Abstract
Certain chemokines act as natural antagonists of human immunodeficiency virus (HIV) by blocking key viral coreceptors, such as CCR5 and CXCR4, on the surface of susceptible cells. Elucidating the structural determinants of the receptor-binding and HIV-inhibitory functions of these chemokines is essential for the rational design of derivative molecules of therapeutic value. Here, we identify the structural determinants of CCR5 recognition and antiviral activity of the CC chemokine RANTES, showing that critical residues form a solvent-exposed hydrophobic patch on the surface of the molecule. Moreover, we demonstrate that the biological function is critically dependent on dimerization, resulting in the exposure of a large (∼180 Å2), continuous hydrophobic surface. Relevant to the development of novel therapeutic approaches, we designed a retroinverted RANTES peptide mimetic that maintained both HIV- and chemotaxis-antagonistic functions.
Similar content being viewed by others
Main
Chemokines constitute a growing superfamily of short chemotactic polypeptides involved in leukocyte trafficking, development and inflammation1,2. Following the identification of the CC chemokines RANTES, MIP-1α and MIP-1β as potent natural suppressors of HIV infection3, selected seven-transmembrane-domain, G-protein-coupled chemokine receptors, such as CXCR4 and CCR5, were recognized as critical components of the cellular receptor complex for primate immunodeficiency retroviruses4. The prevalent biological variants of HIV-1 use CCR5 as a coreceptor and are specifically blocked by CCR5-binding chemokines5. Although RANTES, the most potent HIV-suppressive CC chemokine, is being investigated as a potential therapeutic agent, its inherent proinflammatory activity1,2 may limit an in vivo use. Therefore, a precise mapping of the functional regions of RANTES is essential for the development of effective derivative molecules lacking receptor-activating capability.
The NMR solution structure of the RANTES dimer6,7 shows a partially disordered N-terminal region followed by a short β-strand (β0) leading to the signature two-cysteine motif, an extended region (N-loop) ending with a 310 turn, three antiparallel β-strands (β1–3) connected by loops and a C-terminal α-helix. The physiological relevance of the dimeric structure of chemokines is still debated1,2,6,7. According to the two-site model8, two distinct regions of the chemokines participate in the interaction with chemokine receptors: the N-terminus, which is critical for receptor activation9,10,11,12, and another domain responsible for the primary docking event, which was suggested to involve the N-loop region13,14,15,16,17,18,19. The latter site is believed to play a pivotal role in the physiology of HIV infection, because neither chemokine-mediated HIV blockade20 nor the HIV-coreceptor function21 requires the signaling activity of chemokine receptors. In the present study, we identify the primary determinants of CCR5 recognition and HIV-1 blockade in RANTES, demonstrating that the antiviral and signaling activities can be uncoupled.
Peptide scanning analysis of RANTES
Peptide scanning analysis was used as a first approach to identify the antiviral determinants of RANTES. Synthetic dodecameric peptides spanning the entire sequence of the mature chemokine and overlapping by 10 amino acids (Fig. 1a) were screened for anti-HIV activity using a syncytia-inhibition assay based on the CCR5-tropic strain HIV-1BaL. Three contiguous peptides (4–6), centered on the eptameric N-loop sequence Phe 12–Pro 18, effectively blocked syncytia formation (Fig. 1b); only one additional peptide (12), centered on the β1-strand, displayed antiviral activity, although of lesser potency.
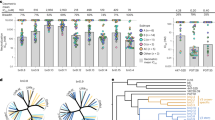
a, Sequence of overlapping dodecameric peptides spanning the entire RANTES molecule. The complete sequence of the mature protein, with the N-loop and the β-strand segments highlighted, is reported at the bottom. b, Inhibition of HIV-1-induced syncytia formation. The test was performed by coculturing PM1 cells chronically infected with HIV-1BaL, a CCR5-tropic isolate, with uninfected homologous cells. The peptides were reconstituted in water containing 66% (v/v) DMSO, at a final stock concentration of 0.8 mM, and tested at 25 μM. Syncytia formation was graded blindly by two independent observers as follows: 100% = large-size, abundant syncytia; 75% = medium-size, abundant syncytia; 50% = medium-size, rare syncytia; 25% = small-size, rare syncytia; and 0% = no syncytia. The data are representative of three independent experiments with similar outcome. c, Binding to CCR5-expressing cells, as determined by fluorocytometric analysis on a human glioma cell line (U87) persistently expressing human CCR5. The peptides were directly conjugated to biotin through a tetrapeptide linker (GSGK) at their C-terminus. Phycoerythrin (PE)-labeled streptavidin was used as a secondary reagent. The data represent the average of the mean fluorescence intensity (MFI) recorded in three independent experiments.
Next, we investigated the determinants of CCR5 recognition using a binding assay based on CCR5-expressing U87 cells. A major cluster of CCR5-binding peptides corresponded to the three contiguous N-loop dodecamers (4–6) that exhibited high antiviral activity; a second cluster of CCR5-binding peptides (12, 14, 16) was instead discontinuous (Fig. 1c). No significant binding was observed using CCR5-negative cells, except for a low-level signal with peptide 16. These findings implicated the N-loop region of RANTES as a primary receptor-binding interface. Attempts to measure the binding affinity of the N-loop-derived peptides were unsuccessful because of a paradox superinduction effect — that is, increasing binding at high competitor concentrations (not shown) — which is consistent with the formation of dimeric or multimeric peptide complexes.
Alanine scanning of the N-loop region
The role of the N-loop region in the antiviral activity of RANTES was further investigated by alanine scanning mutagenesis of the full length recombinant molecule expressed in a baculovirus system. Substitution of Phe 12 induced a dramatic loss of function in an acute HIV-1 infection assay using primary human peripheral blood mononuclear cells (PBMC), with a mean half maximal inhibitory dose (ID50) >2 logs higher than that of the wild type chemokine (Fig. 2). A functional impairment, although less pronounced, was also seen upon substitution of Tyr 14 or Ile 15, whereas downstream mutations had lesser effect.
Two CCR5-tropic HIV-1 strains were tested (HIV-1BaL and HIV-16366) in an acute infection assay using primary human PBMC. The activity is expressed as the ratio between the half-maximal inhibitory dose (ID50) of each mutant and the ID50 of the wild type molecule. The data represent the mean (± SD) of 3–6 independent experiments performed with each mutant using PBMC from different donors (the ID50 of WT RANTES ranged between 0.8 and 5.1 nM against HIV-1BaL and between 0.8 and 3.6 nM against HIV-16366). Of note, analog H23A was underestimated by ELISA quantitation, most likely as a consequence of a lack of reactivity with the monoclonal antibody used in the commercial kit. Thus, accurate quantification of this mutant was made by capillary electrophoresis and confirmed by Western blot followed by densitometric analysis. The proteins were purified as described28 and kept frozen at −80 °C in PBS supplemented with 0.1% human serum albumin.
Structure-function relationship
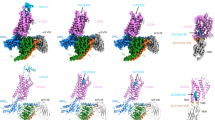
The functional mapping of RANTES was correlated with the NMR solution structure (Fig. 3a). Analysis of the surface electrostatic potential of the RANTES dimer shows that the N-loop residues critical for the antiviral activity (Phe 12, Tyr 14 and Ile 15) are displayed on the surface of the molecule, where they contribute to the formation of a large (∼180 Å2) hydrophobic, solvent-exposed patch (Fig. 3b). This structural feature is consistent with a potential role as a receptor-docking site. Phe 12 is structurally equivalent in the related CC chemokine MCP-1 to Tyr 13, a residue implicated in CCR2 binding and contributing, along with other N-loop residues, to the formation of a hydrophobic surface groove18. Similarly, the equivalent residue in MIP-1β, Phe 13, is critical for CCR5 binding and chemokine dimerization19. However, the hydrophobic patch in RANTES also encompasses a cluster of aromatic residues (Tyr 27–Tyr 29) lying at the center of the β1-strand peptide that displayed antiviral activity (Fig. 1b). Thus, the structural analysis suggests that both the N-loop and β1-strand residues contribute to the formation of the putative receptor interface.
a, Ribbon model of the RANTES dimer displaying the location of selected residues (Phe 12–Ile 15 and Tyr 27–Tyr 29) that were identified as critical for CCR5 recognition and anti-HIV activity. b, Mapping of the surface electrostatic potential of RANTES calculated on the molecular surface of the dimeric form. A large, solvent-exposed hydrophobic patch encompassing two clusters of aromatic residues within the N-loop and the β1-strand regions is displayed. The orientation of the RANTES dimer is approximately the same as in panel (a). Electrostatic potentials were calculated with GRASP29.
Experimental validation of the model
To provide an experimental basis for the model of receptor-interactive site derived from structural analysis, we designed synthetic peptides encompassing both the N-loop and β1-strand residues that form the hydrophobic patch (Fig. 3b). In the acute infection assay, the anti-HIV potency of a 19-mer spanning both the N-loop and the β1-strand aromatic cluster (AcCys 11–Tyr 29NH 2) was significantly higher than that of the N-loop-derived dodecamer AcCys 11–Ala 22NH2 (mean ID50 = 8.9 ± 3.8 μM versus 51.5 ± 6.3 μM; P < 0.0001) (Fig. 4a). Two shorter peptides lacking the β1-strand aromatic cluster (AcCys 11–Lys 25NH2 and AcCys 11–Glu 26NH2) were less effective (mean ID50 = 48.1 ± 6.8 μM and 44.7 ± 3.3 μM, respectively), confirming the importance of such cluster for the antiviral activity. We also designed a cyclic 24-mer spanning the entire region between the second and third cysteines (cyCys 11–Cys 34): such peptide inihibited HIV more potently than the original dodecamer (mean ID50 = 29.8 ± 1.6 μM), although less than the 19-mer AcCys 11–Tyr 29NH2. No effect was seen with a colinear control cyclic peptide derived from the CXCR4 ligand SDF-1 (SDF1 cyCys 12–Cys 35). These findings conclusively demonstrated that the aromatic cluster within the β1 strand of RANTES is crucial for the antiviral activity, confirming that this region, along with the N-loop, is involved in the CCR5 receptor interface.
a, Inhibition of acute HIV-1 infection by RANTES-derived synthetic peptides. In vitro-activated human PBMC were infected with cell-free HIV-1BaL in the presence or absence of the synthetic peptides at the indicated concentrations. The peptides were maintained at the original concentration throughout the culture period. Extracellular HIV-1 p24 antigen release was measured at day five postinfection. The results denote the percent inhibition of HIV-1 expression, relative to that measured of control, untreated cultures in each experiment. The data represent the mean (± SEM) from 3–6 experiments performed for each peptide using PBMC from different donors. A control retroinverted 20-mer peptide derived from the rat acetylcholine receptor α-subunit, residues 97–116 (Ac-IHGTYDLLVRTFKVIAFDGD-NH2), showed no inhibitory activity. b, Agonistic and antagonistic activity of RANTES-derived peptides on primary human lymphocyte chemotaxis. The activity of the full-length chemokine, used at 25 nM, is shown for reference. The test was performed with IL-2-activated primary human lymphocytes, which express high levels of surface CCR5. Expression of CCR5 was monitored at two-day intervals and the cells were analyzed for chemotaxis between days seven and 14, during which the level of CCR5 was high on most of the cells. The chemotactic index was calculated as the ratio between the number of cells migrated toward the chemoattractant and those spontaneously migrated through the membrane in the absence of chemotactic stimuli. The data represent the mean of duplicate tests and are representative of three experiments with similar outcome.
Uncoupling HIV blockade and receptor activation
Next, we investigated whether the determinants of CCR5 recognition and anti-HIV activity can be uncoupled from those involved in receptor activation. None of the peptides tested (AcCys 11–Ala 22NH2, cyCys 11–Cys 34 and AcCys 11–Tyr 29NH2) were able to induce CCR5-mediated intracellular calcium mobilization (not shown) or lymphocyte chemotaxis (Fig. 4b). Moreover, all three peptides effectively antagonized RANTES-elicited lymphocyte chemotaxis with a relative potency comparable to that seen in HIV-inhibition assays. By contrast, the control SDF-1 peptide showed neither agonistic nor antagonistic activity. These data indicated that, despite a direct interaction with CCR5, determinants within the RANTES N-loop/β1-strand are not sufficient for inducing receptor activation.
Biological activity of a retroinverted peptide mimetic
Peptide mimetics with inverted amino acid chirality are inherently resistant to protease-mediated digestion and thus represent suitable candidates for therapeutic applications. We synthesized an inverted mimetic of the N-loop/β1-strand 19-mer with a reversed direction of the peptide bonds in order to preserve the original side chain topochemistry22. The retroinverted peptide (AcdTyr 29–dCys 11NH2) effectively inhibited HIV-1 envelope-mediated cell fusion with a potency (mean ID50 = 13.3±2.7 μM) comparable to that of its l-counterpart (Fig. 4a); moreover, it antagonized RANTES-elicited chemotaxis (Fig. 4b). By contrast, no effect was seen with a control retroinverted peptide with similar amino acid composition and hydrophobicity value (data not shown). These biological activities confirmed the prediction that retroinversion resulted in the preservation of the original spatial orientation of the peptide lateral chains.
Critical role of dimerization for the biological activity
As all the bioactive peptides derived from the RANTES N-loop contained at least one Cys residue with a free sulphydryl group, we hypothesized that they could undergo spontaneous dimerization. Thus, after reconstitution and storage, we reanalyzed by mass spectrometry the entire series of overlapping peptides that were used for scanning analysis (Fig. 1a). Two distinct peaks, with molecular masses corresponding to the monomeric and dimeric forms, were observed with all Cys-containing peptides, indicating that partial dimerization had occurred, whereas no dimeric forms were detectable with peptides lacking Cys residues (data not shown). Quantitative analysis by reverse phase (RP)-HPLC demonstrated variable rates of dimerization (range: 0.10–0.28); in particular, significant levels were detected with all the HIV-inhibitory peptides (4, 5, 6 and 12; dimerization rates: 0.18, 0.27, 0.23 and 0.20, respectively).
To definitively elucidate the biological relevance of the dimeric structure, both stably monomeric and stably dimerized forms were obtained for two bioactive peptides (AcCys 11–Glu 26NH2 and AcCys 11–Tyr 29NH2). The monomeric peptides were virtually ineffective against HIV-1 in a cell fusion assay (mean ID50 = >160 μM) (Fig. 5); in sharp contrast, the fully dimerized peptides were highly inhibitory, with a greater specific activity (mean ID50 = 1.8 ± 0.7 μM and 15.7 ± 1.4 μM, respectively) compared to their spontaneously oxidized (partially dimerized) counterparts. The specific activity of these peptides remained markedly lower than that of full length RANTES in the same assay (mean ID50 = 5.8 ± 2.7 nM); however, this discrepancy is not unexpected in light of the extreme flexibility of linear peptides in solution, which makes the bioactive conformation an infrequent event. In other experiments (not shown), stably dimerized peptide AcCys 11–Tyr 29NH2 failed to induce CCR5-mediated intracellular calcium mobilization and antagonized RANTES-elicited chemotaxis. These results demonstrated that the biological function is critically dependent on a dimeric structure, suggesting that the extended hydrophobic surface formed by the coalescence of two monomeric patches (Fig. 3b) is essential for the functional interaction of RANTES with CCR5.
Comparative analysis of the anti-HIV-1 activity of stably monomeric (solid symbols) and stably dimerized (empty symbols) RANTES-derived peptides. An HIV-1 envelope-mediated cell fusion assay based on vaccinia technology, modified from the one originally developed by Berger and coworkers4, was employed. The human CD4+ T-cell clone PM1, chronically infected with HIV-1BaL, was used as an effector; uninfected homologous cells were used as targets. The level of envelope-mediated cell fusion, as revealed by β-galactosidase enzyme activity, was compared with the level detected in the absence of inhibitors. The data represent the mean (± SEM) from 3–6 independent experiments performed with each peptide. Full length recombinant RANTES was a highly effective inhibitor in this assay, with a mean ID50 of 5.8 ± 2.7 nM. CMC = S-carboxy-methyl-cysteine.
Conclusions and prospects
The development of new drugs with the ability to block HIV-1 entry into cells constitutes a primary goal of the current efforts to devise effective AIDS therapies23. However, the rational design of effective inhibitors of the two major HIV-1 coreceptors, CCR5 and CXCR4, has been hampered by the lack of high-resolution structural information because of difficulties inherent in performing crystallization or NMR studies on proteins belonging to the seven-transmembrane-domain, G-protein-coupled receptor superfamily. We have identified the primary determinants of CCR5 recognition and HIV-1 blockade in RANTES, formally demonstrating that the antiviral and signaling functions can be uncoupled. The structural features of such determinants are unique, with critical residues forming a large, solvent-exposed hydrophobic patch. Even more strikingly, we found that the biologically active structure is dimeric, resulting in the formation of a continuous ∼180 Å2 hydrophobic surface that seems to constitute the primary CCR5-binding interface. This finding corroborates the concept that chemokines may act as dimers under physiological conditions6,7, suggesting that chemokine receptors might also dimerize on the cellular surface, as documented for other G-protein-coupled receptors24,25. In keeping with this hypothesis, heterodimerization of chemokine receptors has been reported26; however, it is unclear whether homodimerization may also occur and whether it constitutes a general mode of interaction of chemokine receptors with their nominal ligands.
Our data open new perspectives for the development of safe and effective HIV-entry inhibitors. A first spin-off of potential therapeutic relevance was the design of a retroinverted peptide that maintained the biological properties of its l-counterpart, including not only the anti-HIV activity but also a potential anti-inflammatory effect by a blockade of RANTES-elicited chemotaxis. Considering the peculiar resistance of d-peptides to protease-mediated degradation, this molecule may represent a lead compound for antiviral drug development. Based on the available structural information, derivative compounds featuring a substantial reduction in size and increase in potency might be rationally designed. Finally, novel structural information may derive from the analysis of molecular complexes formed in solution between the RANTES peptides described herein and bioactive peptides derived from CCR5.
Methods
Peptides.
Peptides were synthesized by standard solid phase protocols using Fmoc chemistry and purified by RP-HPLC to >95% purity. Peptide scanning was performed using overlapping MultipinTM peptides (Chiron) that were acylated and amidated at the extremities and conjugated to biotin through a tetrapeptide linker (GSGK) at the C-terminus. For stable dimerization, peptides were incubated overnight in 50% (v/v) DMSO (Sigma) in water; DMSO was then removed by lyophylization, and the dimerized peptides were purified by RP-HPLC. The final stocks were >97% pure and contained no free sulphydryl groups, as assessed by Ellman test27. To obtain stable monomeric peptides, sulphydryl groups were alkylated by treatment with iodoacetamide (Sigma) for 1 h under continuous stirring in 50 mM phosphate buffer, pH 8; following RP-HPLC purification, the final stocks were >97% monomeric. Spontaneous dimerization of MultipinTM peptides was documented by MALDI-TOF mass spectrometry and quantitated by RP-HPLC using stably monomeric and stably dimerized peptides as a reference. The dimerization rate was calculated as the ratio between the amount of dimer and the total amount of peptide.
HIV-1 inhibition.
The syncytia assay was performed by coculturing chronically infected PM1BaL cells with uninfected homologous cells at a ratio of 1:5 in 200 μl of RPMI 1640 medium (Gibco) supplemented with 10% fetal bovine serum (FBS) (Hyclone) in duplicate wells of flat bottom 96-well plates (Nunc). Syncytia formation was scored visually after 18 h by two independent observers. To perform the acute infection assay, normal human PBMC was isolated from healthy blood donors and then infected with HIV-1BaL or with HIV-16366 as described28. The cell fusion assay was performed using a modification of the test, based on vaccinia technology, originally developed by Berger and coworkers4. In the modified assay, chronically infected PM1BaL cells were used as effectors and uninfected PM1 cells as targets.
CCR5 binding.
Binding to CCR5 was evaluated by fluorocytometry using U87 cells expressing CCR5 (a gift of D.R. Littman). Trypsinized cells were treated for 2 h at 37 °C with heparinase III (1 U / 106 cells) (Sigma). Then, the cells were washed with cold PBS supplemented with 1% FBS, exposed to biotin-labeled peptides for 30 min at 4 °C, washed again and then incubated for 30 min with phycoerythrin (PE)-conjugated streptavidin (Sigma). Streptavidin alone was used to establish the level of spontaneous fluorescence. The analysis was conducted on a FacScan instrument (Becton Dickinson).
Recombinant RANTES analogs.
Wild type RANTES and mutated analogs were expressed in a baculovirus system as described28. The primers used to introduce specific mutations, in combination with vector-derived primers, were:

Chemotaxis.
The chemotactic activity was assayed on IL-2-stimulated human lymphocytes in duplicate 24-well, 5 μm pore-size TranswellTM chambers (Costar) as described28. To test agonistic activities, chemokines or peptides in the lower chamber were diluted in 500 μl of RPMI containing 0.3% human serum albumin; the cells (1.5 × 105) were added to the upper chamber in 250 ml of RPMI. To test antagonistic effects, the peptides were mixed with the cells prior to addition to the upper chamber.
References
Baggiolini, M., Dewald, B. & Moser, B. Ann. Rev. Immunol. 55, 97–179 (1994).
Schall, T.J. & Bacon, K.B. Curr. Opin. Immunol. 6, 865–873 (1994).
Cocchi, F. et al. Science 270, 1811–1815 (1995).
Berger, E.A., Murphy, P.M. & Farber, J.M. Ann. Rev. Immunol. 17, 657–700 (1999).
Lusso, P. Virology 273, 228–240 (2000).
Skelton, N.J., Aspiras, F., Ogez, J. & Schall, T.J. Biochemistry 34, 5329–5342 (1995).
Chung, C., Cooke, R.M., Proudfoot, A.E. & Wells, T.N.C. Biochemistry 34, 9307–9314 (1995).
Siciliano, S.J. et al. Proc. Natl. Acad. Sci. USA 91, 1214–1218 (1994).
Clark-Lewis, I. et al. J. Leuk. Biol. 57, 703–711 (1995).
Gong, J.-H., Uguccioni, M., Dewald, B., Baggiolini, M. & Clark-Lewis, I. J. Biol. Chem. 271, 10521–10527 (1996).
Proudfoot, A.E. et al. J. Biol. Chem. 271, 2599–2605 (1996).
Arenzana-Seisdedos, F. et al. Nature 383, 400 (1996).
Clark-Lewis, I., Dewald, B., Loetscher, M., Moser, B. & Baggiolini, M. J. Biol. Chem. 269, 16075–16081 (1994).
Schraufstätter, I.U., Ma, M., Oades, Z.G., Barritt, D.S. & Cochrane, C.G. J. Biol. Chem. 270, 10428–10431 (1995).
Lowman, H.B. et al. J. Biol. Chem. 271, 14344–14352 (1996).
Pakianathan, D.R., Kuta, E.G., Artis, D.R., Skelton, N.J. & Hebert, C.A. Biochemistry 36, 9642–8648 (1997).
Crump, M.P. et al. EMBO J. 16, 6996–7007 (1997).
Hemmerich, S. et al. Biochemistry 38, 13013–13025 (1999).
Laurence, J.S., Blanpain, C., Burgner, J.W., Parmentier, M. & LiWang, P.J. Biochemistry 39, 3401–3409 (2000).
Cocchi, F. et al. Nature Med. 2, 1244–1247 (1996).
Farzan, M. et al. J. Biol. Chem. 272, 6854–6857 (1997).
Chorev, M. & Goodman, M. Trends Biotechnol. 13, 438–445 (1995).
Ferrer, M. et al. Nature Struct. Biol. 6, 953–960 (1999).
Romano, C., Yang, W.L. & O' Malley, K.L. J. Biol. Chem. 271, 28612–28616 (1996).
Jordan, B.A. & Devi, L.A. Nature 399, 697–700 (1999).
Mellado, M., Rodríguez-Frade, J.M., Vila-Coro, A.J., De Ana A.M. & Martínez, A.C. Nature 400, 723–724 (1999).
Ellman, G.L. Arch. Biochem. Biophys. 82, 70–77 (1959).
Polo, S., et al. Eur. J. Immunol. 30, 3190–3198, 2000.
Nichols, A.J., Sharp, K. & Honig, B. Proteins 11, 281–296 (1991).
Acknowledgements
We thank D.R. Littman for CCR5-transfected U87 cells (Ghost-CD4 cells), E.A. Berger for vaccinia virus vectors, E. Frittoli for technical assistance and S. Laus for editorial assistance. This work was supported by grants from the ISS AIDS Program, Rome, Italy.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Nardese, V., Longhi, R., Polo, S. et al. Structural determinants of CCR5 recognition and HIV-1 blockade in RANTES. Nat Struct Mol Biol 8, 611–615 (2001). https://doi.org/10.1038/89653
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/89653
This article is cited by
-
Rational CCL5 mutagenesis integration in a lactobacilli platform generates extremely potent HIV-1 blockers
Scientific Reports (2018)
-
Protein and oligonucleotide delivery systems for vaginal microbicides against viral STIs
Cellular and Molecular Life Sciences (2015)
-
Elucidating a Key Anti-HIV-1 and Cancer-Associated Axis: The Structure of CCL5 (Rantes) in Complex with CCR5
Scientific Reports (2014)
-
Biophysical and structural investigation of bacterially expressed and engineered CCR5, a G protein-coupled receptor
Journal of Biomolecular NMR (2013)
-
Foot-and-mouth disease virus leader proteinase inhibits dsRNA-induced RANTES transcription in PK-15 cells
Virus Genes (2011)