Abstract
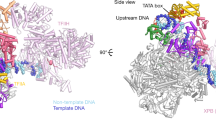
RNA polymerase from the hyperthermophile archaeon Pyrococcus furiosus (Pfu) forms specific and transcriptionally active complexes with its conjugate transcription factors TBP (the archaeal TATA binding protein homolog) and TFB (the archaeal homolog of eukaryotic RNA polymerase II and III transcription factors TFIIB and Brf) at the Pfu glutamate dehydrogenase promoter. A photochemical crosslinking method was used to map the vicinity of the catalytic subunits of Pfu RNA polymerase to DNA locations distributed along the polymerase–promoter interface. The largest component of this archaeal polymerase is split into two subunits, A′ and A″, whose relatively sharp boundary of DNA crosslinking (probed on the transcribed strand) is centered five to six base pairs downstream of the transcriptional start site. A strong argument based on this information, on the well-defined homology between the core bacterial, archaeal and eukaryotic RNA polymerase subunits, and on the recently determined structure of a bacterial RNA polymerase specifies the directionality of DNA in the archaeal transcription complex and its trajectory downstream of the transcriptional start site.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Langer, D., Hain, J., Thuriaux, P. & Zillig, W. Proc. Natl. Acad. Sci. USA 92, 5768–5772 ( 1995).
Thomm, M. FEMS Microbiol. Rev. 18, 159–171 (1996).
Bell, S.D. & Jackson, S.P. Cold Spring Harb. Symp. Quant. Biol. 63, 41–51 ( 1998).
Berghöfer, B. et al. Nucleic Acids Res. 16, 8113– 8128 (1988).
Zhang, G. et al. Cell 98, 811–824 (1999).
Hethke, C., Geerling, A.C.M., Hausner, W., De Vos, W.M. & Thomm, M. Nucleic Acids Res. 24, 2369–2376 (1996).
Qureshi, S.A., Bell, S.D. & Jackson, S.P. EMBO J. 16, 2927– 2936 (1997).
Hausner, W., Wettach, J., Hethke, C. & Thomm, M. J. Biol. Chem. 271, 30144–30148 (1996).
Kosa, P.F., Ghosh, G., Dedecker, B.S. & Sigler, P.B. Proc. Natl. Acad. Sci. USA 94, 6042– 6047 (1997).
Littlefield, O., Korkhin, Y. & Sigler, P.B. Proc. Natl. Acad. Sci. USA 96, 13668–13673 (1999).
Bartholomew, B., Tinker, R.L., Kassavetis, G.A. & Geiduschek, E.P. Methods Enzymol. 262, 476–494 (1995).
Hethke, C., Bergerat, A., Hausner, W., Forterre, P. & Thomm, M. Genetics 152, 1325–1333 (1999).
Straney, D.C. & Crothers, D.M. Cell 43, 449–459 (1985).
Mayer, A.N. & Barany, F. Gene 153, 1–8 (1995).
Yang, S.W. & Nash, H.A. Proc. Natl. Acad. Sci. USA 91, 12183–12187 (1994).
Lagrange, T. et al. Proc. Natl. Acad. Sci. USA 93, 10620 –10625 (1996).
Persinger, J., Sengupta, S.M. & Bartholomew, B. Mol. Cell. Biol. 19, 5218– 5234 (1999).
Fu, J. et al. Cell 98, 799–810 (1999).
Cramer, P. et al. Science 288, 640–649 (2000).
Poglitsch, C.L. et al. Cell 98, 791–798 (1999).
Nudler, E., Avetissova, E., Markovtsov, V. & Goldfarb, A. Science 273, 211–217 ( 1996).
Naryshkin, N., Revyakin, A., Kim, Y.G., Mekler, V. & Ebright, R.H. Cell 101, 601– 611 (2000).
Korzheva, N. et al. Science, 289, 619– 625 (2000).
Bartholomew, B., Durkovich, D., Kassavetis, G.A. & Geiduschek, E.P. Mol. Cell. Biol. 13, 942–952 (1993).
Korzheva, N., Mustaev, A., Nudler, E., Nikiforov, V. & Goldfarb, A. Cold Spring Harb. Symp. Quant. Biol. 63, 337–345 (1998).
Landick, R. Science 284, 598–599 ( 1999).
Kim, T.K. et al. Proc. Natl. Acad. Sci. USA 94, 12268– 12273 (1997).
Microbial genomes BLAST data base at NCBI: http://www.ncbi.nlm.nih.gov/Microb_blast/unfinishedgenome.html
Lannutti, B.J., Persinger, J. & Bartholomew, B. Biochemistry 35, 9821– 9831 (1996).
Forget, D. et al. Proc. Natl. Acad. Sci. USA 94, 7150 –7155 (1997).
Robert, F. et al. Mol. Cell. 2, 341– 351 (1998).
Kassavetis, G.A., Kumar, A., Ramirez, E. & Geiduschek, E.P. Mol. Cell. Biol. 18, 5587–5599 ( 1998).
Sayle, R. & Milner-White, E.J. Trends Biochem. Sci. 20, 374 (1995).
Acknowledgements
We are grateful to S.A. Darst for providing coordinates of Taq RNA polymerase, to V. Nikiforov for sharing data on sequence alignments, to W. Liu for generous help in generating Fig. 3, to J. Buschdorf and B. Goede for valuable materials and to G.A. Kassavetis for advice. Research support at UCSD from the NIGMS, at Kiel from the DFG and the Fonds der Chemischen Industrie, and a National Research Service postdoctoral fellowship from the NIH to M.S.B. are also gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bartlett, M., Thomm, M. & Geiduschek, E. The orientation of DNA in an archaeal transcription initiation complex . Nat Struct Mol Biol 7, 782–785 (2000). https://doi.org/10.1038/79020
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/79020
This article is cited by
-
Machine learning and statistics shape a novel path in archaeal promoter annotation
BMC Bioinformatics (2022)
-
Mechanisms of Evolutionary Innovation Point to Genetic Control Logic as the Key Difference Between Prokaryotes and Eukaryotes
Journal of Molecular Evolution (2015)
-
Finding the right spot to start transcription
Nature Structural & Molecular Biology (2007)