences on community composition.
de Rezende, J. R. et al. Dispersal of thermophilic Desulfotomaculum endospores into Baltic Sea sediments over thousands of years. ISME J. 7, 72–84 (2012).
Müller, A. L. et al. Endospores of thermophilic bacteria as tracers of microbial dispersal by ocean currents. ISME J. 8, 1153–1165 (2013).
Gilbert, J. A. et al. The seasonal structure of microbial communities in the Western English Channel. Environ. Microbiol. 11, 3132–3139 (2009).
Gilbert, J. A. et al. Defining seasonal marine microbial community dynamics. ISME J. 6, 298–308 (2011).
Caporaso, J. G., Paszkiewicz, K., Field, D., Knight, R. & Gilbert, J. A. The Western English Channel contains a persistent microbial seed bank. ISME J. 6, 1089–1093 (2012). Deep sequencing of a single time point, compared to a 6-year time series, resulted in the identification of recruitment from the rare biosphere or microbial seed bank rather than extinction and recolonization.
Brazelton, W. J. et al. Archaea and bacteria with surprising microdiversity show shifts in dominance over 1,000-year time scales in hydrothermal chimneys. Proc. Natl Acad. Sci. USA 107, 1612–1617 (2010).
Sjöstedt, J. et al. Recruitment of members from the rare biosphere of marine bacterioplankton communities after an environmental disturbance. Appl. Environ. Microbiol. 78, 1361–1369 (2012).
Anderson, D. M., Cembella, A. D. & Hallegraeff, G. M. Progress in understanding harmful algal blooms: paradigm shifts and new technologies for research, monitoring, and management. Annu. Rev. Mar. Sci. 4, 143–176 (2012).
Gibbons, S. M. et al. Evidence for a persistent microbial seed bank throughout the global ocean. Proc. Natl Acad. Sci. USA 110, 4651–4655 (2013). Single-location (English Channel station L4) deep sequencing shows substantial taxonomic overlap with global marine sampling and strongly suggests that there is a rich and persistent microbial seed bank.
Webster, N. S. et al. Deep sequencing reveals exceptional diversity and modes of transmission for bacterial sponge symbionts. Environ. Microbiol. 12, 2070–2082 (2010).
Taylor, M. W. et al. “Sponge-specific” bacteria are widespread (but rare) in diverse marine environments. ISME J. 7, 438–443 (2012).
Walke, J. B. et al. Amphibian skin may select for rare environmental microbes. ISME J. 8, 2207–2217 (2014).
Crump, B. C., Amaral-Zettler, L. A. & Kling, G. W. Microbial diversity in Arctic freshwaters is structured by inoculation of microbes from soils. ISME J. 6, 1629–1639 (2012).
Kirchman, D. L., Cottrell, M. T. & Lovejoy, C. The structure of bacterial communities in the western Arctic Ocean as revealed by pyrosequencing of 16S rRNA genes. Environ. Microbiol. 12, 1132–1143 (2010).
Hugoni, M. et al. Structure of the rare archaeal biosphere and seasonal dynamics of active ecotypes in surface coastal waters. Proc. Natl Acad. Sci. USA 110, 6004–6009 (2013).
Shade, A. et al. Fundamentals of microbial community resistance and resilience. Front. Microbiol. 3, 417 (2012).
Yachi, S. & Loreau, M. Biodiversity and ecosystem productivity in a fluctuating environment: the insurance hypothesis. Proc. Natl Acad. Sci. USA 96, 1463–1468 (1999).
Shade, A. et al. Resistance, resilience and recovery: aquatic bacterial dynamics after water column disturbance. Environ. Microbiol. 13, 2752–2767 (2011).
Wardle, D. A. et al. Ecological linkages between aboveground and belowground biota. Science 304, 1629–1633 (2004).
Miki, T., Ushio, M., Fukui, S. & Kondoh, M. Functional diversity of microbial decomposers facilitates plant coexistence in a plant–microbe–soil feedback model. Proc. Natl Acad. Sci. USA 107, 14251–14256 (2010).
Montoya, J. P. et al. High rates of N2 fixation by unicellular diazotrophs in the oligotrophic Pacific Ocean. Nature 430, 1027–1032 (2004).
Ze, X., Duncan, S. H., Louis, P. & Flint, H. J. Ruminococcus bromii is a keystone species for the degradation of resistant starch in the human colon. ISME J. 6, 1535–1543 (2012).
Berry, D. & Widder, S. Deciphering microbial interactions and detecting keystone species with co-occurrence networks. Front. Microbiol. 5, 219 (2014).
Williams, R. J., Howe, A. & Hofmockel, K. Demonstrating microbial co-occurrence pattern analyses within and between ecosystems. Front. Microbiol. 5, 358 (2014).
Morris, J. J., Lenski, R. E. & Zinser, E. R. The Black Queen Hypothesis: evolution of dependencies through adaptive gene loss. mBio 3, e00036–12 (2012).
Giovannoni, S. J. & Stingl, U. Molecular diversity and ecology of microbial plankton. Nature 437, 343–348 (2005).
Pholchan, M. K., Baptista, J. C., Davenport, R. J., Sloan, W. T. & Curtis, T. P. Microbial community assembly, theory and rare functions. Front. Microbiol. 4, 68 (2013).
Bartram, A. K. et al. Exploring links between pH and bacterial community composition in soils from the Craibstone Experimental Farm. FEMS Microbiol. Ecol. 87, 403–415 (2014).
Pester, M., Bittner, N., Deevong, P., Wagner, M. & Loy, A. A 'rare biosphere' microorganism contributes to sulfate reduction in a peatland. ISME J. 4, 1591–1602 (2010).
Neufeld, J. D. et al. Stable-isotope probing implicates Methylophaga spp. and novel Gammaproteobacteria in marine methanol and methylamine metabolism. ISME J. 1, 480–491 (2007).
Jørgensen, B. B. Big sulfur bacteria. ISME J. 4, 1083–1084 (2010).
Jørgensen, B. B. & Gallardo, V. A. Thioploca spp.: filamentous sulfur bacteria with nitrate vacuoles. FEMS Microbiol. Ecol. 28, 301–313 (1999).
Schulz, H. et al. Dense populations of a giant sulfur bacterium in Namibian shelf sediments. Science 284, 493–495 (1999).
Jørgensen, B. B., Dunker, R., Grünke, S. & Røy, H. Filamentous sulfur bacteria, Beggiatoa spp., in arctic marine sediments (Svalbard, 79° N). FEMS Microbiol. Ecol. 73, 500–513 (2010).
Hol, W. H. G. et al. Reduction of rare soil microbes modifies plant–herbivore interactions. Ecol. Lett. 13, 292–301 (2010).
Fisher, C. K. & Mehta, P. Identifying keystone species in the human gut microbiome from metagenomic timeseries using sparse linear regression. PLoS ONE 9, e102451 (2014).
Reeder, J. & Knight, R. The 'rare biosphere': a reality check. Nature Methods 6, 636–637 (2009).
Mello, A., Murat, C. & Bonfante, P. Truffles: much more than a prized and local fungal delicacy. FEMS Microbiol. Lett. 260, 1–8 (2006).
Rinke, C. et al. Insights into the phylogeny and coding potential of microbial dark matter. Nature 499, 431–437 (2013). A targeted study of the genomics of novel lineages (microbial dark matter) that greatly expands genomic representation of microbial life.
Wu, D. et al. Stalking the fourth domain in metagenomic data: searching for, discovering, and interpreting novel, deep branches in marker gene phylogenetic trees. PLoS ONE 6, e18011 (2011).
Jensen, S., Lynch, M. D. J., Ray, J. L., Neufeld, J. D. & Hovland, M. Norwegian deep-water coral reefs: cultivation and molecular analysis of planktonic microbial communities. Environ. Microbiol. http://dx.doi.org/10.1111/1462-2920.12531 (2014).
Hugenholtz, P., Goebel, B. M. & Pace, N. R. Impact of culture-independent studies on the emerging phylogenetic view of bacterial diversity. J. Bacteriol. 180, 4765–4774 (1998).
Goodman, A. L. et al. Extensive personal human gut microbiota culture collections characterized and manipulated in gnotobiotic mice. Proc. Natl Acad. Sci. USA 108, 6252–6257 (2011).
Pinhassi, J., Zweifel, U. L. & Hagstroëm, A. Dominant marine bacterioplankton species found among colony-forming bacteria. Appl. Environ. Microbiol. 63, 3359–3366 (1997).
Zengler, K. et al. Cultivating the uncultured. Proc. Natl Acad. Sci. USA 99, 15681–15686 (2002).
Joseph, S. J., Hugenholtz, P., Sangwan, P., Osborne, C. A. & Janssen, P. H. Laboratory cultivation of widespread and previously uncultured soil bacteria. Appl. Environ. Microbiol. 69, 7210–7215 (2003).
Leadbetter, J. R. Cultivation of recalcitrant microbes: cells are alive, well and revealing their secrets in the 21st century laboratory. Curr. Opin. Microbiol. 6, 274–281 (2003).
Nichols, D. Cultivation gives context to the microbial ecologist. FEMS Microbiol. Ecol. 60, 351–357 (2007).
Lee, K. C.-Y. et al. Chthonomonas calidirosea gen. nov., sp. nov., an aerobic, pigmented, thermophilic micro-organism of a novel bacterial class, Chthonomonadetes classis nov., of the newly described phylum Armatimonadetes originally designated candidate division OP10. Int. J. Syst. Evol. Microbiol. 61, 2482–2490 (2011).
Neufeld, J. D., Li, J. & Mohn, W. W. Scratching the surface of the rare biosphere with ribosomal sequence tag primers. FEMS Microbiol. Lett. 283, 146–153 (2008).
Derakshani, M., Lukow, T. & Liesack, W. Novel bacterial lineages at the (sub) division level as detected by signature nucleotide-targeted recovery of 16S rRNA genes from bulk soil and rice roots of flooded rice microcosms. Appl. Environ. Microbiol. 67, 623–631 (2001).
Nakai, R. et al. Microflorae of aquatic moss pillars in a freshwater lake, East Antarctica, based on fatty acid and 16S rRNA gene analyses. Polar Biol. 35, 425–433 (2012).
Couradeau, E. et al. An early-branching microbialite cyanobacterium forms intracellular carbonates. Science 336, 459–462 (2012).
Di Rienzi, S. C. et al. The human gut and groundwater harbor non-photosynthetic bacteria belonging to a new candidate phylum sibling to Cyanobacteria. eLife 2, e01102 (2013).
Engelbrektson, A. et al. Experimental factors affecting PCR-based estimates of microbial species richness and evenness. ISME J. 4, 642–647 (2010).
Kennedy, K., Hall, M. W., Lynch, M. D. J., Moreno-Hagelsieb, G. & Neufeld, J. D. Evaluating bias of Illumina-based bacterial 16S rRNA gene profiles. Appl. Environ. Microbiol. 80, 5717–5722 (2014).
Pinto, A. J. & Raskin, L. PCR biases distort bacterial and archaeal community structure in pyrosequencing datasets. PLoS ONE 7, e43093 (2012).
Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012).
Kowalchuk, G. A., Speksnijder, A. G. C. L., Zhang, K., Goodman, R. M. & Veen, J. A. Finding the needles in the metagenome haystack. Microb. Ecol. 53, 475–485 (2007).
Albertsen, M. et al. Genome sequences of rare, uncultured bacteria obtained by differential coverage binning of multiple metagenomes. Nature Biotech. 31, 533–538 (2013).
Frisli, T., Haverkamp, T., Jakobsen, K., Stenseth, N. C. & Rudi, K. Estimation of metagenome size and structure in an experimental soil microbiota from low coverage next-generation sequence data. J. Appl. Microbiol. 114, 141–151 (2013).
Gans, J., Wolinsky, M. & Dunbar, J. Computational improvements reveal great bacterial diversity and high metal toxicity in soil. Science 309, 1387–1390 (2005).
Schloss, P. D. & Handelsman, J. Toward a census of bacteria in soil. PLoS Comput. Biol. 2, 786–793 (2006).
Marcy, Y. et al. Dissecting biological “dark matter” with single-cell genetic analysis of rare and uncultivated TM7 microbes from the human mouth. Proc. Natl Acad. Sci. USA 104, 11889–11894 (2007).
Podar, M. et al. Targeted access to the genomes of low-abundance organisms in complex microbial communities. Appl. Environ. Microbiol. 73, 3205–3214 (2007).
Stoffels, M., Ludwig, W. & Schleifer, K. H. rRNA probe-based cell fishing of bacteria. Environ. Microbiol. 1, 259–271 (1999).
Binga, E. K., Lasken, R. S. & Neufeld, J. D. Something from (almost) nothing: the impact of multiple displacement amplification on microbial ecology. ISME J. 2, 233–241 (2008).
Neufeld, J. D. et al. DNA stable-isotope probing. Nature Protoc. 2, 860–866 (2007).
Verastegui, Y. et al. Multisubstrate isotope labeling and metagenomic analysis of active soil bacterial communities. mBio 5, e01157–14 (2014).
Green, S. J. & Minz, D. Suicide polymerase endonuclease restriction, a novel technique for enhancing PCR amplification of minor DNA templates. Appl. Environ. Microbiol. 71, 4721–4727 (2005).
Barberán, A., Bates, S. T., Casamayor, E. O. & Fierer, N. Using network analysis to explore co-occurrence patterns in soil microbial communities. ISME J. 6, 343–351 (2011).
Dufrêne, M. & Legendre, P. Species assemblages and indicator species: the need for a flexible asymmetrical approach. Ecol. Monogr. 67, 345–366 (1997).
Blainey, P. C. The future is now: single-cell genomics of bacteria and archaea. FEMS Microbiol. Rev. 37, 407–427 (2013).
Shade, A. et al. Culturing captures members of the soil rare biosphere. Environ. Microbiol. 14, 2247–2252 (2012).
Whittaker, R. H. Evolution and measurement of species diversity. Taxon 21, 213–251 (1972).
Whittaker, R. H. Vegetation of the Siskiyou mountains, Oregon and California. Ecol. Monogr. 30, 279–338 (1960).
Woodcock, S. et al. Neutral assembly of bacterial communities. FEMS Microbiol. Ecol. 62, 171–180 (2007).
Bell, G. The co-distribution of species in relation to the neutral theory of community ecology. Ecology 86, 1757–1770 (2005).
Ofiţeru, I. D. et al. Combined niche and neutral effects in a microbial wastewater treatment community. Proc. Natl Acad. Sci. USA 107, 15345–15350 (2010).
Wang, J. et al. Phylogenetic beta diversity in bacterial assemblages across ecosystems: deterministic versus stochastic processes. ISME J. 7, 1310–1321 (2013).
Soininen, J. Macroecology of unicellular organisms — patterns and processes. Environ. Microbiol. Rep. 4, 10–22 (2012).
Green, J. & Bohannan, B. J. M. Spatial scaling of microbial biodiversity. Trends Ecol. Evol. 21, 501–507 (2006).
Hughes, J. B. & Bohannan, B. J. M. in Molecular Microbial Ecology Manual Vol. 1, Ch. 7.01 (eds Kowalchuk, G. A. et al.) (Springer, 2004).
Connolly, S. R. et al. Commonness and rarity in the marine biosphere. Proc. Natl Acad. Sci. USA 111, 8524–8529 (2014).
McGill, B. J. et al. Species abundance distributions: moving beyond single prediction theories to integration within an ecological framework. Ecol. Lett. 10, 995–1015 (2007).
Van de Peer, Y., Chapelle, S. & De Wachter, R. A quantitative map of nucleotide substitution rates in bacterial rRNA. Nucleic Acids Res. 24, 3381–3391 (1996).
Wang, Q., Garrity, G. M., Tiedje, J. M. & Cole, J. R. Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 73, 5261–5267 (2007).
Human Microbiome Project Consortium. A framework for human microbiome research. Nature 486, 215–221 (2012).
Klein, D. Microbial communities in nature: a postgenomic perspective. Microbe 2, 591–595 (2007).
Levy-Booth, D. J. et al. Cycling of extracellular DNA in the soil environment. Soil Biol. Biochem. 39, 2977–2991 (2007).
Nogva, H. K., Dromtorp, S., Nissen, H. & Rudi, K. Ethidium monoazide for DNA-based differentiation of viable and dead bacteria by 5′-nuclease PCR. Biotechniques 34, 804–813 (2003).
Rudi, K., Moen, B., Drømtorp, S. M. & Holck, A. L. Use of ethidium monoazide and PCR in combination for quantification of viable and dead cells in complex samples. Appl. Environ. Microbiol. 71, 1018–1024 (2005).
Nocker, A., Cheung, C.-Y. & Camper, A. K. Comparison of propidium monoazide with ethidium monoazide for differentiation of live versus dead bacteria by selective removal of DNA from dead cells. J. Microbiol. Methods 67, 310–320 (2006).
Pan, Y. & Breidt, F. Enumeration of viable Listeria monocytogenes cells by real-time PCR with propidium monoazide and ethidium monoazide in the presence of dead cells. Appl. Environ. Microbiol. 73, 8028–8031 (2007).
Bachar, A. et al. Soil microbial abundance and diversity along a low precipitation gradient. Microb. Ecol. 60, 453–461 (2010).
Wagner, A. O., Malin, C., Knapp, B. A. & Illmer, P. Removal of free extracellular DNA from environmental samples by ethidium monoazide and propidium monoazide. Appl. Environ. Microbiol. 74, 2537–2539 (2008).
Flekna, G. et al. Insufficient differentiation of live and dead Campylobacter jejuni and Listeria monocytogenes cells by ethidium monoazide (EMA) compromises EMA/real-time PCR. Res. Microbiol. 158, 405–412 (2007).
Kobayashi, H., Oethinger, M., Tuohy, M., Hall, G. & Bauer, T. Unsuitable distinction between viable and dead Staphylococcus aureus and Staphylococcus epidermidis by ethidium bromide monoazide. Lett. Appl. Microbiol. 48, 633–638 (2009).
Keer, J. & Birch, L. Molecular methods for the assessment of bacterial viability. J. Microbiol. Methods 53, 175–183 (2003).
Blazewicz, S. J., Barnard, R. L., Daly, R. A. & Firestone, M. K. Evaluating rRNA as an indicator of microbial activity in environmental communities: limitations and uses. ISME J. 7, 2061–2068 (2013).
Goodrich, J. K. et al. Conducting a microbiome study. Cell 158, 250–262 (2014).
Patin, N. V., Kunin, V., Lidström, U. & Ashby, M. N. Effects of OTU clustering and PCR artifacts on microbial diversity estimates. Microb. Ecol. 65, 709–719 (2013).
Schloss, P. D., Gevers, D. & Westcott, S. L. Reducing the effects of PCR amplification and sequencing artifacts on 16S rRNA-based studies. PLoS ONE 6, e27310 (2011). An evaluation of PCR and sequencing artefacts in SSU rRNA-based studies, including a proposed quality-filtering pipeline.
Lee, C. K. et al. Groundtruthing next-gen sequencing for microbial ecology — biases and errors in community structure estimates from PCR amplicon pyrosequencing. PLoS ONE 7, e44224 (2012).
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C. & Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27, 2194–2200 (2011).
Huber, T., Faulkner, G. & Hugenholtz, P. Bellerophon: a program to detect chimeric sequences in multiple sequence alignments. Bioinformatics 20, 2317–2319 (2004).
Quince, C., Lanzen, A., Davenport, R. J. & Turnbaugh, P. J. Removing noise from pyrosequenced amplicons. BMC Bioinformatics 12, 38 (2011).
Quince, C. et al. Accurate determination of microbial diversity from 454 pyrosequencing data. Nature Methods 6, 639–641 (2009).
Claesson, M. J. et al. Comparison of two next-generation sequencing technologies for resolving highly complex microbiota composition using tandem variable 16S rRNA gene regions. Nucleic Acids Res. 38, e200 (2010).
Eren, A. M., Vineis, J. H., Morrison, H. G. & Sogin, M. L. A filtering method to generate high quality short reads using Illumina paired-end technology. PLoS ONE 8, e66643 (2013).
Schloss, P. D. Secondary structure improves OTU assignments of 16S rRNA gene sequences. ISME J. 7, 457–460 (2013).
Nawrocki, E. P., Kolbe, D. L. & Eddy, S. R. Infernal 1.0: inference of RNA alignments. Bioinformatics 25, 1335–1337 (2009).
Pruesse, E., Peplies, J. & Glöckner, F. O. SINA: accurate high-throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics 28, 1823–1829 (2012).
Cannone, J. J. et al. The comparative RNA web (CRW) site: an online database of comparative sequence and structure information for ribosomal, intron, and other RNAs. BMC Bioinformatics 3, 1–31 (2002).
Doxey, A. C. et al. Aquatic metagenomes implicate Thaumarchaeota in global cobalamin production. ISME J. 9, 461–471 (2015).
Acknowledgements
The authors appreciate comments provided by M. L. Sogin and B. J. Butler during the preparation of this manuscript. They acknowledge support from an Early Researcher Award (Government of Ontario), the Canadian Institutes of Health Research (CIHR) and the Natural Sciences and Engineering Research Council of Canada (NSERC).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
PowerPoint slides
Glossary
- Rare biosphere
-
Although this has been an arbitrarily defined term, subject to variable abundance thresholds (for example, <0.1% of total community relative abundance), the rare biosphere may be considered as the collective of rare viable or dormant microbial taxa that are found in a given environmental sample at a specific time point.
- Bioprospecting
-
The screening of biological systems (for example, genomes or ecosystems) for novel components of industrial, commercial or scientific value.
- Conditionally rare taxa
-
Species that are rare under some conditions but can become abundant when provided with optimal growth conditions.
- Resilience
-
The ability of a community to regain functionality following a disturbance event. This ability is linked to role redundancy conferred through species diversity.
- Operational taxonomic units
-
(OTUs). Commonly used theoretical framework for relating sequence differences to discrete taxonomic entities. The OTU remains a surrogate for a taxonomic rank (for example, species) and is typically based on a specified nucleotide identity (for example, 97% for small subunit ribosomal RNA marker-gene studies).
- Biogeography
-
The study of species distribution through geographic space and time.
- Copiotrophic
-
Microorganisms that grow optimally in nutrient-rich environments and are generally adapted to rapidly exploit available resources.
- Killing-the-winner hypothesis
-
A negative frequency-dependent selection, in which abundant or active bacterial types are affected by viral pressure. This mechanism would promote the survival and viability of rare types of microorganisms, maintaining high diversity.
- Taphonomic gradient
-
A temporal gradient involving decay and fossilization of cells. In the context of studying the rare biosphere, the term refers to the detection of nucleic acids from deceased organisms.
- Microbial seed bank
-
A collection of dormant microorganisms that can respond to favourable environmental conditions.
- r-selected growth
-
Growth strategy favouring rapid reproduction and exponential population size increase. It is common in organisms such as bacteria, insects and weeds.
- K-selected growth
-
Growth strategy in which abundance tends to be stable and close to the maximum capacity in an environment. Populations undergoing K-selected growth often have larger body sizes, slower growth rates and longer life cycles.
- Epilimnion
-
Top layer in a thermally stratified lake, typically with increased dissolved oxygen concentrations.
- Hypolimnion
-
Dense bottom layer of a thermally stratified lake (below the epilimnion).
- Black Queen hypothesis
-
A theory explaining reductive evolution in free-living organisms and dependence on co-occurring microorganisms. Natural selection favours the loss of costly biological functions ('leaky' functions) as long as the function is retained by a subset of the community and provides an indispensable public good.
- Synapomorphic
-
A derived characteristic shared by two or more taxa and their most recent common ancestor.
Rights and permissions
About this article
Cite this article
Lynch, M., Neufeld, J. Ecology and exploration of the rare biosphere. Nat Rev Microbiol 13, 217–229 (2015). https://doi.org/10.1038/nrmicro3400
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro3400
This article is cited by
-
Plastiphily is linked to generic virulence traits of important human pathogenic fungi
Communications Earth & Environment (2024)
-
A comparison of two gene regions for assessing community composition of eukaryotic marine microalgae from coastal ecosystems
Scientific Reports (2024)
-
Gravesoil fungi are more sensitive than bacteria in response to precipitation
Soil Ecology Letters (2024)
-
Fungal communities are more sensitive to mildew than bacterial communities during tobacco storage processes
Applied Microbiology and Biotechnology (2024)
-
Stochastic community assembly of abundant taxa maintains the relationship of soil biodiversity-multifunctionality under mercury stress
Soil Ecology Letters (2024)