Abstract
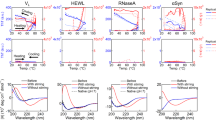
The refolding kinetics of 13 proteins have been studied in the presence of 2,2,2-trifluoroethanol (TFE). Low concentrations of TFE increased the folding rates of all the proteins, whereas higher concentrations have the opposite effect. The extent of deceleration of folding correlates closely with similar effects of guanidine hydrochloride and can be related to the burial of accessible surface area during folding. For those proteins folding in a two-state manner, the extent of acceleration of folding correlates closely with the number of local backbone hydrogen bonds in the native structure. For those proteins that fold in a multistate manner, however, the extent of acceleration is much smaller than that predicted from the data for two-state proteins. These results support the concept that for two-state proteins the search for native-like contacts is a key aspect of the folding reaction, whereas the rate-determining steps for folding of multistate proteins are associated with the reorganization of stable structure within a collapsed state or with the search for native-like interactions within less structured regions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Buck, M. Q. Rev. Biophy. 31, 297–355 (1998).
Hamada, D., Segawa, S. & Goto, Y. Nature Struct. Biol. 3, 868– 873 (1996).
Lu, H., Buck, M., Radford, S.E. & Dobson, C.M. J. Mol. Biol. 265, 112–117 ( 1997).
Kentsis, A. & Sosnick, T.R. Biochemistry 37, 14613–14622 (1998).
Chiti, F. et al. Nature Struct. Biol. 6, 380– 387 (1999).
Main, E.R.G., Fulton, K.F. & Jackson, S.E. J. Mol. Biol. 291, 429– 444 (1999).
Main, E.R.G. & Jackson, S.E. Nature Struct. Biol. 6, 831–835 (1999).
Ionescu, R.M. & Matthews, C.R. Nature Struct. Biol. 6, 304–306 (1999).
Jackson, S.E. & Fersht, A.R. Biochemistry 30, 10428–10443 (1991).
Plaxco, K.W., Simons, K.T. & Baker, D. J. Mol. Biol. 277, 985– 994 (1998).
Colón, W., Elöve, G.A., Wakem, L.P., Sherman, F. & Roder, H. Biochemistry 35, 5538–3349 (1996).
Kuwajima, K., Mitani, M. & Sugai, S. J. Mol. Biol. 206, 547– 561 (1989).
Forge, V. et al. J. Mol. Biol. 288, 673– 688 (1999).
Kuwajima, K. Proteins Struct. Funct. Genet. 6, 87– 103 (1989).
Ptitsyn, O.B. Trends Biochem. Sci. 237, 376–379 (1995).
Jennings, P.A. & Wright, P.E. Science 262, 892–896 (1993).
Radford, S.E., Dobson, C.M. & Evans, P.A. Nature 385, 302– 307 (1992).
Segawa, S. & Sugihara, M. Biopolymers 23, 2473–2488 (1984).
Myers, J.K., Pace, N. & Scholtz, J.M. Protein Sci. 4, 2138– 2148 (1995).
Makhatadze, G.I. & Privalov, P.L. J. Mol. Biol. 226, 491–505 ( 1992).
Dobson, C.M., Săli, A. & Karplus, M. Angew. Chem. Int. Edn Engl. 37, 868–893 (1998).
Guijarro, J.I., Morton, C.J., Plaxco, K.W., Campbell, I.D. & Dobson, C.M. J. Mol. Biol. 276, 657–667 (1998).
Kabsch, W. & Sander, C. Biopolymers 22, 2577–2637 (1983).
Koradi, R., Billeter, M. & Wüthrich, K. J. Mol. Graphics, 14, 51– 55 (1996).
Plaxco, K.W. et al. 37, 2529–2537 (1998).
Goto, Y. & Hamaguchi, K. J. Mol. Biol. 156, 891–910 (1982).
Walkenhorst, W.F., Green, S.M. & Roder, H. Biochemistry 36, 5795– 5805 (1997).
Kragelund, B.B. et al. J. Mol. Biol. 256, 187– 200 (1996).
Plaxco, K.W., Spitzfaden, C., Campbell, I.D. & Dobson, C.M. J. Mol. Biol. 270, 763–770 (1997).
Clarke, J. Hamill, S.J. & Johnson, C.M. J. Mol. Biol. 270, 771– 778 (1997).
Wildegger, G. & Kiefhaber, T. J. Mol. Biol. 270, 294–304 (1997).
Acknowledgements
We are very grateful to G. Ramponi, J. Clarke, Y. Goto, B.B. Kragelund, F.M. Poulsen, J. Bright, I.D. Campbell and J. Zurdo for gifts of proteins. We also acknowledge M. Buck, B.B. Kragelund, S.E. Jackson and K.W. Plaxco for valuable discussions. The Oxford Centre for Molecular Sciences is supported by the United Kingdom BBSRC, EPSRC and MRC. The Dipartimento di Scienze Biochimiche, Università di Firenze, is supported by MURST (fondi ex 40% - Folding e Misfolding di Proteine). A JSPS Postdoctoral Fellowships for Research Abroad supports D.H. This work was also supported in part by an International Research Scholars award from the Howard Hughes Medical Institute and by a programme grant from the Wellcome Trust (C.M.D.), the Grants-in-Aid for Scientific Research from the Ministry of Education, Science and Culture of Japan (M.K.) and the European Community (C.M.D., F.C. and J.I.G.).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hamada, D., Chiti, F., Guijarro, J. et al. Evidence concerning rate-limiting steps in protein folding from the effects of trifluoroethanol. Nat Struct Mol Biol 7, 58–61 (2000). https://doi.org/10.1038/71259
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/71259
This article is cited by
-
Cosolvents Induced Unfolding and Aggregation of Keyhole Limpet Hemocyanin
Cell Biochemistry and Biophysics (2014)
-
Trifluoroethanol-Induced Changes in Activity and Conformation of Manganese-Containing Superoxide Dismutase
Applied Biochemistry and Biotechnology (2012)
-
Amphiphilic α-Helical Potential: A Putative Folding Motif Adding Few Constraints to Protein Evolution
Journal of Molecular Evolution (2011)
-
The Effect of Trifluoroethanol on Tyrosinase Activity and Conformation: Inhibition Kinetics and Computational Simulations
Applied Biochemistry and Biotechnology (2010)