Abstract
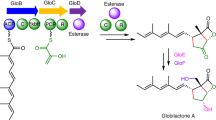
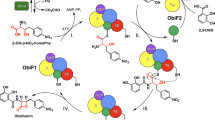
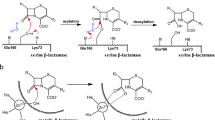
The enzyme β-lactam synthetase (β-LS) catalyzes the formation of the β-lactam ring in clavulanic acid, a clinically important β-lactamase inhibitor. Whereas the penicillin β-lactam ring is generated by isopenicillin N synthase (IPNS) in the presence of ferrous ion and dioxygen, β-LS uses ATP and Mg2+ as cofactors. According to sequence alignments, β-LS is homologous to class B asparagine synthetases (AS-Bs), ATP/Mg2+-dependent enzymes that convert aspartic acid to asparagine. Here we report the first crystal structure of a β-LS. The 1.95 Å resolution structure of Streptomyces clavuligerus β-LS provides a fully resolved view of the active site in which substrate, closely related ATP analog α,β-methyleneadenosine 5′-triphosphate (AMP-CPP) and a single Mg2+ ion are present. A high degree of substrate preorganization is observed. Comparison to Escherichia coli AS-B reveals the evolutionary changes that have taken place in β-LS that impede interdomain reaction, which is essential in AS-B, and that accommodate β-lactam formation. The structural data provide the opportunity to alter the synthetic potential of β-LS, perhaps leading to the creation of new β-lactamase inhibitors and β-lactam antibiotics.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Neu, H.C. Science 257, 1064–1073 (1992).
Levy, S.B. N. Engl. J. Med. 338, 1376–1378 (1998).
Walsh, C. Nature 406, 775–781 (2000).
Baggaley, K.H., Brown, A.G. & Schofield, C.J. Nat. Prod. Rep. 14, 303–333 (1997).
Jensen, S.E. & Paradkar, A.S. Antonie van Leeuwenhoek 75, 125–133 (1999).
Bachmann, B.O., Li, R. & Townsend, C.A. Proc. Natl. Acad. Sci. USA 95, 9082–9086 (1998).
Bachmann, B.O. & Townsend, C.A. Biochemistry 39, 11187–11193 (2000).
McNaughton, H.J. et al. Chem. Commun. 21, 2325–2326 (1998).
Roach, P.L. et al. Nature 387, 827–830 (1997).
Burzlaff, N.I. et al. Nature 401, 721–724 (1999).
Li, R., Stapon, A., Blanchfield, J.T. & Townsend, C.A. J. Am. Chem. Soc. 122, 9296–9297 (2000).
McGowan, S.J., Bycroft, B.W. & Salmond, G.P.C. Trends Microbiol. 6, 203–208 (1998).
Scofield, M.A., Lewis, W.S. & Schuster, S.S. J. Biol. Chem. 265, 12895–12902 (1990).
Richards, N.G.J. & Schuster, S.M. Adv. Enzymol. Relat. Areas Mol. Biol. 72, 145–198 (1998).
Zalkin, H. & Smith, J.L. Adv. Enzymol. Relat. Areas Mol. Biol. 72, 87–143 (1998).
Larsen, T.M. et al. Biochem. 38, 16146–16157 (1999).
Boehlein, S.K., Richards, N.G.J. & Schuster, S.M. J. Biol. Chem. 269, 7450–7457 (1994).
Jones, S. & Thornton, J.M. Proc. Natl. Acad. Sci. USA 93, 13–20 (1996).
Kleywegt, G.J. & Jones, T.A. Acta Crystallogr. D 50, 178–185 (1994).
Altschul, S.F. et al. Nucleic Acids Res. 25, 3389–3402 (1997).
Chakrabarti, R. & Schuster, S.M. Int. J. Pediatr. Hematol. Oncol. 4, 597–611 (1997).
Bruice, T.C. & Benkovic, S.J. Biochemistry 39, 6267–6274 (2000).
Bürgi, H.B. & Dunitz, J.D. Acc. Chem. Res. 16, 153–161 (1983).
Hendrickson, W.A., Horton, J.R. & LeMaster, D.M. EMBO J. 9, 1665–1672 (1990).
Elson, S.W. et al. J. Chem. Soc. Chem. Commun. 15, 1212–1214 (1993).
Otwinowski, Z. & Minor, W. Methods Enzymol. 276, 307–326 (1997).
Brünger, A.T. et al. Acta Crystallogr. D 54, 905–921 (1998).
McRee, D.E. J. Struct. Biol. 125, 156–165 (1999).
Laskowski, R.A. J. Appl. Crystallogr. 26, 283–291 (1993).
Kraulis, P.J. J. Appl. Crystallogr. 24, 946–950 (1991).
Merritt, E.A. & Bacon, D.J. Methods Enzymol. 277, 505–524 (1997).
Nicholls, A., Sharp, K.A. & Honig, B. Proteins 11, 281–296 (1991).
Esnouf, R.M. J. Mol. Graph. Model. 15, 132–134 (1997).
Collaborative Computational Project, Number 4. Acta Crystallogr. D 50, 760–763 (1994).
Acknowledgements
This work was supported by funds from the David and Lucile Packard Foundation to A.C.R., by an NIH grant to C.A.T. and in part by an NIH training grant to M.T.M. The DND-CAT Synchrotron Research Center at the Advanced Photon Source is supported by the E.I. DuPont de Nemours & Co., The Dow Chemical Company, the NSF and the State of Illinois.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Miller, M., Bachmann, B., Townsend, C. et al. Structure of β-lactam synthetase reveals how to synthesize antibiotics instead of asparagine. Nat Struct Mol Biol 8, 684–689 (2001). https://doi.org/10.1038/90394
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/90394
This article is cited by
-
Targeting adenylate-forming enzymes with designed sulfonyladenosine inhibitors
The Journal of Antibiotics (2019)
-
Origins of the β-lactam rings in natural products
The Journal of Antibiotics (2013)
-
Biosynthesis of clavam metabolites
Journal of Industrial Microbiology and Biotechnology (2012)
-
Clavulanic acid biosynthesis and genetic manipulation for its overproduction
Applied Microbiology and Biotechnology (2010)
-
Regulation and biosynthesis of carbapenem antibiotics in bacteria
Nature Reviews Microbiology (2005)